Palmitoylation inhibits immune evasion by enhancing host IFN-I signaling against malaria
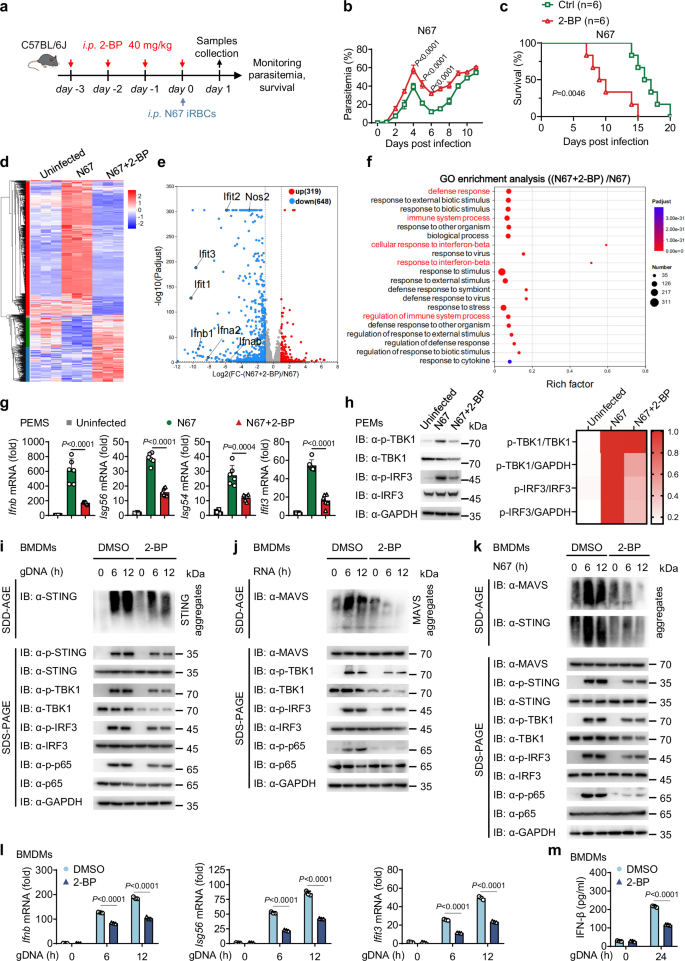
To assess the role of palmitoylation in malaria infection, we employed 2-bromopalmitate (2-BP), an inhibitor of palmitoylation that specifically targets DHHC (Asp-His-His-Cys) protein palmitoyltransferases. C57BL/6 J female mice received intraperitoneal (i.p.) injection of dimethyl sulfoxide (DMSO) or 2-BP (40 mg/kg) for four days30, followed by an injection of Plasmodium. yoelii N67 (N67) infected red blood cells (iRBCs) (1 × 106, i.p.) (Fig. 1a). Mice treated with 2-BP exhibited significantly higher parasitemia and lower survival rate compared to the control group (Fig. 1b, c). RNA-sequencing analysis of peritoneal macrophages (PEMs) revealed that 2-BP treatment down-regulated hundreds of genes, including key interferon-stimulated genes (ISGs) involved in innate immune responses against malarial infection (Fig. 1d, e). Gene Ontology (GO) analysis further indicated an enrichment of genes participating in various immune signaling pathways, notably those related to “defense response” and “response to interferon-beta” after 2-BP treatment (Fig. 1f). In addition, Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR) and immunoblotting demonstrated a marked inhibition of the IFN-I signaling pathway in PEMs from 2-BP-treated N67-infected mice when compared to the N67 infection group (Fig. 1g, h).
a–c C57BL/6 J female mice (n = 6) were injected intraperitoneally (i.p.) with dimethyl sulfoxide (DMSO) or 2-bromopalmitate (2-BP) (40 mg/kg) for 4 days, followed by an injection of Plasmodium. yoelii N67 (N67) infected red blood cells (iRBCs) (1 × 106, i.p.). Parasitemia (b) and survival rate (c) were then evaluated. This was repeated n = 3 independent times with similar results. Figure 1a was adapted from the image “Adult Mouse” by Yunan Chen (SciDraw; CC BY 4.0). d, e Heatmap (d) and volcano plot (e) display the differentially expressed genes in peritoneal macrophages (PEMs) pre-treated with DMSO or 2-BP (100 mM for 24 h), with or without stimulation following freeze-thawed iRBCs. Each group has three biological replicates. (f) Gene ontology (GO) enrichment analysis of the differentially expressed genes shown in panel (e). g, h C57BL/6 J female mice were injected i.p. with DMSO or 2-BP (40 mg/kg) for 4 days, followed by an injection of N67 iRBCs (1 × 106, i.p.) for 24 h. RT-qPCR analysis was performed to measure the mRNA expression levels of Ifnb, Isg56, Isg54 and Ifit3, data shown are from one representative experiment (n = 6 mice per group), consistent results were observed across three biological replicates (g). Immunoblotting was conducted to assess the levels of total IRF3 and TBK1, as well as phosphorylated IRF3 and TBK1, on day 1 after N67 infection in PEMs (h). Representative heatmaps display the results of three independent replicate experiments. The grayscale values from each replicate were quantified, and the average value across the three replicates was calculated. This average was then normalized using the group with the highest grayscale value as the baseline (set to 1), to generate the final heatmap. i–m Bone-marrow derived macrophages (BMDMs) were pre-treated with DMSO or 2-BP (100 mM for 24 h) and then stimulated with gDNA, RNA or freeze-thawed iRBCs at indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (i–k) and RT-qPCR analysis (n = 3 biological replicates) (l). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (m). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (b, g, l, m), and two-sided log-rank test (c). In (e) differential expression analysis between N67 + 2-BP and N67 was performed using DESeq2 (Version 1.42.0). The Wald test (two-tailed) was used, and the resulting p-values were adjusted for multiple testing using the Benjamini-Hochberg method. Genes with an adjusted |log2FC | ≥ 1 and padjust < 0.05 were considered significantly differentially expressed. In (f) Gene Ontology (GO) biological process enrichment analysis was performed on the list of genes that were significantly down-regulated in the N67 + 2-BP group compared to the N67 group (|log2FC | ≥ 1 and padjust < 0.05). The analysis was carried out using the goatools (Version 1.4.4) with Fisher’s exact test (two-tailed). All genes detected in the RNA-seq experiment were used as the background universe. The resulting p-values were adjusted for multiple testing using the Benjamini-Hochberg method. Source data are provided as a Source Data file.
To further validate the role of 2-BP, we stimulated RAW 264.7 cells and bone-marrow-derived macrophages (BMDMs) with N67 genomic DNA (gDNA), RNA or crude antigens from N67 iRBCs subjected to repeated freezing-thaw cycles. 2-BP treatment resulted in a significant inhibition of the IFN-I signaling pathway in both cell types, which is characterized by reduced oligomerization of MAVS and STING, along with decreased phosphorylation of STING, TBK1 and IRF3 (Fig. 1i–k and Supplementary Fig. 1a–c). In addition, reduced TBK1-mediated phosphorylation of p65 in the downstream NF-κB pathway was observed (Fig. 1i–k), alongside lowered expression of Ifnb and ISGs (Fig. 1l and Supplementary Fig. 1d–h) and diminished production of IFN-β (Fig. 1m and Supplementary Fig. 1i–m), indicating that 2-BP inhibits the IFN-I response against N67 infection. Therefore, these findings suggest that palmitoylation enhances the IFN-I signaling and anti-malarial immune response.
ZDHHC9 mediates TBK1 palmitoylation during malarial infection
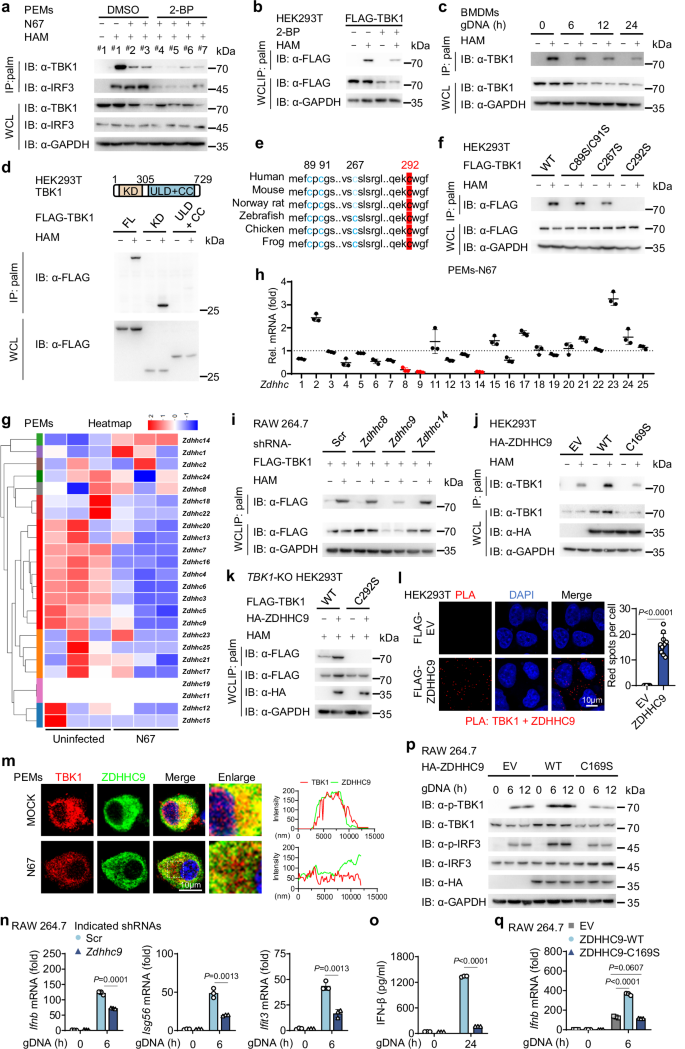
In order to identify the molecular mechanism through which 2-BP inhibits the IFN-I pathway, we conducted an acyl-biotin exchange (ABE) assay to investigate the palmitoylation status of key molecules involved in the IFN-I signaling pathway undergo palmitoylation. The persistence of the signal after hydroxylamine (HAM) treatment indicated that these proteins are palmitoylated via thioester bond formation31. Our results revealed that both TBK1 and IRF3 are susceptible to palmitoylation, while only TBK1 exhibited a decrease in palmitoylation levels following N67 infection (Fig. 2a). Treatment with 2-BP during N67 infection resulted in nearly undetectable palmitoylation levels of TBK1 (Fig. 2a). Exogenous experiments reaffirmed TBK1’s susceptibility to palmitoylation, with treatment with 2-BP substantially reducing its palmitoylation levels (Fig. 2b). Furthermore, stimulation of BMDMs with gDNA, RNA or freeze-thawed iRBCs revealed a time-dependent reduction in TBK1 palmitoylation (Fig. 2c and Supplementary Fig. 2a, b). These results indicate that TBK1 undergoes palmitoylation, which declines during N67 infection.
a Cell lysates from experiments in Fig. 1 (h) were subjected to acyl-biotin exchange (ABE) assays and immunoblot analysis to assess TBK1 palmitoylation. This was repeated n = 3 independent times with similar results. b Human embryonic kidney (HEK) 293 T cells were transfected with Flag-TBK1 for 24 h, followed by the treatment of DMSO (vehicle) or 2-BP (100 mM for 12 h). TBK1 palmitoylation and protein levels were assessed by ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. c BMDMs were stimulated with gDNA at the indicated time points, and TBK1 palmitoylation levels were measured using ABE assay and immunoblotting. This was repeated n = 3 independent times with similar results. d HEK293T cells were transfected with wild-type (WT) or deletion mutants of Flag-TBK1 for 24 h, and TBK1 palmitoylation levels were measured using ABE assay and immunoblotting. This was repeated n = 3 independent times with similar results. e Comparative analysis of potential palmitoylation sites within TBK1 sequences across different species. f HEK293T cells were transfected with WT FLAG-TBK1 or TBK1 palmitoylation-deficient mutants for 24 h. TBK1 palmitoylation levels were detected by ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. g Heatmap analysis of expression changes in the ZDHHC gene family in PEMs, with or without stimulation following freeze-thawed iRBCs. Each group has three biological replicates. h The relative mRNA levels of Zdhhc genes in PEMs were determined by RT-qPCR following N67 infection, with mRNA levels was normalized to Gapdh (n = 3 biological replicates). i RAW 264.7 cells were transfected with Scr shRNA or Zdhhc shRNAs for 24 h, followed by transfection with FLAG-TBK1. TBK1 palmitoylation and protein levels were measured using the ABE assay and immunoblotting. This was repeated n = 3 independent times with similar results. j, k HEK293T cells were transfected with either WT HA-ZDHHC9 or the enzymatically inactive mutant ZDHHC9 (C169S) for 24 h (j). TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h (k). Cell lysates were harvested for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. l Proximity ligation assay (PLA) was performed on HEK293T cells overexpressing FLAG-ZDHHC9 to examine the colocalization of TBK1 and FLAG. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) are presented next to the image. This was repeated n = 3 independent times with similar results. m PEMs were stimulated with freeze-thawed iRBCs for 12 h. The colocalization between TBK1 (red) and ZDHHC9 (green) was examined by confocal microscopy. The nucleus was stained with DAPI (blue). Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. n, o RAW 264.7 cells were transfected with Scr shRNA or Zdhhc9 shRNA for 24 h, followed by stimulation with gDNA at the indicated time points. Cell lysates were used for RT-qPCR analysis (n = 3 biological replicates) (n). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (o). p, q RAW 264.7 cells were transfected with HA-EV, WT HA-ZDHHC9, or the enzymatically inactive mutant of ZDHHC9 (C169S) for 24 h, and then stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblotting. This was repeated n = 3 independent times with similar results (p) and RT-qPCR analysis (n = 3 biological replicates) (q). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (l, n, o, q). Source data are provided as a Source Data file.
TBK1 consists of three main domains: a kinase domain (KD), an ubiquitin-like domain (ULD), and a coiled-coil (CC) domain16. To pinpoint the region of TBK1 that undergoes palmitoylation, HEK293T cells were transiently transfected with TBK1 full-length (FL), TBK1-KD, and TBK1-ULD-CC constructs. Immunoblotting confirmed that both TBK1-FL and TBK1-KD were palmitoylated, while TBK1-ULD-CC was not (Fig. 2d). Notably, four conserved cysteine residues in the KD domain-C89, C91, C267, and C292-were identified across species (Fig. 2e). Only mutating C292 led to a significant reduction in TBK1 palmitoylation levels (Fig. 2f), suggesting that C292 is critical for TBK1’s palmitoylation.
Palmitoylation is catalyzed by DHHC-palmitoyltransferases32. The decrease in TBK1 palmitoylation following N67 infection prompted us to investigate the expression levels of palmitoyltransferases. Heat map analysis in RNA-seq indicated that multiple enzymes of the ZDHHC family in PEMs were downregulated after N67 infection (Fig. 2g). Further verification by RT-qPCR revealed significant down-regulation of Zdhhc8, Zdhhc9, and Zdhhc14 in PEMs after N67 infection (Fig. 2h). Using short hairpin-mediated RNA (shRNA) (Supplementary Fig. 2c) to knockdown Zdhhc9 but not Zdhhc8 or Zdhhc14 markedly decreased TBK1 palmitoylation (Fig. 2i). To further validate the relationship between members of the ZDHHC family and TBK1, exogenous co-immunoprecipitation (co-IP) experiments revealed that, in addition to ZDHHC9, several other members, pacifically ZDHHC13, ZDHHC16, ZDHHC21, and ZDHHC23, demonstrated significant interactions with TBK1. Building on these findings, we simultaneously assessed the impact of these interactions on the palmitoylation level of TBK1. Results indicated that only ZDHHC9 was capable of enhancing the palmitoylation level of TBK1 (Supplementary Fig. 2d, e), indicating that ZDHHC9 is a major palmitoyltransferase for TBK1 palmitoylation. To further explore the role of ZDHHC9 in TBK1 palmitoylation, we generated an enzymatically inactive mutant of ZDHHC9 (C169S)33 and confirmed that enzymatic activity of ZDHHC9 was crucial for TBK1 palmitoylation (Fig. 2j). Moreover, ZDHHC9 was unable to catalyze the palmitoylation of the TBK1 C292S mutant (Fig. 2k). Notably, both proximity ligation assays (PLA) and confocal microscopy consistently demonstrated a strong interaction between flag-ZDHHC9 and endogenous TBK1 (using flag-ZDHHC3 and endogenous TBK1 as controls) (Supplementary Fig. 2f). Furthermore, confocal microscopy demonstrated that stimulation with freeze-thawed iRBCs significantly decreased the co-localization of endogenous ZDHHC9 and endogenous TBK1 (Fig. 2l, m). Therefore, these results suggest that ZDHHC9 interacts with TBK1 and serves as the primary palmitoyltransferase for TBK1, maintaining its palmitoylation levels in the resting state. During N67 infection, the down-regulation of ZDHHC9 leads to a decrease in the interaction between ZDHHC9 and TBK1, which consequently results in a simultaneous reduction in the palmitoylation levels of TBK1.
Next, we investigated whether ZDHHC9 mediated TBK1 palmitoylation affects its function. After silencing ZDHHC9, we observed a reduction in the activity of TBK1-induced IFN-β and ISRE promoter (Supplementary Fig. 2g). Similar results were observed in RAW 264.7 cells when silencing ZDHHC9 expression with shRNA led to decrease activation of IFN-I signaling after stimulation with gDNA, RNA, or freeze-thaw iRBCs (Fig. 2n, o and Supplementary Fig. 2h–m). In contrast, the expression of wild-type (WT) ZDHHC9 markedly augmented the IFN-I signaling pathway, whereas expressing the enzymatically inactive mutant (C169S) ZDHHC9 did not produce this effect (Fig. 2p, q). Thus, these results suggest that ZDHHC9 plays a critical role in the palmitoylation of TBK1, thus activating the IFN-I signaling pathway.
APT2 mediates the depalmitoylation of TBK1 during Plasmodium infection
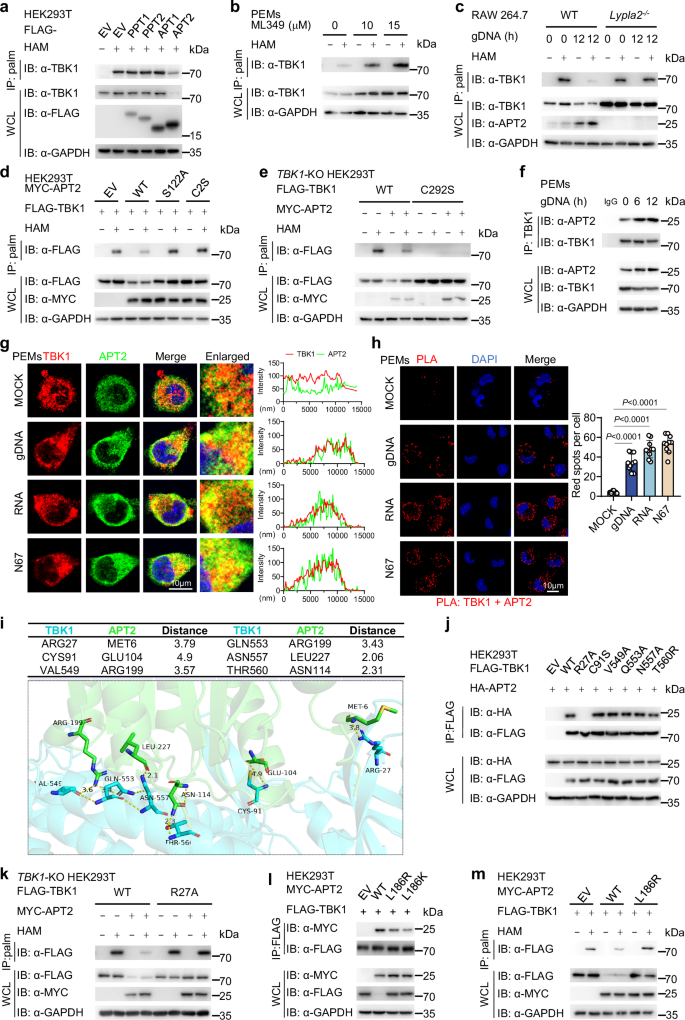
S-palmitoylation is a dynamic process regulated by two opposing classes of enzymes: palmitoyl acyltransferases (PATs) which add palmitate to proteins and acyl protein thioesterases, which remove it. Previous studies have identified several mammalian acyl-protein thioesterases, including APT1 and APT2, as well as palmitoylprotein thioesterases 1 and 2 (PPT1 and PPT2)34. To determine which enzyme is responsible for the depalmitoylation of TBK1 during Plasmodium infection, we expressed four known acyl protein thioesterases and measured the palmitoylation level of endogenous TBK1. Our results demonstrated that only the expression of APT2 led to a significant reduction in TBK1 palmitoylation, while the other APTs showed no such effect (Fig. 3a). Consistent with this finding, pharmacological inhibition of APT2 using ML349 and silencing of Lypla2 with small interfering RNA (siRNA) resulted in increased palmitoylation of TBK1 (Fig. 3b and Supplementary Fig. 3a, b).
a HEK293T cells were transfected with FLAG-EV, -PPT1, -PPT2, -APT1, or -APT2 for 24 h. TBK1 palmitoylation levels were detected by ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. b PEMs were treated with ML349 at the indicated concentrations for 12 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. c WT or Lypla2-knockout (KO) RAW 264.7 cells were stimulated with gDNA for 12 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. d HEK293T cells were transfected with FLAG-TBK1 and MYC-EV, MYC-WT APT2, or its catalytically inactivated variants (S122A and C2S) for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. e TBK1-KO HEK293T cells were reconstituted with plasmids encoding FLAG-WT TBK1 or FLAG-TBK1 C292S mutant and MYC-APT2 for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. f PEMs were stimulated with gDNA at the indicated time points. Cell lysates were collected for IP and immunoblot analysis. This was repeated n = 3 independent times with similar results. g PEMs were stimulated with gDNA, RNA, or repeated freeze-thaw cycles of N67 iRBCs for 12 h. The colocalization between TBK1 (red) and APT2 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. h PEMs were stimulated with gDNA, RNA, or repeated freeze-thaw cycles of N67 iRBCs for 12 h. The colocalization between TBK1 and APT2 was examined by PLA. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) are displayed next to the image. This was repeated n = 3 independent times with similar results. i Protein structure and molecular docking maps of TBK1 and APT2. j HEK293T cells were transfected with various combinations of plasmid encoding FLAG-WT TBK1, -R27A mutant, -C91S mutant, -V549A mutant, -Q553A mutant, -N557A mutant, or -T560R mutant of TBK1 and HA-APT2 for 24 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. k TBK1-KO HEK293T cells were reconstituted with plasmids encoding FLAG-WT TBK1 or FLAG-TBK1 R27A mutant and MYC-APT2 for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. l HEK293T cells were transfected with FLAG-TBK1 and MYC-EV, -WT APT2, -L186R APT2, or -L186K APT2 for 24 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. m HEK293T cells were transfected with FLAG-TBK1 and MYC-EV, -WT APT2, or -L186R APT2 for 24 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. Data are presented as the mean ± SD. P-values were determined by unpaired two-tailed Student’s t tests (h). Source data are provided as a Source Data file.
To further elucidate the in vivo function of APT2, we employed CRISPR/Cas9 technology to create Lypla2-knockout (KO) RAW 264.7 cells. Upon stimulation with gDNA, RNA, or freeze-thawed iRBCs, we observed a significant decrease in TBK1 palmitoylation levels in WT cells but not in Lypla2-KO cells (Fig. 3c and Supplementary Fig. 3c,d). In addition, catalytically inactive mutants of APT2 (S122A and C2S)35 were engineered and it was observed that the expression of WT APT2 significantly decreased TBK1 palmitoylation, while the inactive mutants did not (Fig. 3d). Furthermore, APT2 was ineffective in reducing the palmitoylation of the TBK1 C292S mutant (Fig. 3e). Together, these findings indicate that APT2 is responsible for the depalmitoylation of TBK1.
Furthermore, exogenous Co-IP experiments demonstrated that APT2 can interact with TBK1 when overexpressed (Supplementary Fig. 3e), and continued to bind to a TBK1 mutant with a phosphorylation site alteration (S172A), indicating that APT2 acts on the non-phosphorylated form of TBK1 (Supplementary Fig. 3e). In addition, endogenous Co-IP assay indicated that this physiological interaction is enhanced in PEMs under gDNA or RNA stimulation conditions (Fig. 3f and Supplementary Fig. 3f). Confocal microscopy and PLA analyses confirmed increased colocalization of APT2 with TBK1 in PEMs and BMDMs following stimulation with gDNA, RNA, or freeze-thawed iRBCs (Fig. 3g, h and Supplementary Fig. 3g). Additionally, molecular docking studies identified several potential binding sites on TBK1, including R27, C91, V549, Q553, N557, and T560, as likely candidates for its interaction with APT2 (Fig. 3i). To explore these interactions, we generate mutants of TBK1 and performed Co-IP assays, which revealed that the TBK1 R27 mutant did not bind to APT2 (Fig. 3j). Moreover, APT2 was unable to downregulate palmitoylation in the TBK1 R27A mutant (Fig. 3k). Previous studies indicated that the L186 residue on APT2 as crucial for both substrate binding and catalytic activity35. Our experiments further demonstrated that mutation at this site led to a reduced ability to bind to TBK1 and resulted in the loss of its depalmitoylation function (Fig. 3l, m). Therefore, APT2 mediates the depalmitoylation of TBK1 by binding to the R27 residue on TBK1.
APT2 inhibits TBK1-mediated IFN-I responses
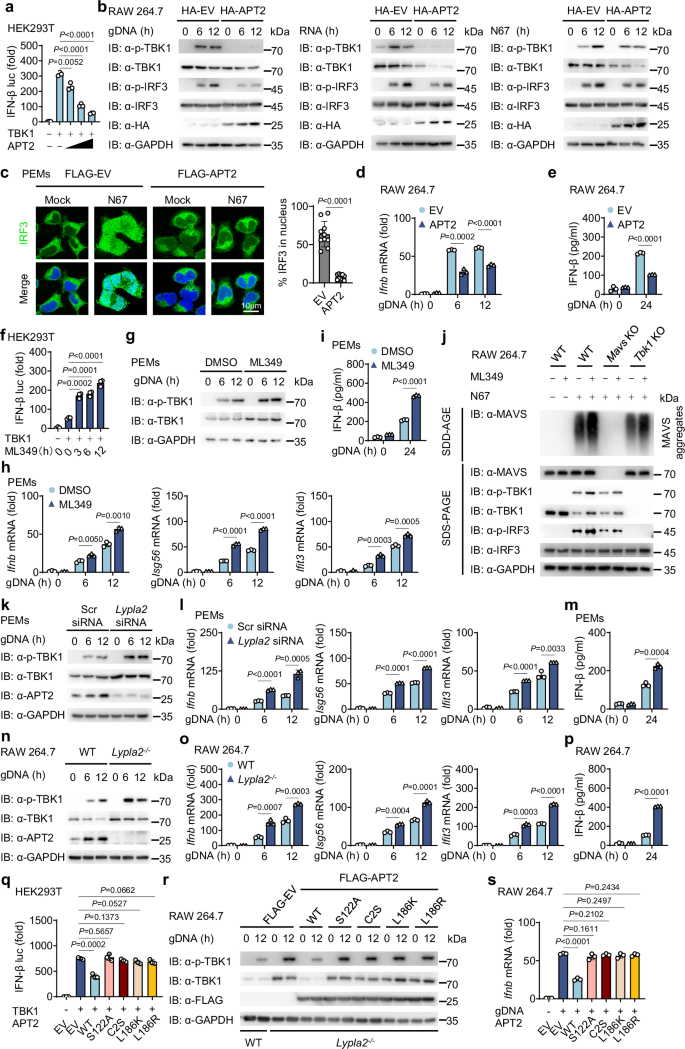
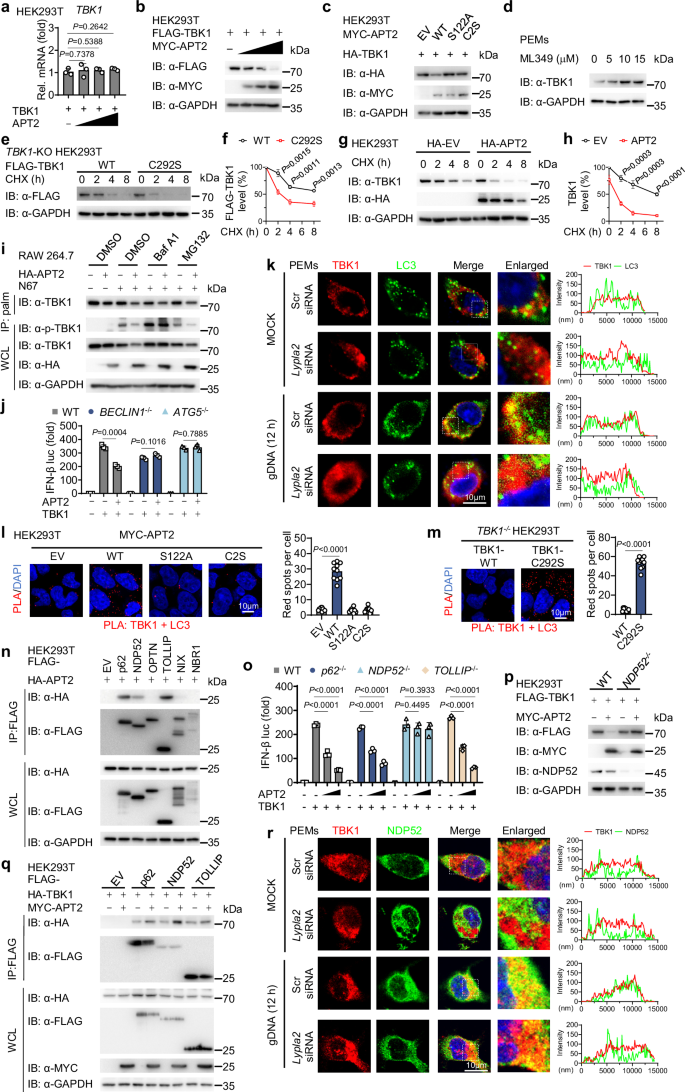
We next investigated whether depalmitoylation of TBK1 by APT2 affects its role in activating IFN-I responses. Overexpressing APT2 in HEK293T cells resulted in a marked inhibition of TBK1-mediated activation of IFN-β reporter gene (Fig. 4a). In parallel, APT2 overexpression significantly suppressed the phosphorylation of TBK1 and IRF3 and inhibited the nuclear translocation of IRF3, as well as the transcription of IFN-responsive genes, including Ifnb, Isg56, and Ifit3, along with the release of IFN-β in RAW 264.7 cells stimulated with gDNA, RNA or freeze-thawed iRBCs (Fig. 4b–e and Supplementary Fig. 4a–d).
a HEK293T cells were transfected with a luciferase reporter for IFN-β-luc, FLAG-TBK1 and increasing doses of MYC-APT2 for 24 h. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). b RAW 264.7 cells were transfected with HA-EV or HA-APT2 for 24 h, then stimulated with gDNA, RNA or freeze-thawed iRBCs at the indicated time points. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. c PEMs were transfected with FLAG-EV or FLAG-APT2 for 24 h, then stimulated with freeze-thawed iRBCs for 12 h. IRF3 (green) and nucleus (DAPI, blue) was examined by confocal microscopy. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) are presented next to the image. This was repeated n = 3 independent times with similar results. d, e RAW 264.7 cells were transfected with HA-EV or HA-APT2 for 24 h, then stimulated with gDNA at the indicated time points. Cell lysates were collected for RT-qPCR analysis (n = 3 biological replicates) (d). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (e). f HEK293T cells were transfected with a luciferase reporter for IFN-β-luc and FLAG-TBK1 for 24 h, followed by treatment with ML349 at different time points. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). g–i PEMs were treated with ML349 (10 μM) for 12 h, then stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (g) and RT-qPCR analysis (n = 3 biological replicates) (h). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (i). j WT, Mavs-KO, or Tbk1-KO RAW 264.7 cells were treated with or without ML349 (10 μM) for 12 h and subsequently stimulated with freeze-thawed iRBCs for 6 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. k–m PEMs were transfected with Scr-siRNA or Lypla2-siRNA for 36 h, then stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (k) and RT-qPCR analysis (n = 3 biological replicates) (l). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (m). n–p WT or Lypla2-KO RAW 264.7 cells were stimulated with gDNA at the indicated time points. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (n) and RT-qPCR analysis (n = 3 biological replicates) (o). IFN-β release in the supernatants was determined by ELISA (n = 3 biological replicates) (p). q HEK293T cells were transfected with a luciferase reporter for IFN-β luc, FLAG-TBK1 and FLAG- WT APT2 or APT2 mutants for 24 h. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). r, s Lypla2-KO RAW 264.7 cells were reconstituted with FLAG- WT APT2 or APT2 mutants for 36 h, then stimulated with gDNA for 12 h. Cell lysates were collected for immunoblot, this was repeated n = 3 independent times with similar results (r) and RT-qPCR analysis (n = 3 biological replicates) (s). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (a, c–f, h, i, l, m, o–q, s). Source data are provided as a Source Data file.
To further elucidate APT2’s role in the IFN-I signaling pathway, we used the inhibitor ML349 to suppress APT2 in HEK293T cells. This manipulation significantly boosted the activity of the IFN-β promoter following the introduction of exogenous TBK1 (Fig. 4f and Supplementary Fig. 4e). In addition, ML349 treatment resulted in increased TBK1 phosphorylation and a strengthened IFN-I signaling response to N67 stimulation (Fig. 4g–i and Supplementary Fig. 4f–i). While ML349 is known to enhance IFN-I signaling pathway via increasing MAVS oligomerization36, the activation of IFN-I by N67 stimulation was markedly diminished in Mavs-KO RAW 264.7 cells compared to WT cells. Nevertheless, ML349 still enhanced IFN-I activation in Mavs-KO RAW 264.7 cells. In contrast, in Tbk1-KO RAW 264.7 cells, ML349 failed to enhance IFN-I activation, resulting in undetectable levels of both basal and stimulated IFN-I activation (Fig. 4j and Supplementary Fig. 4j). In addition, inhibition of APT2, either through knockdown or knockout approaches, led to an enhanced response of IFN-I signals to N67 stimulation (Fig. 4k–p and Supplementary Fig. 4k–r). Notably, WT APT2 overexpression inhibited TBK1-mediated activation of IFN-I, whereas APT2 mutants lacked this inhibitory effect (Fig. 4q), which is consistent with its palmitoyl acyltransferases activity. Next, in Lypla2-KO RAW 264.7 cells, reconstitution of WT APT2 significantly reduced TBK1 phosphorylation and the transcription levels of Ifnb after gDNA stimulation, but this effect was absent with APT2 mutants reconstitution (Fig. 4r, s). Moreover, while APT2 effectively suppressed the activation of WT TBK1-mediated IFN-I, it did not impact the activation of IFN-I mediated by the TBK1 C292S mutant (Supplementary Fig. 4s). Collectively, these results suggest that APT2 promotes the depalmitoylation of TBK1, thereby attenuating TBK1-mediated IFN-I responses.
APT2-mediated depalmitoylation promotes TBK1 autophagic degradation
To elucidate the inhibitory effects of APT2 on IFN-I signaling through TBK1, we considered the role of palmitoylation in protein stability, as previous studies have indicated that this modification is crucial for the stability of proteins such as NLRP337 and MAVS28. We found that increased levels of APT2 led to a significant reduction in TBK1 protein levels without affecting TBK1 mRNA expression (Fig. 5a, b). Notably, only WT APT2, and not the catalytically inactive variant (S122A and C2S), induced degradation of TBK1 (Fig. 5c). Consistently, treatment with APT2 inhibitor ML349 or silencing Lypla2 enhanced TBK1 stability (Fig. 5d and Supplementary Fig. 5a, b). The cycloheximide (CHX) chase assay also revealed that the TBK1 depalmitoylated C292S mutation was more prone to degradation compared to WT TBK1 (Fig. 5e, f). Moreover, APT2 overexpression accelerated endogenous TBK1 degradation, and ML349 treatment stabilized TBK1 protein (Fig. 5g, h and Supplementary Fig. 5c). These findings suggest that APT2-mediated depalmitoylation promotes TBK1 degradation.
a, b HEK293T cells were transfected with FLAG-TBK1 and increasing doses of MYC-APT2 for 24 h. Cell lysates were collected for RT-qPCR analysis (n = 3 biological replicates) (a) and immunoblot analysis, this was repeated n = 3 independent times with similar results (b). (c) HEK293T cells were transfected with HA-TBK1 and WT MYC-APT2 or its catalytically inactivated variants (S122A and C2S) for 24 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. d PEMs were treated with ML349 at indicated concentrations for 12 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. e, f TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h, then treated with cycloheximide (CHX, 100 mg/mL) at the indicated time points. Cell lysates were collected for immunoblot analysis (e). Quantitative analyses (n = 3 biological replicates) (f) of FLAG-TBK1 signals. g, h HEK293T cells were transfected with HA-EV or HA-APT2 for 24 h, then treated with CHX (100 mg/mL) at the indicated time points. Cell lysates were collected for immunoblot analysis (g). Quantitative analyses (n = 3 biological replicates) (h) of TBK1 signals. i RAW 264.7 cells were transfected with HA-EV or HA-APT2 for 24 h, then stimulated with freeze-thawed iRBCs for 12 h and then DMSO (control), bafilomycin A1 (Baf A1) (0.2 μM), or MG132 (10 μM) treatment for 6 h. Cell lysates were collected for ABE assay and immunoblot analysis. This was repeated n = 3 independent times with similar results. j WT, BECLIN1-KO, or ATG5-KO HEK293T cells were transfected with the indicated plasmids. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). k PEMs were transfected with Scr-siRNA or Lypla2 siRNA for 36 h, then stimulated with gDNA for 12 h. The colocalization between TBK1 (red) and LC3 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. l, m HEK293T cells were transfected with MYC-EV, WT MYC-APT2 or its catalytically inactivated variants (S122A and C2S) for 24 h (l). TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h (m). The colocalization between TBK1 and LC3 was examined by PLA. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) is next to the image. This was repeated n = 3 independent times with similar results. n HEK293T cells were transfected with the indicated FLAG-tagged cargo receptors along with HA-APT2 for 24 h, then treated with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. o WT, p62-KO, NDP52-KO or TOLLIP-KO HEK293T cells were transfected with the indicated plasmids. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). p WT or NDP52-KO HEK293T cells were transfected with FLAG-TBK1 in the presence of MYC-APT2 or not for 24 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. q HEK293T cells were transfected with HA-TBK1 and indicated FLAG-tagged cargo receptors in the presence of MYC-APT2 or not for 24 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. r PEMs were transfected with Scr-siRNA or Lypla2 siRNA for 36 h, then stimulated with gDNA for 12 h. The colocalization between TBK1 (red) and NDP52 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (a, f, h, j, l, m, o). Source data are provided as a Source Data file.
TBK1 degradation primarily occurs through the autophagy-lysosomal and proteasome pathways16,38,39. Our immunoblotting results indicated that the degradation of TBK1 mediated by 2-BP or APT2 was inhibited by autophagy-lysosome inhibitors such as 3-methyladenine (3-MA) and bafilomycin A1 (Baf A1), but not by the proteasome inhibitor MG132 (Fig. 5i and Supplementary Fig. 5d), which suggests that 2-BP and APT2-mediated TBK1 degradation occurs predominantly through the autophagy-lysosomal pathway. Meanwhile, our findings revealed that following treatment with Baf A1, APT2 or 2-BP no longer lead to a decrease in p-TBK1 levels (Fig. 5i and Supplementary Fig. 5d). Moreover, upon application of CHX to inhibit new protein synthesis, we observed that the palmitoylation-deficient mutant TBK1-C292S was more prone to degradation compared to WT TBK1, and its p-TBK1 levels also exhibited a concurrent decline. Importantly, when using Baf A1 to stabilize TBK1 protein levels, the reduction in p-TBK1 was effectively blocked (Supplementary Fig. 5e). These results collectively indicate that the degradation of TBK1 protein is a direct cause for the decrease in its phosphorylated form (p-TBK1). To further verify the role specificity of the proteasome and autophagosome in APT2-mediated TBK1 degradation, we used Lactacystin, a more selective proteasome inhibitor than MG132. Western blot analysis showed that treatment with Lactacystin also failed to effectively block APT2-induced TBK1 degradation. However, when we added Baf A1 on the basis of the presence of Lactacystin, the degradation of TBK1 by APT2 was significantly blocked (Supplementary Fig. 5f). These results indicate that simultaneously inhibiting the autophagic lysosomal pathway can effectively prevent APT2-mediated TBK1 degradation, suggesting that the autophagic pathway is crucial for TBK1 degradation mediated APT2. To further assess the degradation of TBK1 by APT2 via autophagy, we examined cells deficient in autophagy-related genes. In BECLIN1– or ATG5-KO HEK293T cells, APT2 lost its ability to impair IFN-I signaling activation (Fig. 5j). Confocal microscopy revealed an interaction between TBK1 and the autophagy marker LC340 following gDNA stimulation, and APT2 knockdown reduced the colocalization between TBK1 and LC3 (Fig. 5k). Subsequent immunoblotting and PLA demonstrated that overexpression of WT APT2 significantly enhanced the co-localization of TBK1 and LC3, whereas the catalytically inactive variants (S122A and C2S) did not (Fig. 5l and Supplementary Fig. 5g). Furthermore, the TBK1 C292S mutant exhibited increased co-localization with LC3 compared to WT TBK1 (Fig. 5m and Supplementary Fig. 5h). Taken together, these findings indicate that APT2-mediated depalmitoylation leads to TBK1 degradation via the autophagy pathway.
Emerging evidence underscores the essential role of cargo receptors in the selective autophagy degradation of substrates16,41,42,43. To investigate this further, we aimed to identify cargo receptors that mediate APT2-induced autophagic degradation of TBK1. Co-IP experiments revealed that APT2 interacts with several cargo receptors, p62, NDP52, and TOLLIP (Fig. 5n). Using the IFN-β luciferase reporter assay and immunoblotting, we found that APT2 failed to inhibit IFN-I signaling or promote TBK1 degradation in NDP52-KO cells, but not in p62- or TOLLIP-KO cells (Fig. 5o, p and Supplementary Fig. 5i, j). In addition, APT2 enhanced the interaction between TBK1 and NDP52, with no significant effects on TBK1 interactions with p62 or TOLLIP (Fig. 5q). Confocal microscopy confirmed the interaction between TBK1 and NDP52 in response to gDNA, and silencing APT2 expression reduced their co-localization (Fig. 5r). Moreover, overexpression of WT APT2 significantly increased the TBK1-NDP52 interaction, while catalytically inactivated variants (S122A and C2S) did not (Supplementary Fig. 5g). Interestingly, the interaction between TBK1 C292S mutant and NDP52 was stronger than that of WT TBK1 (Supplementary Fig. 5h), indicates that depalmitoylated TBK1 is more susceptible to cargo-mediated autophagic degradation. These findings suggest that APT2 facilitates the interaction between TBK1 and NDP52, thereby promoting the selective autophagy degradation of TBK1.
APT2 modulates K48-linked polyubiquitination of TBK1 through TRIM27
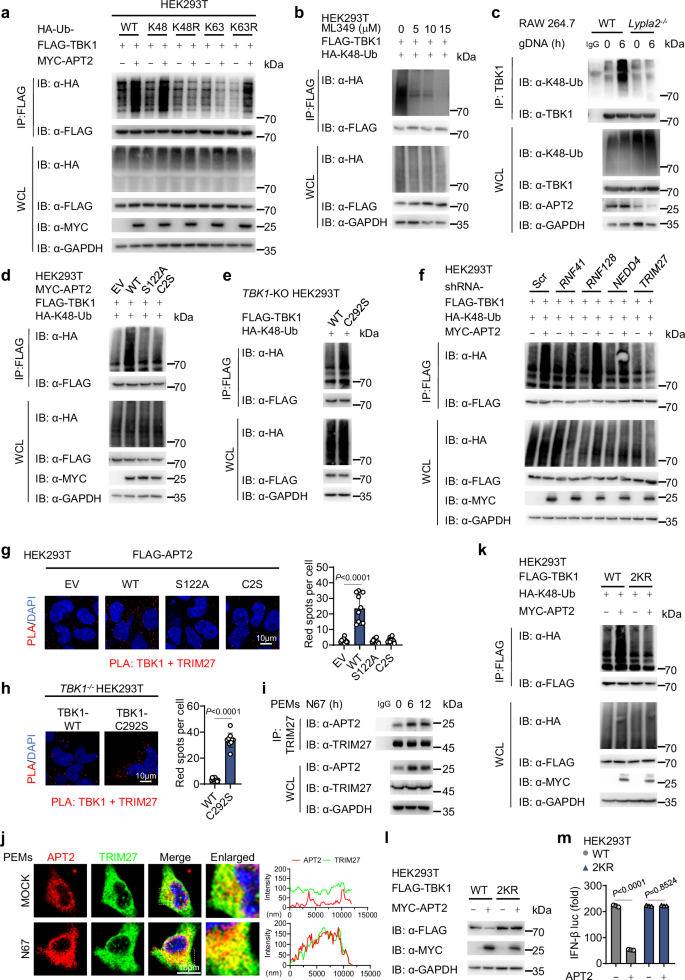
A key step in the selective autophagy degradation process is cargo proteins recognize ubiquitinated substrates, which signals for autophagy-lysosomal degradation. To elucidate APT2’s role in this process, we investigated its effect on TBK1 polyubiquitination and the results revealed that APT2 specifically enhances K48-linked polyubiquitination of TBK1 without affecting other ubiquitin linkages (Fig. 6a and Supplementary Fig. 6a). We also investigated the impact of ZDHHC9 on the ubiquitination of TBK1 and discovered that ZDHHC9 specifically reduced K48-linked polyubiquitination of TBK1 without affecting K63 ubiquitin linkage (Supplementary Fig. 6b). Furthermore, in an in vivo infection model using N67, we observed a notable increase in K48-linked polyubiquitination of TBK1 in the spleen tissues of mice 12 to 24 h post-infection, while levels of K63 ubiquitination remained relatively unchanged (Supplementary Fig. 6c). These results indicate that Plasmodium infection disrupts the dynamic balance between palmitoylation and depalmitoylation of TBK1, and APT2 further depalmitoylates TBK1, rendering it more susceptible to recognition and modification by K48-linked ubiquitin chains, ultimately leading to autophagic degradation pathways. Furthermore, treatment with ML349 decreased K48-linked polyubiquitination of TBK1 in both concentration- and time-dependent manners (Fig. 6b and Supplementary Fig. 6d), further confirming the role of APT2, as silencing APT2 with siRNA also led to reduced K48-linked polyubiquitination of endogenous TBK1 (Supplementary Fig. 6e). In addition, Lypla2 depletion significantly diminished K48-linked polyubiquitination of TBK1 induced by gDNA in RAW 264.7 cells (Fig. 6c). Consistently, APT2 enhances the K48-linked polyubiquitination of TBK1, whereas catalytically inactivated variants (S122A and C2S) failed to produce this effect (Fig. 6d). Notably, the TBK1 C292S mutant exhibited a higher level of K48-linked polyubiquitination compared to WT TBK1 (Fig. 6e), underscoring APT2’s role in promoting the K48-linked polyubiquitination of TBK1.
a HEK293T cells were transfected with FLAG-TBK1 and HA-tagged WT ubiquitin (HA-Ub) or its mutants, together with MYC-EV or MYC-APT2 for 24 h, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. b HEK293T cells were transfected with FLAG-TBK1 and HA-K48-linked ubiquitin for 24 h, followed by treatment with ML349 at indicated concentrations for 12 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. c WT or Lypla2-KO RAW 264.7 cells were stimulated with gDNA for 6 h, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-TBK1) and immunoblot analysis. This was repeated n = 3 independent times with similar results. d, e WT (d) and TBK1-KO (e) HEK293T cells were transfected with the indicated plasmids, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. f HEK293T cells were transfected with Scr shRNA or other E3 ligase-specific shRNAs for 24 h, followed by transfected with FLAG-TBK1, together with HA-K48-linked ubiquitin and MYC-EV or MYC-APT2 for 24 h, and then followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. g, h HEK293T cells were transfected with FLAG-EV, WT FLAG-APT2 or its catalytically inactivated variants for 24 h (g). TBK1-KO HEK293T cells were reconstituted with plasmids encoding WT FLAG-TBK1 or FLAG-TBK1 C292S mutant for 24 h (h). The colocalization between TBK1 and TRIM27 was examined by PLA. The nucleus was stained with DAPI. Scale, 10 μm. Quantitative analyses of the colocalization (10 cells per group) is next to the image. This was repeated n = 3 independent times with similar results. i PEMs were stimulated with freeze-thawed iRBCs at the indicated time points. Cell lysates were collected for IP and immunoblot analysis. This was repeated n = 3 independent times with similar results. j PEMs were stimulated with freeze-thawed iRBCs for 12 h. The colocalization between APT2 (red) and TRIM27 (green) was examined by confocal microscopy. The nucleus was stained with DAPI. Scale, 10 μm. The intensity analysis is next to the image. This was repeated n = 3 independent times with similar results. k HEK293T cells were transfected with WT FLAG-TBK1 or its K251R/K372R (2KR) mutant, together with HA-K48-linked ubiquitin and MYC-EV or MYC-APT2 for 24 h, followed by treatment with Baf A1 (0.2 μM) for 6 h. Cell lysates were collected for IP (with anti-FLAG) and immunoblot analysis. This was repeated n = 3 independent times with similar results. l HEK293T cells were transfected with WT FLAG-TBK1 or its 2KR mutant, together with MYC-EV or MYC-APT2 for 24 h. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. m HEK293T cells were transfected with WT FLAG-TBK1 or its 2KR mutant, together with a luciferase reporter for IFN-β-luc and MYC-EV or MYC-APT2 for 24 h. Cell lysates were collected for luciferase reporter assays (n = 3 biological replicates). Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (g, h, m). Source data are provided as a Source Data file.
Ubiquitin ligase (E3) plays a pivotal role in selecting substrates, and this process is essential for promoting the ubiquitination and subsequent degradation of the substrate protein44. However, APT2 lacks E3 ligase activity, leading us to hypothesize that APT2 enhances TBK1’s K48-linked polyubiquitination by recruiting an E3 ligase. Previous studies have identified several E3 ligases, including RNF41, RNF128, NEDD4, TRIM27, DTX4, TRIP, and TRIM23, as mediators of TBK1 polyubiquitination and degradation15,16,41,45,46. We tested this hypothesis by using shRNA to knockdown the expression of these E3 ligases. Notably, silencing TRIM27 reversed the effects of APT2 on TBK1-induced IFN-β promoter activity, while knockdowns of RNF41, RNF128, and NEDD4 offered only partial rescue (Supplementary Fig. 6f, g). Furthermore, our investigation revealed that APT2 did not enhance K48-polyubiquitination of TBK1 after TRIM27 silencing (Fig. 6f). Immunoblotting analysis and PLA indicated that APT2 facilitates the binding between TBK1 and TRIM27, whereas APT2 S122A and C2S mutants had no effect (Fig. 6g and Supplementary Fig. 6h). In addition, the TBK1 C292S mutant showed increased interaction with TRIM27 (Fig. 6h). Exogenous and endogenous Co-IP experiments directly confirmed the specific binding of APT2 to TRIM27 (Fig. 6i and Supplementary Fig. 6i). In addition, confocal microscopy analysis clearly demonstrated that the interaction between APT2 and TRIM27 was significantly enhanced following stimulation with freeze-thawed iRBCs (Fig. 6j), suggesting that APT2 inhibits the IFN-I signaling pathway by promoting K48-linked polyubiquitination of TBK1 through recruiting TRIM27.
Prior research has established that TRIM27 catalyzes ubiquitination of TBK1 at the K251 and K372 residues47. Our immunoblotting results indicated that mutating these two sites abolished APT2’s ability to enhance K48 ubiquitination of TBK1, culminating in impaired degradation of TBK1 and activation of IFN-I signaling (Fig. 6k–m). These findings highlight the critical role of K48-linked polyubiquitination at K251 and K372 of TBK1 for APT2-mediated autophagic degradation.
ML349 enhances mice resistance to N67 infection
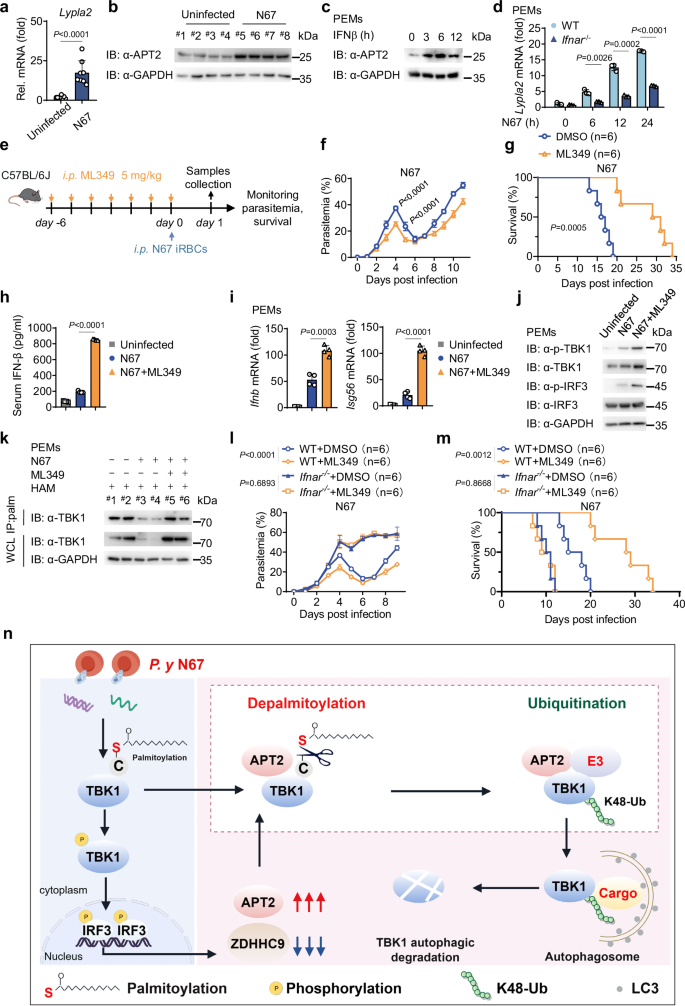
Finally, we examined the impact of N67 infection on the expression of APT2. Both RT-qPCR and immunoblotting analyses revealed a significant increase in APT2 mRNA and protein levels following N67 infection (Fig. 7a, b). Furthermore, stimulation of RAW 264.7 cells or PEMs with freeze-thawed iRBCs also resulted in elevated APT2 expression (Supplementary Fig. 7a–c), confirming that Plasmodium infection promotes the expression of APT2. Notably, recombined mouse IFN-β was able to induce APT2 expression in PEMs (Fig. 7c), and freeze-thawed iRBCs stimulation-induced expression of APT2 was markedly reduced in PEMs derived from Ifnar-deficient (Ifnar-/-) mice (Fig. 7d and Supplementary Fig. 7d). These results indicate that IFN-I can up-regulate the expression of APT2.
a, b C57BL/6 J female mice treated with phosphate-buffered saline (PBS) or infected with N67 via intraperitoneal injection for 24 h. The RT-qPCR analysis of APT2 mRNA presented here is derived from one representative experiment (n = 8 mice per group), and consistent results were observed across three biological replicates (a). In addition, immunoblotting (b) analysis was conducted to assess APT2 protein expression levels, this was repeated n = 3 independent times with similar results. c PEMs were treated with recombined mouse IFN-β at the indicated time points. Cell lysates were collected for immunoblot analysis. This was repeated n = 3 independent times with similar results. d WT and Ifnar-/- PEMs were stimulated with freeze-thawed iRBCs at the indicated time points. Cell lysates were collected for RT-qPCR analysis (n = 3 biological replicates). e–g C57BL/6 J female mice (n = 6 per group) were intraperitoneally (i.p.) injected with DMSO or ML349 (5 mg/kg) for 7 days, followed by an injection of N67 iRBCs (1 × 106, i.p.). Parasitemia (f) and survival rate (g) were then evaluated. This was repeated n = 3 independent times with similar results. Figure 7e was adapted from the image “Adult Mouse” by Yunan Chen (SciDraw; CC BY 4.0). (h) C57BL/6 J female mice were intraperitoneally (i.p.) injected with DMSO or ML349 (5 mg/kg) for 7 days, followed by an injection of N67 iRBCs (1 × 106, i.p.) for 24 h. IFN-β in serum was determined by ELISA. Data shown are from one representative experiment (n = 4 mice per group), consistent results were observed across three biological replicates. i–k PEMs from mice as in (h). Cell lysates were then collected and analyzed by RT-qPCR; the data shown are from one representative experiment (n = 4 mice per group). Consistent results were observed across three biological replicates (i). Lysates were also subjected to immunoblot analysis (j, k), this was repeated n = 3 independent times with similar results. l, m WT and Ifnar-/- C57BL/6 J female mice (n = 6 per group) were intraperitoneally (i.p.) injected with DMSO or ML349 (5 mg/kg) for 7 days, followed by an injection of N67 iRBCs (1 × 106, i.p.). Parasitemia (l) and survival rate (m) were then evaluated. This was repeated n = 3 independent times with similar results. n Graphical abstract created with Figdraw (www.figdraw.com) depicting the mechanism by which Plasmodium evades the host immune response. Data are presented as the mean ± SD. P-values were determined by unpaired two tailed Student’s t tests (a, d, f, h, i), Two-way ANOVA (two tailed) with Tukey’s post hoc test (l) and two-sided log-rank test (g, m). Source data are provided as a Source Data file.
To further explore the role of APT2 in the host immune response to N67 infection, we employed ML349 as a model drug for pharmacological inhibition of APT225. Pretreatment with ML349 significantly reduced parasitemia, improved survival rates, and enhanced the IFN-I response after N67 infection, suggesting that ML349 treatment effectively promoted anti-malarial immune responses (Fig. 7e–j). Subsequent immunoblotting revealed that pretreatment with ML349 markedly increased the palmitoylation level and protein stability of TBK1 (Fig. 7k), suggesting that ML349 exerts its protective effects through modulating TBK1 stability. Next, we investigated whether the resistance conferred by ML349 to N67 infection is influenced by IFN-I by comparing the effect of ML349 in WT and Ifnar-/- mice. Our findings indicated that in the WT mice group, ML349 pretreatment resulted in a lower infection rate and higher survival rate after N67 infection, whereas these benefits were not observed in the Ifnar-/- group (Fig. 7l, m). These findings highlight that the protective effects of ML349 on host anti-malarial immunity depend on the IFN-I signaling pathway. Collectively, these data demonstrate that Plasmodium evades the host immune response by up-regulating APT2 and inhibiting the palmitoylation of TBK1 (Fig. 7n).