Pseudotyped-VSV particles for the analysis of viral entry
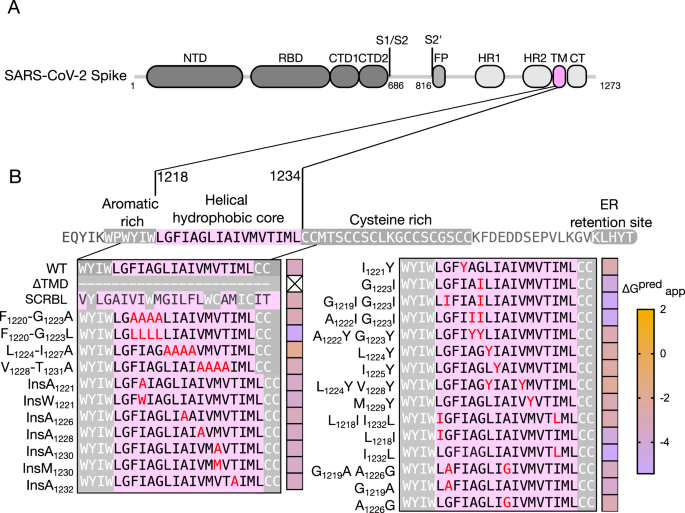
The SARS-CoV-2 S protein contains a single TMD in the C-terminal region of the protein (Fig. 1A). It spans 23 amino acids from Trp1214 to Cys1236 with a predicted ΔGapp of −3.4 kcal/mol for its insertion into the membrane, according to the ΔG Prediction Server20,21. The predicted TMD includes an aromatic-rich region (Trp1214-Tyr-Ile-Trp1217) in its N-terminal end and a Cys pair (Cys1235–Cys1236) in its C-terminal end. However, the structure of the S trimer in its post-fusion state19 shows a helical secondary structure from Leu1218 to Leu1234. Although the aromatic region and the C-terminal Cys-rich section on the TMD most likely interact with the membrane either as an interfacial membrane proximal region19,22 or through palmitoyl post-translational modifications23,24,25,26, in the present study, we focus exclusively on the helical hydrophobic core of the TMD, that is, from Leu1218 to Leu1234. The sequence of all the mutations included in this paper and their insertion potential can be found in Fig.1B.
A Schematic representation of SARS-CoV-2 Spike protein domain architecture. The position of key domains is highlighted: NTD N-terminal domain, RBD receptor binding domain, CTD1 C-terminal domain 1, CTD2 C-terminal domain 2, S1/S2, S1/S2 cleavage site, S2′, S2′ cleavage site, FP fusion peptide, HR1 heptad repeat 1, HR2 heptad repeat 2, TM transmembrane domain, CT cytoplasmic tail. B Sequence of the SARS-CoV-2 S protein TMD and the mutations included in this study. The sequence of the SARS-CoV-2 S protein from Glu1206 to Thr1273 is shown. Membrane-associated regions, including an aromatic-rich section and a cysteine-rich domain, are highlighted with light letters on a gray background. The TMD hydrophobic core is shown with dark letters on a pink background. The TMD, as predicted by the ΔG Prediction Server (WT), and the variations included in this work are highlighted. A heat map (right of the sequence) shows the predicted ΔGapp for each TMD mutant.
To study the role of the SARS-CoV-2 S protein TMD during viral entry, we utilized pseudotyped vesicular stomatitis virus (VSV) particles coated with different S protein mutants (Fig. 2A)10,15,27,28. Briefly, HEK293T cells were transfected with plasmids encoding the different S mutants and subsequently infected with a recombinant VSV in which the glycoprotein gene has been replaced with GFP (VSVΔG-GFP). Next, the infectivity of the resulting particles was assayed on VeroE6 cells expressing the TMPRSS2 cofactor by counting GFP-positive cells. VSVΔG-GFP particles were pseudotyped with the native SARS-CoV-2 Wuhan-Hu-1 S protein as a positive control and reference value (WT). No infection was observed when the TMD of the SARS-CoV-2 S protein was eliminated (ΔTMD) or was substituted by a scrambled version of the TMD (SCRBL) where the composition of the residues included in the TMD was preserved but their position was randomly varied, confirming the importance of the SARS-CoV-2 S protein TMD in the entry process and indicating a role beyond membrane anchoring (Fig. 2B). Expression of the chimeras included in this work can be found in Supplementary Fig. 1.
A Generation of VSVΔG-GFP pseudotyped particles. HEK293T cells were transfected with the corresponding SARS-CoV-2 S protein mutant and infected with a VSV lacking the G gene and expressing GFP (VSVΔG-GFP). The resulting virus, incorporating the S mutants on its surface (VSVΔG-GFP -S), was recovered from the media and used to infect Vero-TMPRSS2 cells. Viral infectivity was then analyzed by counting GFP-positive cells. B–D Focus forming units (FFU) measured after infection with VSVΔG-GFP pseudotyped with different mutants of the SARS-CoV-2 S protein TMD. All samples were normalized to the wild-type SARS-CoV-2 S protein (WT). The p-values (<0.05) for individual one-sample t-tests (vs. WT) are indicated above each bar. B FFU from VSVΔG-GFP particles decorated with the wild-type SARS-CoV-2 S protein (WT), the SARS-CoV-2 S protein where TMD was eliminated (ΔTMD), and a scrambled version of the SARS-CoV-2 S protein TMD (SCRBL). C FFU from pseudotyped VSVΔG-GFP with the SARS-CoV-2 S protein with substitutions of amino acid stretches, Phe1220 to Gly1223, Leu1224 to Ile1227, and Val1228 to Thr1231, with alanine (Phe1220-Gly1223A, Leu1224-Ile1227A, and Val1228-Thr1231A, respectively). Substitutions of amino acids Phe1220 to Gly1223 to leucine (Phe1220 to Gly1223L) were also included. D FFU from pseudotyped VSVΔG-GFP with the SARS-CoV-2 S protein, including alanine insertions at positions 1221, 1226, 1228, 1230, and 1232 (InsA1221, InsA1226, InsA1228, InsA1230, and InsA1232), tryptophan insertion at position 1221 (InsW1221), and methionine insertion at position 1230 (InsM1230). Four independent biological replicates were performed.
Stretches of four amino acids, corresponding to one helical turn, within the SARS-CoV-2 S protein TMD beginning at Phe1220 were substituted with alanine residues (Fig. 2C). Substitution of the Phe1220-Gly1223 and Leu1224-Ile1227 segments (F1220-G1223A and L1224-I1227A, respectively) significantly reduced the infectivity of pseudotyped VSVΔG-GFP particles. In contrast, substitution of Val1228-Thr1231 (V1228-T1231A) with alanines did not decrease viral entry; a non-significant increase was observed. To control for potential bias from alanine substitutions, the Phe1220-Gly1223 segment was also replaced with leucine residues (F1220-G1223L), yielding similar results.
These findings indicate that sequence and structural determinants relevant for viral entry are distributed throughout the TMD, with a greater contribution from the N-terminal region. Notably, none of the modifications tested significantly altered the predicted membrane insertion potential (ΔGapp) of the SARS-CoV-2 S protein TMD (Fig. 1B).
To further analyze the role of SARS-CoV-2 S protein TMD in the viral entry process, we inserted alanine residues in key positions of the hydrophobic segment (Fig. 2D). Inserting a residue not only increases the length of the TMD but also changes the relative orientation of the residues positioned before and after the insertion point due to the α-helical nature of this segment29. We first inserted an alanine between Phe1220 and Ile1221 (Ins-A1221), which alters the relative orientation of the aromatic membrane proximal region and the TMD. Additionally, Ins-Ala1221 disrupts the G1223xxxG1229 motif found in the TMD14, forcing a 100° rotation for Gly1219 relative to Gly1223 (Supplementary Fig. 2A vs. B). As shown in Fig. 2D, Ins-Ala1221 greatly reduced the infectivity of pseudo-typed VSVΔG-GFP particles, suggesting that either the relative orientation of the aromatic stretch vs the TMD, the GxxxG interaction motif, or both are important for particle entry.
Insertion of an alanine residue in positions 1226 or 1228 (Ins-A1226 and Ins-A1228) negatively affected viral entry, indicating that the relative orientation of the N-terminal and C-terminal sections of the TMD might be important for infectivity (Fig. 2D). These two insertion points were selected with the work of Fu and Chou16 in mind, who proposed a trimerization hydrophobic zipper involving residues 1221, 1225, 1229, and 1233. Thus, the observed reduction in viral entry could also result from disrupting the oligomerization motif they described.
Insertions at positions 1230 and 1232 (Ins-A1230 and Ins-A1232) also disturb the hydrophobic trimerization motif proposed by Fu and Chou (Supplementary Fig. 2), but reduced particle entry to a lesser extent than Ins-A1226 or Ins-A1228 (Fig. 2D). Of these two insertions, Ins-A1230 reduced pseudo-typed VSVΔG-GFP entry more than Ins-A1232, suggesting that the relative orientation of the N-terminal and C-terminal sections of the TMD is more important for the protein function than the trimerization hydrophobic zipper. Note that while the authors16 included leucine residues on positions 1229 and 1233, we incorporated the methionine residues found in the SARS-CoV-2 S protein consensus sequence (Uniprot: P0DTC2). Once again, to ensure that the observed results are not biased by the choice of Ala residues, we inserted a Trp at position 1221 (InsW1221) and a Met at position 1230 (InsM1230). Regarding the insertion of Ala or any other residue, we observed similar results (Fig. 2D).
A closer look at the SARS-CoV-2 TMD reveals a highly hydrophobic surface composed of Leu1218, Ile1221, Leu1224, Ile1225, Val1228, and Ile1232 and, an opposing surface rich in small residues including, Gly1219, Ala1222, Gly1223, and Ala1226 which could participate in TMD-TMD interactions15,27,28,29,30 (Fig. 3A). To analyze the importance of these regions, we first analyzed the impact of the GxxxG and the hydrophobic zipper as oligomerization motifs, we selectively mutated key residues in the SARS-CoV-2 S protein. First, we replaced Gly1223 with isoleucine (G1223I), as the equivalent substitution is sufficient to break the dimerization of Glycophorin A (GpA) TMD30,31. The GFP count associated with pseudo-typed VSVΔG-GFP entry revealed that the G1223I mutation is sufficient to reduce viral entry (Fig. 3B). Substituting Gly1219 and Gly1223 with isoleucine (G1219I G1223I) further decreases pseudo-typed VSVΔG-GFP infectivity. Alternatively, we replaced Ile1221, Ile1225, or Met1229 from the hydrophobic surface with tyrosine (I1221Y, I1225Y, and M1229Y), changes that were previously suggested to modify TMD homo-oligomerization16. While the I1221Y substitution slightly reduced viral entry, the I1225Y and M1229Y substitutions did not (Fig. 3B).
A Structural model of the SARS-CoV-2 S protein TMD. AlphaFold3 model of residues 1218-1234 (top) and helical wheel projection (bottom) of the same sequence. The surface of the helix containing small side chain residues is highlighted in orange, while the opposite surface containing highly hydrophobic residues is highlighted in purple. B FFU from pseudotyped VSVΔG-GFP with the SARS-CoV-2 S protein bearing the following point mutations Gly1223 to Ile, Gly 1219 to Ile and Gly1223 to Ile, Ile1221 to Tyr, Ile1225 to Tyr, Met1229 to Tyr, Ala1222 to Ile and Gly1223 to Ile, Ala1222 to Tyr, and Gly1223 to Tyr, Leu1224 to Tyr, Leu1224 to Tyr, and Val1228 to Tyr (G1223I, G1219I G1223I, I1221Y, I1225Y, M1229Y, A1222I G1223I, A1222Y G1223Y, L1224Y, L1224Y V1228Y, respectively). C Analysis of the Leu1218 and Ile1232 (L1218I I1232L) and Gly1219 and Ala1226 (G1229A A1226G) residue swapping. The single mutations associated with these residue swapping Leu1218, Ile1232, Gly1219, and Ala1226 (L1218I, I1232L, G1229A, and A1226G respectively) were also analyzed. Four independent biological replicates were performed.
Then, we replaced Ala1222 and Gly1223 with isoleucine (A1222I G1223I) and measured the impact on the pseudo-typed VSVΔG-GFP infectivity. Our results revealed that the A1222I G1223I mutant is less able to promote viral entry than the previously tested G1223I mutant (Fig. 3B). When Ala1222 and Gly1223 were replaced with tyrosine (A1222Y G1223Y), a larger and less hydrophobic amino acid, we observed a greater reduction in VSVΔG-GFP infectivity. Similarly, substituting Leu1224 with tyrosine (L1224Y) on the hydrophobic surface also reduced VSVΔG-GFP entry. Double substitution of Leu1224 and Val1228 to tyrosine (L1224Y V1228Y) diminished VSVΔG-GFP infectivity further. To further test our hypothesis, we swapped Leu1218 and Ile1232 (L1218I I1232L) on one hand and Gly1219 and Ala1226 (G1229A A1226G) on the other (Fig. 3C). Neither of these changes nor the single mutations associated with these residue swapping negatively affected VSVΔG-GFP entry. Therefore, according to our results, both the small residues and the hydrophobic bulky residues aligned on opposing faces of the helix (Fig. 3A) are relevant for the entry process, probably because they participate in protein-protein interactions or in TMD-lipid interactions. Protein levels for all the chimeras were probed by Western blotting to avoid misinterpretation of the data (Supplementary Fig. 1).
To incorporate the SARS-CoV-2 S protein into VSVΔG-GFP pseudo-particles, it must be located at the plasma membrane. Thus, we investigated whether those events in which no VSVΔG-GFP pseudo-particle infectivity was observed were the consequence of SARS-CoV-2 S protein absence at the cell surface. We determined whether the SARS-CoV-2 S protein was located at the plasma membrane by surface-staining with an anti-RBD antibody followed by flow cytometry analysis (Fig. 4). All SARS-CoV-2 S protein mutants that showed a decreased VSVΔG-GFP infectivity were included in this assay. Additionally, the WT S protein was used as a positive control while the ΔTMD was included as a negative control (Fig. 4A). Our results indicate that all SARS-CoV-2 S protein mutants were present at the plasma membrane at similar levels except for Ins-A1228 (Fig. 4B and Supplementary Fig. 3A). Thus, validating the results of the VSVΔG-GFP pseudo-particle assay. The absence of Ins-A1228 at the cell surface suggests that the SARS-CoV-2 S protein TMD might play a role in protein trafficking.
A Spike surface expression of WT and ΔTMD controls was analyzed by flow cytometry. HEK293T cells were transfected with plasmids encoding the corresponding S protein alongside a constitutively GFP-expressing plasmid. After 48 h, cells were stained with an anti-RBD antibody. Levels of the surface SARS-CoV-2 S protein (red) and GFP (green) were analyzed by flow cytometry. B Relative surface expression of all Spike mutants tested in this assay. GFP was used as a transfection control. The figure shows the percentage of SARS-CoV-2 S protein vs GFP signal ratio. Samples were normalized to the SARS-CoV-2 S protein WT. The significative p-values (<0.05) for individual one-sample t-tests vs. WT are indicated above each bar. Three independent biological replicates were performed.
BiMuC assay for the analysis of membrane fusion
The role of the SARS-CoV-2 S protein TMD during the viral and cellular membrane fusion process was corroborated with a syncytia-based assay known as BiMuC32,33. Briefly, the Venus fluorescent protein (VFP) can be split into two fragments, VN and VC, neither of which is fluorescent. However, when these two fragments are fused to a pair of interacting proteins that bring them together, such as cJun and bFos, the structure of the VFP is reconstituted and the fluorescence is recovered. The VN-cJun and VC-bFos chimeras were expressed in separate cell pools together with the viral machinery required for membrane fusion. Therefore, the two chimeras would only meet in the event of membrane fusion (Fig. 5A). The complete SARS-CoV-2 S WT protein was used as a positive control and reference value. The ΔTMD and SCRBL were included as negative controls (Fig. 5C). Measurements of the area of the syncytia can be found in Supplementary Fig. 3B.
A Schematic representation of the BiMuC assay. The Venus fluorescence protein (VFP) can be split into two fragments, VN and VC, respectively, neither of which is fluorescent. These two fragments were fused to cJun and bFos (VN-cJun and VC-bFos) and transfected into HEK 293 T cells in separate cell pools. Cells were co-transfected with the viral machinery required for membrane fusion, the corresponding SARS-CoV-2 S protein, and the ACE2 receptor. Additionally, both cell pools were transfected with the Renilla Luciferase (luciferase) to normalize the fluorescent values. Only functional SARS-CoV-2 S proteins facilitated the S-ACE2 recognition and membrane fusion, allowing VFP reconstitution and fluorescence. B Representative images of the assay where cells have been transfected with the SARS-CoV-2 S protein (WT) or the SARS-CoV-2 S protein bearing a scrambled version of the TMD (SCRBL). Scale bar = 100 microns. C–E The fluorescence/luciferase signal ratio (GFP/Luc ratio) was measured for the SARS-CoV-2 S protein mutants tested on this assay. Samples were normalized to the SARS-CoV-2 S protein WT. Significative p-values (<0.05) for individual one-sample t-tests are indicated above each bar. C GFP/Luc ratio for the SARS-CoV-2 S protein (WT), the SARS-CoV-2 S protein where TMD was eliminated (ΔTMD), and a chimera bearing a scrambled version of the SARS-CoV-2 S protein TMD (SCRBL). As a negative control, we also included cells that did not incorporate the SARS-CoV-2 S protein (ΔS). D GFP/Luc ratio for the SARS-CoV-2 S protein with substitutions of 4 amino acid stretches Phe1220 to Gly1223, and Val1228 to Thr1231, by alanines (Phe1220-Gly1223A and Val1228-Thr1231A), and the SARS-CoV-2 S protein including alanine insertions in positions 1221, 1228, and 1232 (InsA1221, InsA1228, and InsA1232). E GFP/Luc ratio for the SARS-CoV-2 S protein bearing the following point mutations: Gly 1219 to Ile and Gly1223 to Ile, Ile1221 to Tyr, Ile1225 to Tyr, Met1229 to Tyr, Ala1222 to Tyr and Gly1223 to Tyr, and Leu1224 to Tyr and Val1228 to Tyr (G1219I G1223I, I1221Y, I1225Y, M1229Y, A1222Y G1223Y, and L1224Y V1228Y, respectively). All panels include the WT values used to normalize the data (dotted line) and the SCRBL samples. F Correlation of FFU measured in the VSVΔG-GFP infection assay against values of reconstituted GFP/Luc ratio in the BiMuC fusion assay. Colors show samples that have lower signals than WT values in both systems (pink background), higher in both systems (green background), or different between the two assays (white background). At least three independent biological replicates were performed.
Next, we selected some of the previously described mutants of the S protein and tested them using the BiMuC methodology. Substituting the Phe1220-Gly1223 helical turn with alanines (F1220-G1223A) diminished syncytia formation, though not significantly (Fig. 5D). On the other hand, V1228-T1231A did not reduce syncytia formation (Fig. 5D). Alanine insertions Ins-A1221 altered the observed syncytia-derived fluorescence (Fig. 5D). However, only a small non-significant reduction in syncytia formation was observed for Ins-A1232. We also tested the influence of point mutations on the ability of the SARS-CoV-2 S protein to induce syncytia formation (Fig. 5E). In this case, the G1219I G1223I, I1221Y, or the M1229Y substitutions did not perturb the SARS-CoV-2 S protein function. However, I1225Y, A1222Y G1223Y, and L1224Y V1228Y reduced the formation of syncytia. Altogether, the BiMuC-derived results correlate with the pseudo-typed VSVΔG-GFP assay34 (Fig. 5F).
Analysis of TMD oligomerization
Some of the modifications included in the BiMuC and the pseudo-typed VSVΔG-GFP assay could alter the previously described homo-oligomerization of the SARS-CoV-2 TMD12,16,35. To directly analyze the SARS-CoV-2 TMD potential oligomer, we used a bimolecular fluorescent complementation (BiFC) approach adapted to study intramembrane interactions36,37,38,39. Briefly, the tested TMDs were fused to a split VFP, either to its N-terminal (VN) or its C-terminal (VC). If the interaction between the tested TMDs occurs, it will bring the VN and VC ends together, facilitating the reconstitution of the VFP structure and causing fluorescence (Fig. 6A). The TMD of GpA was used as a positive control and normalization value across experimental replicates. The TMD of Tomm20, a monomeric hydrophobic segment, was used as a negative control38,40.
A Reconstitution of the Venus fluorescent protein (VFP) mediated by the SARS-CoV-2 S protein TMD oligomerization. Different S TMD mutants were fused to both the N and C terminal sections of the VFP (VN and VC, respectively) and expressed in HEK 293T cells together with the Renilla Luciferase. If the SARS-CoV-2 S protein TMD can oligomerize the VFP will be reconstituted B GFP/Luc ratios measured for different chimeras bearing Glycophorin A TMD (GpA), E. coli Lep H2, wild-type SARS-CoV-2 S protein TMD (WT), and the SARS-CoV-2 S protein TMD with the following point mutations G1219I Gly1223 to Ile, Gly1219 to Tyr and Gly1223 to Tyr, Ala1222 to Tyr and Gly1223 to Tyr, Ala1222 to Tyr, Gly1223 to Ile, le1221 to Tyr, Ile1225 to Tyr, Met1229 to Tyr, and Leu1224 to Tyr and Val1228 to Tyr (G1219I G1223I, G1219Y G1223Y, A1222Y G1223Y, A1222Y, G1223I, I1221Y, I1225Y, M1229Y, and L1224Y V1228Y, respectively). All values are normalized against the GpA homo-oligomer. The significant p-values (<0.05) for individual one-sample t-tests are indicated above each bar. At least three independent biological replicates were performed.
Using this BiFC assay, we tested the SARS-CoV-2 TMD. Our results indicate that this TMD is sufficient for inducing oligomerization (Fig. 6B). Next, we focused on the role of the surface with small residues on the oligomerization of the TMD. The G1219I G1223I substitution did not decrease the SARS-CoV-2 S protein’s TMD oligomerization potential; however, the G1219Y G1223Y did, a result that correlates with the outcome of the BiMuC assay. The A1222Y G1223Y double substitution further reduced oligomerization. Interestingly, the substitution of only Ala1222 or Gly1223 with Y (A1222Y and G1223Y) did not alter the oligomerization capabilities of the SARS-CoV-2 TMD, suggesting a synergistic effect of these two residues.
We also tested the role of the bulky hydrophobic helix interface on the SARS-CoV-2 S protein TMD on oligomerization. The single substitutions I1221Y, I1225Y, or M1229Y did not alter oligomerization. Furthermore, the double substitution L1224Y V1228Y, which had proven fundamental for VSVΔG-GFP entry and membrane fusion in the BiMuC assay, did not decrease the observed fluorescence, suggesting that the hydrophobic surface does not participate in the oligomerization of the SARS-CoV-2 S protein TMD.
We modeled the potential SARS-CoV-2 S protein TMD homo-trimer using TMHOP41. TMHOP uses Rosetta symmetric all-atom ab initio fold-and-dock simulations in an implicit membrane environment to predict low-energy conformations based on the empirical measurement of amino acid insertion propensities into E. coli inner membrane42. These conformations are clustered based on energy and structural properties (Supplementary Fig. 4A), and a representative model of each cluster is shown (Fig. 7A). In each model, Gly1216, Ala1222, Gly1223, and Ala1226 are located on the interaction surface of the potential trimer, confirming our experimental results. We also modeled the potential TMD trimer using AlphaFold343, a recent evolution of the AlphaFold2 architecture and training procedure, which is capable of predicting the joint structure of complexes. Once again, the predicted models present, in most cases, Gly1216, Ala1222, Gly1223, and Ala1226 at the interior of a TMD homo-trimer (Fig. 7B and Supplementary Fig. 4). The AlphaFold3 server enabled us to model larger sequences, including the full-length S protein. When the TMD region was modeled in the context of the full-length protein, accuracy was low, and thus the models were discarded (Supplementary Fig. 5). However, the server modeled with sufficient confidence the TMD flanked by the N-terminal aromatic-rich region and the Cys-rich C-terminal end. The retrieved models for this section included, once again, Gly1216, Ala1222, Gly1223, and Ala1226 on the internal surface of a TMD trimer (Fig. 7C and Supplementary Fig. 4).
A and B SARS-CoV-2 S protein TMD trimer modeled by the TMHOP server (A) and by the AlphaFold3 server (B). The top five models are shown. The input sequence was 1212WYIWLGFIAGLIAIVMVTIMLCC1235. Models show the TMD hydrophobic core, residues 1218-1234 shown in pink. The secondary structure, as well as the surface of the peptide, is shown. Gly1216, Ala1222, Gly1223, and Ala1226 are highlighted in red on the secondary structure representation. C AlphaFold3 server model of the C-terminal region of the SARS-CoV-2 S protein, including the aromatic-rich section, the TMD, and the cysteine-rich domain. The five top models are shown. Models include just the TMD hydrophobic core, residues 1218-1234 shown in pink. The secondary structure, as well as the surface of the TMD hydrophobic core, is shown. Gly1216, Ala1222, Gly1223, and Ala1226 are highlighted in red.