PNP was identified to play hub roles in viral-host interaction upon H1N1 infection
In order to discover novel regulators within the viral-host interaction, the peripheral plasma of 41 IAV-infected patients and 19 healthy volunteers were collected and subjected to proteomics analysis (Fig. 1a). Compared to the healthy controls, 325 differentially expressed proteins (DEPs) were identified in IAV-infected patients, which were enriched in multiple metabolic pathways based on Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis (Fig. 1a, c).
PNP was identified as a potential hub gene in IAV infection via VI-PPI network. a Overview of the workflow for the proteomic analysis of the plasma derived from H1N1-infected patients and healthy controls. The peripheral plasma of 19 healthy controls and 41 H1N1-infected patients were collected, followed by proteomics analysis. b The volcano map of the plasma proteomic data. The differentially expressed proteins (DEPs) were labeled as blue (downregulation) and red (upregulation). P < 0.05, fold change > 1.5 or <0.6667. c The KEGG pathway enrichment analysis of DEGs between the healthy and H1N1-infected individuals. The top 15 KEGG pathways enriched by DEGs (H1N1-infected vs. Healthy controls) were displayed. d Data collection for VI-PPI. VI-PPI network was constructed on the basis of protein-protein interaction, containing molecules related to viral infection, host inflammation, and drug targets, with 6449 nodes (genes) and 74103 edges (PPI pairs). e The landscape of VI-PPI. A total of 22 functional domains were enriched and marked with different colors by SAFE tools in Cytoscape. Each cluster was annotated with the most relevant gene ontology (GO) term. f The algorithm for the identification of the key hub genes during influenza progression. Hub genes by VI-PPI network, defined as genes that were significantly located at the shortest paths from H1N1-related genes to pneumonia-related genes, were extracted by calculating the local betweenness centrality (the upper panel). g The Venn plot of shared hub genes among infection of different strains of influenza virus by VI-PPI. 348 hub genes with potential broad-spectrum among the 4 strains of influenza virus (H1N1, H7N7, H5N1, H3N2, Supplementary Fig. 1e) by VI-PPI prediction were used to overlap with a total of 325 differentially expressed proteins (DEPs) derived from the plasma proteome of clinical cohort (H1N1 infected vs. healthy control). 18 overlapped genes were identified by Venn plot. h The overview of the expression of the overlapping genes by the plasma proteomic data of the clinical cohort. The expression of the 18 overlapping DEPs based on the proteomic data of H1N1-infected vs. healthy control was shown by box plot. i, j The ScRNA-seq analysis of lungs derived from the control and H1N1 mice models. The scRNA-seq data of H1N1-infected mice were obtained from previously published data (https://ngdc.cncb.ac.cngsabrowse/CRA013573). t-SNE plot revealed 19 clusters, and the expression of the 18 overlapping DEPs in alveolar type 2 (AT2) cells was analyzed. Differentially expressed genes (DEGs) with significance were labeled as red. All data are presented as mean ± SD; unless otherwise indicated, N = 3 biologically independent experiments; Statistical analysis was performed by two-tailed Student’s t-test (h) and Wilcoxon rank-sum test (j). *P < 0.01 versus Mock (j)
To efficiently explore nexus molecules within the viral-host interaction during IAV infection from a large quantities of candidate molecules, we firstly created a screening model called Viral-Inflammation Protein-protein Interaction network (VI-PPI), through manually collecting genes related to viral infection and host inflammation, as well as high-quality physical PPI pairs, from multiple published datasets (Fig. 1d, e, Supplementary Fig. 1a, b, and Supplementary Table 1). As shown in Fig. 1e, all the nodes in VI-PPI were enriched in pathways associated with fundamental homeostasis maintenance function based on SAFE analysis (Supplementary Table 2), yielding a scale-free network,26 with higher density compared to other published PPI networks (Supplementary Fig. 1c, d). Secondly, we proposed a novel strategy for screening potential host targets with hub roles. The hub genes were defined as genes that exhibited significance calculated by local betweenness centrality (see Methods) and located at the shortest paths between host factors interacting with IAV proteins and molecules related to pneumonia. Then, to gain insights into broad-spectrum host targets, hub genes that might function during the infection of four subtypes of IAV, including H5N1 (A/goose/Guangdong/1/1996), H3N2 (A/NewYork/392/2004), and H7N7 (A/mallard duck/Korea/VI142063/2014), were screened via VI-PPI network, and 348 shared genes were identified (Supplementary Fig. 1e).
Furthermore, by comparing the DEPs lists from IAV-infected patients with the 348 shared genes (Fig. 1f), 18 overlapping genes were identified (Fig. 1g, h). Since alveolar type II (AT2) cells were the primary targets for IAV, we further focused on the change of the 18 genes in AT2 cells by the previously published scRNA-seq data of the lungs derived from the control and H1N1-infected mice (Fig. 1i).27 Galectin 3 binding protein (LGALS3BP), granulin precursor (GRN), C-C motif chemokine ligand 5 (CCL5), secretory leukocyte peptidase inhibitor (SLPI), and purine nucleoside phosphorylase (PNP) were revealed with the most significant variations (Fig. 1j). LGALS3BP, GRN, CCL5, and SLPI have been suggested as anti-IAV targets,28,29,30,31 yet no data has been reported regarding the potential role of PNP in IAV infection. Moreover, since the investigation of purine salvage, an energy-efficient manner for nucleotides production, has not been adequately addressed, PNP, as the critical enzyme in purine salvage, was selected for further study.
PNP contributed to H1N1 replication in AECs both in vitro and in vivo
Furthermore, the expression of PNP in the lung tissues of mice infected with H1N1, H3N2, and H5N1 viruses, respectively, were analyzed, showing a significant increase at both the mRNA and protein levels (Supplementary Fig. 1f). We then evaluated the effect of PNP on H1N1-infected AECs by interfering PNP expression through either siRNA transfection or PNP-overexpression vector (Supplementary Fig. 1g, h). PNP knockdown significantly decreased the viral titer and enhanced cell viability in response to H1N1, compared to the mock-infected cells, whereas overexpression of PNP increased the viral titer and reduced cell viability (Fig. 2a, b). We further employed fluorescence in situ hybridization assay (FISH) assays to verify the association of PNP with viral replication. Inhibition of PNP significantly diminished viral genomic RNA (vRNA) expression in both challenged A549 and BEAS-2B cells, whereas overexpression of PNP increased vRNA expression (Fig. 2c, d).
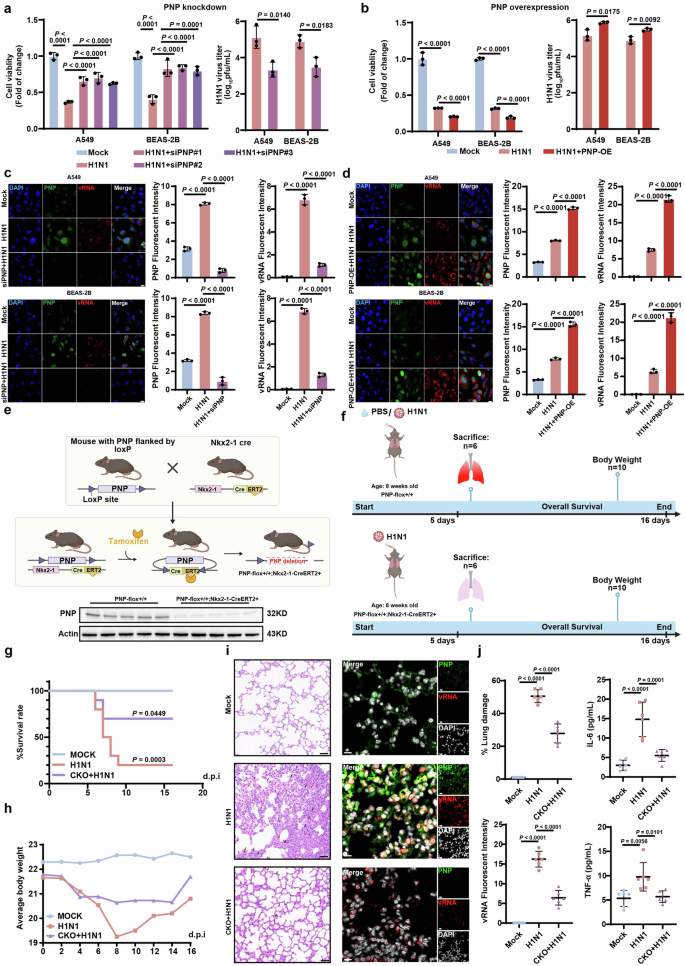
PNP knock down/knock out in alveolar epithelial cells (AEC) exhibit anti-influenza capacity in vitro/in vivo. The effect of PNP inhibition (a) or PNP overexpression (b) on cell viability and virus titer upon 24 h post H1N1 infection. A549 or BEAS-2B cells were transfected with NC/PNP siRNA at 30 pmol (a) or 2 µg control vector/PNP-overexpression vector (b) before challenged with H1N1 (MOI = 5). Cell viability and viral titer of H1N1-infected A549 cells and BEAS-2B cells with or without PNP knockdown/overexpression were evaluated at 24 h post-infection. FISH and IF analysis of vRNA and PNP expression in alveolar epithelial cells with or without PNP knockdown/overexpression upon H1N1. Both A549 and BEAS-2B cells were transfected by NC/PNP siRNA (c) or control vector/PNP-overexpression vector (d), followed by FISH of H1N1 viral genomic RNA (vRNA) and IF assay of PNP, respectively. DAPI was used to stain nuclei of the cells (Scale bar: 20 μm). e Schematic representation of the generation of AEC-specific PNP conditional knockout mice models. PNP expression in the lung tissue of PNPflox/flox and Nkx2-1-CreERT2, PNPflox/flox mice was analyzed by Western blot. f Diagram of the experimental procedures. Briefly, 8-week-old PNPflox/flox mice and PNP conditional knockout mice were intranasally challenged with 2LD50 (h) or 4LD50 (g) H1N1. The survival rate and body weight change upon IAV infection. The survival rate (g) and body weight change (h) of the different groups (n = 10) were observed daily for a course of 16 days. The inflammation lesion and viral infection within the lungs. After 6 days of infection, 6 mice from each group were sacrificed, with the lungs dissociated and subjected to H&E staining (N = 6, Scale bar: 50μm, (i), FISH of vRNA and IF assay of PNP. DAPI was used to indicate nucleus (N = 6, Scale bar: 10 μm, i). The area of lung damage and intensity of vRNA were shown. The concentration of TNF-ɑ and IL-6 levels in mice plasma (N = 6) was measured by ELISA on day 6 post infection (j). All data are presented as mean ± SD; unless otherwise indicated, N = 3 biologically independent experiments; Statistical analysis was performed by one-way ANOVA (c, d, j), two-tailed Student’s t test (a, b) and Log-rank test (g)
Next, we validated the in vivo role of PNP during IAV infection. Tamoxifen (TAM)-inducible AEC-specific homozygous PNP conditional knockout (CKO) mice (Nkx2-1-CreERT2, PNPflox/flox) were established by mating Nkx2-1-CreERT2 mice with PNPflox/flox mice (Fig. 2e). TAM (75 mg/Kg) was administered to 6-week-old PNP-CKO and PNPflox/flox mice through intraperitoneal injection once a day for 6 consecutive days. One week later, PNP deficiency in lung was determined by Western blot (Fig. 2e). PNP-CKO mice intranasally infected with H1N1 (Fig. 2f) displayed a significantly extended survival rate and an accelerated body weight restoration when compared to PNPflox/flox mice (Fig. 2g, h). Pulmonary pathological injury was also alleviated in PNP-CKO mice at 6 days post-infection (dpi), evidenced by reduced hemorrhage, necrocytosis of epithelial cells, infiltration of inflammatory cells within the lung tissues, and suppressed proinflammatory cytokines production (IL-6, TNF-ɑ) in the peripheral blood (Fig. 2i, j). Moreover, vRNA was significantly diminished in the lungs of PNP-CKO mice by FISH analysis (Fig. 2i, j).
Taken all these lines of evidence together, our results indicated that the VI-PPI-predicted PNP played critical roles in H1N1 infection in both in vivo and in vitro models.
PB1-F2 of IAV acted as a novel vTF to activate host PNP by TATA box binding
Subsequently, we tried to unveil the molecular mechanism of H1N1-induced PNP upregulation. The genome of viruses encodes multiple proteins to strictly control viral replication and transmission to other host cells, among which viral transcription factors (vTFs) deserve more attention as they were responsible for regulating host genes.32 To reveal the potential association between H1N1 proteins and host PNP, A549 cell lines stably expressing the control pLVX-Puro vector and various of H1N1 protein (HA, NA, NP, PA, PB1, PB1-F2, PB2, NS1, M1, M2) were established (Fig. 3a). The mRNA and protein level of PNP in these cell lines were analyzed, showing that PB1-F2 overexpression led to a significant increase of PNP expression (Fig. 3b, c). Moreover, the PB1-F2-overexpressing vector for H5N1, H3N2, and H7N7 which were also used in the screening of the broad‐spectrum target in the VI-PPI network (Supplementary Fig. 1e) was transfected into A549 cells, respectively, revealing that the increase of PNP by viral PB1-F2 was universal among different subtypes of IAV (Fig. 3d, e).
IAV PB1-F2 transcriptionally activated PNP expression by binding to the TATA-box of the promoter. a Schematics of the establishment of A549 cells stably transfected by various of H1N1 proteins and the transient transfection of PB1-F2-overexpressing vectors to A549 cells. A549 cells stably overexpressed different H1N1 proteins were used to screen H1N1 protein most relevant to PNP expression. Moreover, PB1-F2-overexpressing vectors of other strains of IAV (H5N1, H7N7, H3N2) were transiently transfected into A549 cells to analyze their effect on PNP expression. The relative expression of PNP in A549 cells stably expressed different H1N1 proteins. The mRNA (b) and protein (c) level of PNP were quantified by qRT-PCR and Western blot, respectively. The effect of PB1-F2 proteins of other IAV strains ((H5N1, H7N7 and H3N2)) on PNP expression. A549 cells were transiently transfected with vectors overexpressing PB1-F2 of other strains of IAV for 24 h, followed by examining the mRNA (d) and protein (e) level of PNP via qRT-PCR and Western Blot analysis, respectively. f, g Effect of PB1-F2 on the luciferase activity of PNP-promoter reporter gene by the dual luciferase assay. The PB1-F2-overexpressing plasmid was co-transfected with pGL3-PNP-promoter reporter gene and Renilla luciferase reporter in 293 T cells for 24 h. Renilla luciferase activities were used as an internal reference. The assay was performed in triplicate. The values are expressed as mean ± SD. h Diagram of motif prediction based on Alphafold 3 and DeepPBS. The exact binding site of PB1-F2 on PNP promoter was predicted by combining Alphafold 3 and DeepPBS analysis. i Schematics of the −50 ~ + 1 sequence of PNP promoter. The TATA-box was shown in red. j Predicted PB1-F2-binding motif on PNP promoter within the −50 ~ + 1 sequence. The PB1-F2-binding consensus sequence is shown in red. k Dual luciferase assay of the wild type or TATA-box-mutated PNP luciferase reporter plasmid upon PB1-F2 overexpression. The PNP luciferase reporter plasmid containing the predicted motif (50 bp) with or without the mutated TATA-box were co-transfected with PB1-F2-expressing plasmid and Renilla luciferase reporter into 293T cells for 24 h. Renilla luciferase activities were used as an internal reference. The assay was performed in triplicate. The values are expressed as mean ± SD. l The binding of PB1-F2 to the TATA-box in PNP promoters was examined by ChIP-qPCR. 293T cells were transfected by PB1-F2-Flag overexpression plasmid and luciferase reporter gene for PNP promoter, followed by anti-Flag antibody pulldown (IgG was used as negative control) and subsequent ChIP-qPCR analysis. ChIP-qPCR was conducted using primers flanking TATA-box in PNP promoters. The occupancy of PB1-F2 on the binding site was calculated as percentage of respective input DNA concentration. m Conserved phenylalanine residues among different strains of IAV. Phe-83 was identified to be conserved within PB1-F2 protein across H1N1, H3N2, H5N1, and H7N7 viruses. n The structure of PB1-F2 of different strains of viruses was predicted using AlphaFold 3. The phenylalanine residue was highlighted in red. o F83Y-mutated PB1-F2 overexpression plasmids of different strains of viruses failed to upregulate PNP expression. A549 cells were transiently transfected with vectors overexpressing either wild-type or mutated PB1-F2 proteins of various strains of IAV for 24 h, followed by examination of the mRNA and protein level of PNP via qRT-PCR and Western Blot analysis, respectively. All data are presented as mean ± SD; unless otherwise indicated, N = 3 biologically independent experiments; Statistical analysis was performed by one-way ANOVA (b–e, g, k, o), two-way ANOVA (l)
Furthermore, the potential of PB1-F2 as a novel viral transcription factor for PNP was analyzed by dual luciferase gene reporter assay. The luciferase reporter containing the full-length of PNP promoter (pGL3-PNP-promoter) was constructed and then co-transfected with Renilla luciferase reporter and the control or PB1-F2 overexpressing vector into 293 T cells (Fig. 3f). The pGL3-PNP-promoter was significantly activated by PB1-F2 overexpressing vector (Fig. 3g). To verify the exact binding site of PB1-F2 on PNP promoter, gene fragments of 50 bp in length, taken at 25 bp intervals upstream of the transcription start site (TSS) of PNP, were analyzed by AlphaFold 333 model for predicting the structure of the PB1-F2-DNA complex and DeepPBS, a geometric deep-learning model designed to predict binding specificity from protein-DNA structure,34 was used to identify the predicted binding motif between PB1-F2 and PNP promoter (Fig. 3h). Our data indicated that PB1-F2 closely interacted with the 50-bp DNA fragment nearest to the TSS site (−50 ~ + 1) of PNP promoter (Fig. 3i). A TATA-box (located at 20 ~ 30 bp upstream TSS) within the predicted motif was most likely to be the binding core of PB1-F2-DNA complex (Fig. 3j).
To prove this, we constructed luciferase reporter plasmids containing the predicted motif (50 bp) of PNP promoter with or without the mutated TATA-box and found that PB1-F2 overexpression was unable to activate the reporter gene with a mutated TATA box (Fig. 3k). By ChIP-qPCR, the enriched signal of PB1-F2 was also found to be specific within the predicted binding motif of PNP (Fig. 3l). Previous studies have indicated that viruses may possess a universally conserved residue (Phe) within their TATA-box-binding protein-like domain.35 We found that Phe-83 of PB1-F2 was conserved in H1N1, H3N2, H5N1, and H7N7, and was likely to be involved in DNA binding Fig. 3m). Furthermore, the structure of PB1-F2 in each viral strain was predicted using AlphaFold 3, revealing that Phe-83 was situated in the alpha helix of PB1-F2 (Fig. 3n). To test the functional significance of this residue, we constructed PB1-F2 overexpression plasmids harboring the F83Y mutation. Our results demonstrated that this mutation significantly diminished the upregulation of PNP, supporting that Phe-83 within PB1-F2 across different viral strains was critical for its interaction with PNP (Fig. 3o).
To our knowledge, our data is the first to reveal that H1N1 protein PB1-F2 serves as a viral transcription factor to transcriptionally activate the expression of host PNP, contributing to the viral-host interaction to promote disease progression.
PNP knockdown suppresses viral replication via compromised purine salvage
As PNP is a critical enzyme of purine salvage, we next investigated the alterations of host purine nucleotide metabolism (PNM) during H1N1 infection. Accordingly, by analyzing the scRNA-seq data of the lungs of IAV-infected mice, expression of multiple purine salvage enzymes, especially PNP, were identified to be upregulated in alveolar type II (AT2) cells during H1N1 infection, while phosphoribosylaminoimidazole succinocarboxamide synthetase (Paics), a de novo purine biosynthetic enzyme, was slightly downregulated (Fig. 4a). In line with AT2 cells, AT1 cells had PNP expression dramatically upregulated upon H1N1 (Supplementary Fig. 2a). These results strongly suggested that H1N1 enhanced the activity of host purine salvage in alveolar epithelial cells, in which PNP played a critical role.
Compromised PNP activity suppresses H1N1-hijacked host purine nucleotide salvage. a (ScRNA-seq analysis of lungs derived from control and H1N1 mice models. The expression of enzymes involved in both purine de novo and salvage biosynthesis in alveolar type 2 (AT2) cells. b Schematic of PNP-catalized purine salvage pathway. c–e LC-MS-based targeted metabolomics analysis of AEC-specific PNP conditional knockout mice after challenged for 6 days. Heat map showing the pulmonary metabolites landscape of nucleotides biosynthesis (c), dedes synthesis (d), and salvage pathway (e) based on metabolomic data. The experiments are performed in 3 replicates. Metabolites with P < 0.05 were defined as biologically significant. N = 3. f–h LC-MS-based targeted metabolomics analysis of A549 cells with or without PNP knockdown after challenged for 24 h. A549 cells, transfected by 30 pmol NC or PNP siRNA for 12 h, were challenged with H1N1 at an MOI of 5 for another 24 h, followed by metabolomic analysis. Heat map showing the intracellular metabolites landscape of ribonucleotides biosynthesis (f), de novo ribonucleotides synthesis (g), and salvage pathway (h) based on metabolomic data. The experiments are performed in three replicates. Metabolites with P < 0.05 were defined as biologically significant. i PNP convert purine nucleosides to purines upon H1N1 infection. A549 cells were treated with [U-15N] guanosine, [U-15N] adenosine and [U-15N] inosine for 4 h, respectively. Intracellular 15N-Guanine, 15N-adenine and 15N-hypoxanthine were measured with LC-MS. All data are presented as mean ± SD; unless otherwise indicated, N = 3 biologically independent experiments; Statistical analysis was performed by two-tailed Student’s t-test (a) and one-way ANOVA (c–i). *P < 0.01 versus Mock (a)
We further carried out LC-MS-based targeted metabolomic analysis of PNP-CKO mice and A549 cells, in order to quantify the change of metabolites involved in nucleotide metabolism. We found that IAV infection robustly induced nucleotide metabolism rewiring, and PNP inhibition potently reversed the metabolic disorders (Fig. 4c-h). Particularly, though both purine and pyrimidine ribonucleotide biosynthesis pathways were dramatically affected after H1N1 infection, PNP suppression tended to rescue ATP over-consumption in purine nucleotide metabolism rather than pyrimidine ribonucleotide biosynthesis (Fig. 4c, e). Of note, inosine monophosphate (IMP) was not remarkably changed after H1N1 infection (Fig. 4c, f), suggesting a balance between the consumption and production of IMP in purine synthesis. This balance might be attributed to a host ‘shut-off’ activity of H1N1 to salvage nucleotide supply for viral replication at the price of the reduced synthesis of cellular mRNA, which has been reported in infection of severe acute respiratory syndrome (SARS) coronaviruses.36,37
One of the most striking changes in H1N1-infected A549 metabolomics was the reduction of purine salvage nucleosides (guanosine; inosine; adenosine) and the accumulation of de novo purine synthesis substrates (glutamine) (Fig. 4d, g). As salvage pathway can restrain the use of ATP needed for de novo synthesis and ensure efficient usage of recycled nucleobases and nucleosides,21 our results indicated that H1N1 infection re-routes host purine metabolism to achieve rapid replication via salvage. Conversely, PNP knockdown by siRNA (siPNP) resulted in an increased level of guanosine, inosine, and adenosine and a reduction of glutamine upon infection (Fig. 4e, h). Moreover, an upregulation of a de novo purine synthesis intermediate 1-(5’-Phosphoribosyl)-5-amino-4-imidazolecarboxamide (AICAR) was observed in infected CKO mices and infected siPNP cells (Fig. 4d, g). Thus, our data indicated that purine salvage pathway in AECs was dramatically activated by H1N1 infection, which could be switched to de novo synthesis via PNP deficiency.
Of note, in eukaryotic cells, adenosine was not considered as the principal substrate of PNP (Fig. 4b).38 LACC1 (laccase domain containing (1) has been reported to combine activity analogous to PNP in catalyzing the phosphorolysis of adenosine, guanosine, and inosine.39 However, LACC1 was almost non-detectable in AT2 cells (Fig. 4a). As shown in Fig. 4e, h, PNP knockdown restored H1N1-induced adenosine downregulation, and thus we speculated that the massive nucleosides demand of H1N1 replication directly promoted the conversion of adenosine by PNP. We found that, similar to the change of metabolomic analysis, H1N1 markedly increased the accumulation of 15N-guanine, 15N-adenine, 15N-hypoxanthine in A549 cells, which can be reversed by PNP knockdown (Fig. 4i).
Collectively, our results revealed that IAV infection controlled the host purine metabolism through PNP, and PNP deficiency significantly inhibited viral replication by reducing the activity of H1N1-enhanced purine salvage.
PNP inhibition re-routes metabolic flux from purine salvage to de novo synthesis during IAV infection
To illuminate the effect of PNP on H1N1 infection-altered host nucleotide metabolism more straightforwardly, metabolic flux analysis was performed to trace the source of the purine ring in the production of ATP and GTP in A549 cells and BEAS-2B cells. H1N1-infected A549 and BEAS-2B cells were incubated with stable isotope-labeled 15N-amide-glutamine, U-15N inosine, U-15N adenosine, and U-15N guanosine, all of which contributed to the purine ring synthesis. The peak area of 15N-glutamine, the principal substrate of purine de novo synthesis, was much higher in infected cells when compared to the mock, revealing a decreased utilization of 15N-glutamine (Fig. 5a, b and Supplementary Fig. 8), while PNP inhibition enhanced the utilization of 15N-glutamine in challenged A549 and BEAS-2B cells (Fig. 5b and Supplementary Fig. 8). Accordingly, the peak area of 15N-labeled purine intermediates (IMP, AMP, ADP, ATP, GMP, GDP, GTP) during de novo synthesis were decreased in response to H1N1 infection, yet significantly increased by PNP knockdown (Fig. 5b and Supplementary Fig. 8). These results indicated that H1N1 inhibited de novo purine synthesis in alveolar epithelial cells, which could be reversed by PNP suppression.
PNP knockdown re-routes metabolic flux from purine salvage to de novo synthesis upon IAV infection. a, c, e, g Schematic of metabolic flux assay for purine synthesis. The de novo purine synthesis was tracked by isotype-labeled [Amide-15N] glutamine (a), inosine salvage by [U-15N] inosine (c), adenosine salvage by [U-15N] adenosine (e), and guanosine salvage by [U-15N] guanosine (g). b, d, f, h Metabolic flux analysis of metabolites involved in de novo purine synthesis and purine salvage pathways. The control and H1N1 infected A549 cells were cultured in media containing 15N-labeled starting material for 4 h before mass spectrometry analysis of isotype-labeled glutamine (b), inosine (d), adenosine (f), and guanosine (h), as well as related purine nucleosides. The experiment was conducted in triplicate. M + (n): the gross (n) of incorporated 15N. All data are presented as mean ± SD; unless otherwise indicated, N = 3 biologically independent experiments; Statistical analysis was performed by one-way ANOVA (b, d), f (15N-Adenosine) and h (15N-Guanosine); Two-way ANOVA in f (15N-ATP, 15N-ADP and 15N-AMP) and h (15N-GTP, 15N-GDP and 15N-GMP)
We next explored the effect of H1N1 infection on purine salvage. In contrast to its restriction on de novo pathway, H1N1 infection increased the consumption of U-15N inosine and the generation of 15N-labeled purine intermediates (IMP, ATP) in both A549 and BEAS-2B cells (Fig. 5c, d and Supplementary Fig. 8). In addition, compared to the control group, a lower level of 15N-labeled ATP was detected in PNP-knockdown cells upon infection (Fig. 4d, Supplementary Fig. 8). As for adenosine salvage, H1N1 enhanced the utilization of 15N-labeled adenosine and 15N-labeled AXP (ATP, ADP, AMP) production (Fig. 5e, f and Supplementary Fig. 8). Even though the phosphorolytical conversion of adenosine to adenine was not catalyzed by PNP in eukaryotic cells, PNP knockdown still reduced 15N-labeled adenosine consumption and AXP production (AMP, ADP, ATP, Fig. 4e, f and Supplementary Fig. 8), which was consistent to the increased level of adenosine in PNP-knockdown A549 and BEAS-2B cells (Fig. 4e, Supplementary Fig. 8). During guanosine salvage, suppressing PNP activity reduced the generation of 15N-labeled purine products increased by H1N1 (Fig. 5g, h, and Supplementary Fig. 8). Moreover, the branch of adenosine and guanosine salvage (M + 4) pathway was not significantly affected by H1N1 or PNP (Fig. 5f, h and Supplementary Fig. 8).
Collectively, by metabolite flux analysis, we further demonstrated that, upon infection, PNP inhibition blocked H1N1-enhanced purine salvage while reactivated purine de novo synthesis in alveolar epithelial cells.
PNP-catalyzed purine nucleosides recycling benefits H1N1 propagation
Given that purine nucleotides rewiring in response to H1N1 infection was mediated by host PNP, we employed FISH assays to further verify the PNP-mediated purine salvage in vitro and in vivo. The addition of methotrexate (MTX), a dihydrofolate reductase (DHFR) competitive inhibitor used as an inhibitor for one-carbon metabolism involved in nucleotide synthesis, inhibited vRNA production and diminished viral titer in both A549 and BEAS-2B cells (Fig. 6a, c). This was accompanied by an increase in cell viability (Fig. 6c). Importantly, the inhibition of vRNA and virus production can be reversed by the addition of purine salvage substrate adenosine, guanosine, or inosine (Fig. 6a, c). These results implied that IAV efficiently utilized purine salvage to support viral propagation. However, when PNP was knocked down in infected cells, the addition of extra guanosine or inosine was unable to restore vRNA generation, viral titer, or cell viability, highlighting that the effect of purine salvage on viral replication was relied on PNP (Fig. 6b, d). We further verified the PNP-mediated purine salvage in vivo (Fig. 6e). The administration of the wild-type mice with purine nucleosides after H1N1 infection dramatically aggravated lung damage and intensity of vRNA (Fig. 6f). While PNP-knockout mice significantly suppressed the rapid growth of H1N1 virus and severe lung damage induced by nucleosides supplementation (Fig. 6g).
PNP-mediated purine salvage is essential for H1N1 propagation. a, c The effect of nucleosides on viral replication and cell viability upon MTX treatment in challenged cells. H1N1-challenged A549 and BEAS-2B cells were cultured in medium with 2 μM MTX for 12 h, and then treated by adenosine (50 μM), guanosine (50 μM), or inosine (50 μM) for another 24 h, followed by FISH analysis of vRNA (Scale bar: 100 μm, a). Cell viability and viral titer were also examined (c). b, d The effect of nucleosides on viral replication and cell viability was dependent on PNP activity. H1N1-infected A549 and BEAS-2B cells, transfected by NC siRNA or PNP siRNA, were cultured in the addition of 50 μM guanosine, inosine, or adenosine for 24 h. FISH analysis of vRNA, IF assay of PNP, and DAPI staining were performed (Scale bar: 100 μm, (b). Cell viability and viral titer were also analyzed (d). All the experiments were conducted in triplicates. e Diagram of the experimental procedures. Briefly, 8-week-old PNPflox/flox mice and PNP conditional knockout mice were intranasally challenged with 2LD50 H1N1. H1N1-infected mice were intragastric administered with adenosine (8 mg/Kg), guanosine (8 mg/Kg), and inosine (8 mg/Kg), respectively. f, g The inflammatory lesion and viral infection within the lungs of H1N1-infected mice. After 6 days of infection, 3 mice from each group were sacrificed, with the lungs dissociated and subjected to H&E staining (f, Scale bar: 50 μm), FISH of vRNA, and IF assay of PNP (g). DAPI was used to indicate nucleus (N = 6, Scale bar: 10 μm). The area of lung damage and intensity of vRNA (f, g, the right plots) were shown. All data are presented as mean ± SD; unless otherwise indicated, N = 3 biologically independent experiments; Statistical analysis was performed by one-way ANOVA (c, d, f, g)
Taken all the above results together, we have revealed the critical role of host purine salvage in H1N1 replication and host cell viability, demonstrating that the sensitivity of viral infection to purine nucleosides was dependent on PNP activity.
PNP inhibition exhibits anti-inflammation roles via APRT-AICAR-AMPK axis upon H1N1 infection
Consistent with the effect of H1N1 infection on purine synthesis and the critical role of PNP, a series of purine salvage enzymes were found to be decreased in PNP-knockdown A549 cells, compared to the control group in response to H1N1 infection. However, the de novo purine synthesis enzymes, such as polyfluoroalkyl substances (PFAS), 5-Amino-4-imidazolecarboxamide ribonucleotide transformylase/IMP cyclohydrolase (ATIC), were increased by transcriptomic data (Supplementary Fig. 3a–c). Furthermore, gene set enrichment analysis (GSEA) demonstrated that AMP-activated protein kinase (AMPK) pathway, whose anti-inflammation effect has been extensively studied40,41 was activated, while a series of downstream proinflammation signaling (phosphoinositide 3-kinase (PI3K)-Akt, TNF, Janus kinase/signal transducer and activator of transcription (JAK/STAT) and nuclear factor kappa B (NF-κB) pathway) were inhibited (Supplementary Figs. 3d, 4a). To better illuminate the regulatory role of PNP-AMPK axis in influenza-associated inflammation, we further inhibited AMPK phosphorylation by compound C (a well-defined AMPK antagonist) and found that compound C significantly reduced the anti-inflammation response of PNP inhibition, which was evidenced by a much higher level of IL-6 and TNF-α in PNP-knockdown A549 and BEAS-2B cells (Supplementary Figs. 3e, 4b).
Next, we set to uncover the molecular mechanism of PNP-modulated AMPK activation. As discussed in Fig. 4, AICAR, a de novo purine synthesis intermediate, was increased by PNP inhibition. Actually, AICAR was well-known for its activating effect on AMPK.42 Moreover, hypoxanthine phosphoribosyltransferase 1 (HPRT1) and adenine phosphoribosyltransferase (APRT), playing central roles in purine salvage and AICAR production,42,43 were also found to be upregulated in PNP-knockdown cells (Supplementary Fig. 3c). By knockdown of APRT or HPRT1, we found that APRT, rather than HPRT1, was responsible for PNP-deficiency-induced accumulation of AICAR upon H1N1 infection (Supplementary Fig. 3f). Furthermore, the suppression of APRT inhibited AMPK activation and reversed the expression of proinflammatory cytokines (Supplementary Fig. 3g, 4c). These results indicated that PNP suppression-induced AMPK activation under H1N1 infection was mediated by APRT-catalyzed AICAR accumulation.
Thus, our results unveiled that PNP inhibition activated APRT-AICAR-AMPK signaling to suppress hyperinflammation induced by H1N1 infection, suggesting the hub role of PNP in connecting viral replication and inflammation via purine salvage.
Purine nucleotides in the blood of IAV-infected patients is correlated with infection and peripheral inflammation
Given the above discussed role of host purine salvage in IAV infection, we further explored the variation of purine metabolism in IAV-infected patients. The peripheral plasma derived from 19 healthy controls and 41 H1N1-infected individuals were subjected to widely-targeted metabolomic analysis. 326 metabolites in positive ion mode and 331 in negative were identified (Fig. 7a). Via orthogonal partial least squares discriminant analysis (OPLS-DA), healthy control (HC) group and IAV-infected group could be distinguished clearly (Fig. 7b and Supplementary Fig. 5a). 146 out of 657 metabolites were significantly associated with IAV infection (P < 0.05; Fig. 7c). Consistent with our in vitro/in vivo data, nucleotide metabolism was significantly enriched in H1N1 patients by Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis (Fig. 7d). In particular, all the mononucleotides were significantly increased, implying that H1N1 infection reprogramed host nucleotide metabolism to meet the massive demand for viral replication (Supplementary Fig. 5b). Similar to the alteration pattern revealed in IAV-infected AECs, in H1N1-infected patients, glutamine of de novo synthesis was also accumulated, while metabolites of purine salvage including all the purine bases (adenine, hypoxanthine, guanine) and inosine were decreased (Fig. 7e).
Purine nucleotides in the blood of IAV-infected patients is correlated with infection and peripheral inflammation. a The schematic diagram of widely-targeted metabolomic profiling of plasma from H1N1 patients (H1N1 positive) and healthy control (HC). Briefly, the plasma of the peripheral blood from 19 HC and 41 H1N1 patients were subjected to widely-targeted metabolomic analysis. b Orthogonal partial least square discriminant analysis (OPLS-DA) for the data from widely targeted metabolomic profiling of HC and H1N1 positive individuals. The HC and H1N1 group were discriminated into two separated clusters by OPLS-DA analysis. c Heatmap of identified metabolites. The expression of 657 identified metabolites from the widely-targeted metabolomic data were represented by heatmap. d KEGG pathway enrichment analysis of the differentiated expressed metabolites between HC and infected individuals. The top 20 KEGG pathways were enriched, with the annotation of nucleotide metabolism pathway being labeled as red. e An overview of purine metabolism pathway alterations and metabolites involved in de novo purine synthesis and purine salvage. Red arrows indicated upregulated metabolites, while blue arrows indicated downregulated ones. f AUC-ROC curve analysis of the correlation between purine nucleotide and H1N1 infection. The discriminative capability of purine nucleotide GMP, IMP, and AMP in distinguishing H1N1-infected and uninfected individuals was quantified by area under ROC curve (AUC), with GMP showing the highest AUC. g The expression of TNF-ɑ and IL-6 in the peripheral plasma. The concentration of TNF-ɑ and IL-6 of the control and H1N1-infected patients were measured by ELISA. h The correlation analysis between inflammatory cytokines and purine nucleotides. The correlation of purine nucleotides (GMP, IMP, AMP) with inflammatory cytokines (TNF-ɑ, IL-6) in the peripheral blood of H1N1 patients. All data are presented as mean ± SD; Statistical analysis was performed by two-tailed Student’s t-test (g) and Spearman rank correlation coefficient (h)
Next, we generated ROC curves to assess if the nucleotide metabolites signatures could be used as diagnostic biomarkers for H1N1. The ROC curves revealed that GMP, with the AUC value of 0.990, was the most efficient metabolite in discriminating H1N1 positive patients from uninfected controls (Fig. 7f). The concentration of IL-6 and TNF-ɑ in the peripheral blood were also dramatically increased in patients with H1N1 (Fig. 7g). Moreover, the levels of GMP, IMP and AMP were positively correlated with the peripheral expression of IL-6 and TNF-ɑ (Fig. 7h). To further investigate the relationship between the biomarkers and disease outcomes, we stratified the enrolled patients into mild and severe groups according to their clinical characteristics(Supplementary Table 5).44 Subsequently, we re-evaluated the correlation between the biomarkers (GMP, AMP, and IMP) and clinical outcomes. The results indicated that GMP was positively correlated with the severity of the disease, suggesting that GMP may serve as a potential biomarker for disease progression (Supplementary Fig. 5c, d).
Overall, these observations validated the transition of purine metabolism from de novo to salvage pathway in H1N1-infected patients, and provided evidence for the potential biomarker role of nucleotide metabolite in disease diagnosis.
PNP inhibitors are promising host-targeted anti-influenza drugs
Multiple PNP inhibitors (PNPIs) have been developed, exhibiting potential therapeutic effects in clinical. Peldesine is a PNPI for treatment of autoimmune disorders and has entered clinical trials for human immunodeficiency virus (HIV) infections. Ulodesine is an investigational PNPI used to treat hyperuricemia and gout.45 8-aminoinosine is reported to exert diuretic and natriuretic activity by inhibiting purine nucleoside phosphorylase.46 All the three PNPI showed marked inhibitory effect on the viral copy number in the supernatant of H1N1-infected MDCK cells, indicating the anti-influenza potential of PNPIs (Fig. 8a).
VI-PPI-predicted dihydroartemisinin (DHA) plays an anti-influenza role in vitro/in vivo by targeting host PNP. a The anti-H1N1 activities of the verified PNPIs. MDCK cells infected with H1N1 at an MOI of 0.01 were treated with PNPIs (peldesine, ulodesine, ganciclovir) at different doses for 24 h, followed by analysis of the viral copy number in the supernatant of H1N1-infected cells. b The effect of DrugBank-predicted PNPIs on H1N1 replication. MDCK cells infected with H1N1 at an MOI of 0.01 were treated with cladribine or didanosine at different concentrations for 24 h, followed by analysis of the viral copy number in the supernatant of H1N1-infected cells. c The anti-H1N1 activity of VI-PPI-predicted PNPI. H1N1-infected MDCK cells were treated with different doses of DHA, which was predicted by VI-PPI network, and then the viral copy number in the supernatant was analyzed. d Molecular operating environment (MOE) analysis of the interaction between PNP and DHA. The potential binding sites in PNP are highlighted in red and DHA is indicated in green. e The kinetic profile of DHA-PNP binding reaction by surface plasmon resonance (SPR) assay. PNP was immobilized to the surface of the sensor chip. The binding and dissociation of DHA at different concentrations with PNP chip were monitored. The predicted docking score and affinity parameters of DHA binding to PNP are displayed in the lower chart. (f–i) The in vivo effect of DHA on H1N1 infection. f The schematic of the in vivo assay of the effect of DHA on H1N1-infected mice models. Briefly, mice were randomized into 3 groups (mock, H1N1 plus vehicle (90% corn oil and 10% DMSO), H1N1 plus DHA (12.5 mg/Kg, 25 mg/Kg, 50 mg/Kg)). Vehicle or DHA were intraperitoneally administered for 6 consecutive days. g The survival rate was monitored for 16 consecutive days after infection (N = 10). h The lung tissue of the mice was dissected for H&E analysis 6 days after vehicle or DHA treatment (N = 6, Scale bar: 50 μm). i Expression level of nuclear protein (NP) of H1N1 in mice lung (N = 6) measured by qRT-PCR on day 6 post infection. The concentration of TNF-ɑ and IL-6 levels in mice plasma (N = 6) was measured by ELISA on day 6 post infection. All data are presented as mean ± SD; unless otherwise indicated, N = 3 biologically independent experiments; Statistical analysis was performed by Log-rank test (g) and one-way ANOVA (i)
However, there is no FDA-approved PNPIs available to date. As discussed before, that VI-PPI network incorporated drug targets data, we tried to screen potential PNP-targeting drugs (FDA-approved) via this newly created model. The top 200 PNP-related genes were extracted using Random Walk with Restart algorithm (RWR), followed by an enrichment analysis based on the approved drug targets derived from DrugBank database. As a result, dihydroartemisinin (DHA, also known as artenimol), ranked the first place with the lowest p value and the most overlapped genes between PNP-related molecules and drug targets, was capable of inhibiting H1N1 replication (Fig. 8c and Supplementary Fig. 6a). Moreover, DHA increased the viability of infected A549 and BEAS-2B cells in a dose-dependent manner, peaking at the concentration of 1 µM. And thus, 1 µM DHA was used afterwards. The viral load in AECs was also markedly inhibited by DHA at 1 µM (Supplementary Fig. 6b). Additionally, cladribine (deoxyadenosine analog) and didanosine (deoxynucleoside analog), suggested by the DrugBank database to be capable of interacting with PNP, were also evaluated for their effect on H1N1-infected cells. However, no significant protective roles were found in cells treated with these two drugs (Fig. 8b), indicating that VI-PPI-predicted DHA might be a novel PNP-targeting compound in IAV treatment.
To demonstrate if there was a direct inhibition of DHA on PNP, we evaluated the protein level of PNP upon DHA treatment, and found that IAV-induced upregulation of PNP in A549 and BEAS-2B cells were reduced by DHA (Supplementary Fig. 6c). Additionally, overexpression of PNP significantly reversed DHA-induced increase of cell viability and reduction of H1N1 viral titer, indicating that the anti-viral effect of DHA was mainly mediated by its effect on PNP (Supplementary Fig. 6d). Then, molecular docking analysis predicted a docking score of −5.99 to imply a promising binding between DHA and PNP. Also, a potential pocket consisting of 14 amino acids within PNP were identified, and two H-bonds were implied to be responsible for the binding (Fig. 8d and Supplementary Fig. 6e). Importantly, surface plasmon resonance (SPR) assay confirmed the direct interaction between DHA and PNP, showing a dissociation equilibrium constant (KD) of \(3.282\times 10-8\,\)M (Fig. 8e).
Subsequently, the anti-influenza effect of DHA was examined in vivo. H1N1-challenged mice were intraperitoneally administered with DHA or vehicle (90% corn oil and 10% DMSO) daily for a consecutive six days (Fig. 8f). DHA induced a robust extended survival rate and an accelerated body weight restoration upon IAV when compared to vehicle-treated group (Fig. 8g and Supplementary Fig. 6f), with PNP downregulation and AMPK activation observed in the lungs (Supplementary Fig. 6g). DHA administration (50 mg/Kg/day) also significantly mitigated the pulmonary pathological changes of the infected mice (Fig. 8h), reducing leukocytes infiltration, proinflammatory cytokines production (IL-6, TNF-ɑ), and viral nucleoprotein (NP) expression (Fig. 8i).
Collectively, our results clearly demonstrated the potential of PNP as an anti-influenza host target, with PNP inhibitors being efficient to combat IAV. More importantly, DHA, predicted by VI-PPI as a novel PNP-interacting molecule, could directly bind PNP to constrain viral replication and host hyperinflammation.