Study design and strategy for congruence analysis
Multiple European laboratories, from different countries and sectors, employed, whenever possible, their different bioinformatics pipelines for genomic surveillance of foodborne bacterial pathogens (Fig. 1) on the WGS datasets of L. monocytogenes (n = 3300 isolates50,51), S. enterica (n = 2974 isolates52,53), E. coli (n = 2307 isolates54,55), and C. jejuni (n = 3686 isolates56,57) (details in the Methods section and in Supplementary Data 1). This collaborative effort covered a broad variety of pipelines (hereinafter used as a proxy for the combination of software pipeline and schema/reference), including the most commonly used cg/wgMLST schemas, allele/SNP-callers and clustering methods (Table 1). In order to assess pipeline clustering congruence and comparability, it was essential to obtain clustering information at all possible distance thresholds for each pipeline (both allele- and SNP-based pipelines). Given the heterogeneity of end-point pipeline outputs, this harmonization was achieved with ReporTree58, which also allowed the application of the clustering methods used by the original laboratory, either single-linkage hierarchical clustering (HC) or Minimum-Spanning Tree (MST) generation through MSTreeV2 model of GrapeTree (GT)59 (Table 1, details in the Methods section). In addition, whenever an allele matrix was provided, clustering was performed with both algorithms, reinforcing the magnitude of the comparison (Table 1).
Listeria monocytogenes
Listeria monocytogenes dataset (Fig. 2a) was analyzed with five allele-based and three SNP-based pipelines (Table 1 and Tables S1.1 and S1.2 in Supplementary Data 2).
a Composition of the L. monocytogenes dataset used in this study in terms of ST in comparison with datasets of previous studies (Maury et al. 2016139 and Moura et al. 201660), the LiSeq project140 and the BIGSdb database, as of November 202123. A GrapeTree59 visualization of the MST obtained with the INNUENDO-like pipeline is shown. Nodes (i.e., samples) are collapsed at the threshold with the highest congruence with CC (508 ADs for this pipeline) and colored according to the ST classification. b Number of partitions obtained by each pipeline at each possible distance threshold. c Clustering stability regions determined for each pipeline. To better distinguish each region (represented by separated rectangle blocks), the different blocks are vertically phased, starting in a different line. Distance thresholds (x axis) are presented in log2 scale. d Barplot (top) with the number of samples of the top represented STs (≥50 samples) in L. monocytogenes dataset, with a swarmplot (bottom) indicating the AD threshold at which each pipeline clusters together all samples of each ST. e Distribution of the AD thresholds at which each pipeline clusters together all samples of a given ST (n = 219). Boxplots show the interquartile range (25% to 75%) and median, and whiskers extend 1.5 times the range, with outliers (diamond symbol) plotted separately. The outlier STs are indicated above the respective symbol. Source data are provided as a Source Data file.
Evaluation of allele-based clustering and comparison of stability regions
Following quality control (QC) of the initial 3300 L. monocytogenes isolates (Supplementary Data 1), all pipelines retained >99% of the samples, with the exception of Bionumerics, which only retained ~95% (Table S1.1 in Supplementary Data 2). Despite the intrinsic differences between the pipelines, they generally provided very similar clustering patterns in terms of number of partitions across all possible distance thresholds (Fig. 2b). Independently of the schema (Moura60 or Ruppitsch61) and clustering algorithm (GT or HC), the pipelines consistently revealed low stability (i.e., cluster composition considerably changes in proximal distance thresholds) in the highest resolution region (spanning the outbreak level), and two main plateau regions of high stability (i.e., yielding similar cluster number and composition across a given threshold range), likely reflecting the pathogen population structure and dataset diversity (Fig. 2c).
Evaluation of allele-based clustering congruence with traditional typing
Our results revealed a good correspondence between the first large plateau of stability of all allele-based pipelines and the L. monocytogenes Sequence Type (ST) classification (Fig. 2c and d), with the highest congruence point being identified between 143 and 190 ADs (Table S1.1 in Supplementary Data 2). The maximum CS was very high (~2.9) but not the maximum possible (3.0), indicating that some of the STs were divided into more than one phylogenetic branch at this level of resolution. Between 75% to 90% (depending on the pipeline) of the 106 STs with two or more samples had all samples grouped in the same cluster, and around half of these were fully isolated, i.e., they were not clustering with another ST. Less than 2% of the STs were split into three or more groups regardless of the pipeline. We then identified the lowest threshold level at which all samples of the same ST cluster together in each pipeline (Supplementary Data 3). The majority of the STs congruently cluster below the highest congruence point (albeit at different scales), including prevalent and/or epidemiologically relevant STs, such as ST1, ST5, ST6, ST8 and ST121 (Fig. 2d). This in-depth ST-specific analysis also suggested that some STs were consistently polyphyletic regardless of the pipeline, as it is the case of ST7 and ST325 due to the presence of a few same-ST samples (one and four at the highest CS, respectively) clearly clustering apart (Fig. 2a and e).
Similar results were found when comparing the cgMLST clustering results with L. monocytogenes clonal complex (CC) typing. In this case, the highest congruence point was identified between 388 and 508 ADs, with the maximum CS (>2.97) being even higher than the one obtained with the ST (Supplementary Data 4 and Table S1.1 in Supplementary Data 2). From the 70 CCs with at least two samples, between 60% to 83% (depending on the pipeline) had all samples grouped in the same cluster at the highest congruence point. On average, 89% of these CCs were fully clustered apart, with some of them, including the majority of the CCs with the higher amount of samples, forming a single cluster at thresholds lower than the highest congruence point in all pipelines (Fig. S1.1 in Supplementary Data 2). In contrast, a few CCs were consistently polyphyletic regardless of the pipeline (Fig. S1.1 in Supplementary Data 2), although with different signatures. For instance, while CC5 and CC2 had just a few divergent samples leading to the polyphyletic signature, CC4 and CC14 were divided into two main clusters that could only be merged at high threshold levels (considerably above the highest CS) (Fig. S1.1 in Supplementary Data 2). When comparing the clustering of each CC with the corresponding STs, we noted that CC8 clustered all its samples together at higher AD thresholds than the individual STs (Fig. S1.1 in Supplementary Data 2), particularly due to ST8 and ST16, which differ from each other by more than 200 ADs but reveal low intra-ST diversity.
Evaluation of cluster congruence between different pipelines at all threshold levels
Our in-depth pairwise congruence analysis showed a general high concordance between all allele-based pipelines (as exemplified for a pairwise comparison in Fig. 3a and b, and detailed in Section 2 of Supplementary Data 2). Indeed, the AD threshold points with highest concordance (assessed as CS ≥ 2.85) between every two pipelines (corresponding points) were observed across all levels of resolution and followed a linear trend (r2 ≥ 0.99) in all comparisons (Fig. 3c and d and Supplementary Datas 2, 5 and 6). The slight deviation from a y = x scenario (i.e., theoretical situation in which clustering at one level with a pipeline is concordant with the clustering at the exact same level in the other one) revealed differences in their discriminatory power (Fig. 3d), which corroborated the need for a fine evaluation at outbreak level (next section).
a Heatmap with the CS of two pipelines (details on each pairwise comparison are in Supplementary Data 2, with chewieSnake vs. Bionumerics using the HC algorithm being presented here as an example). The inverted dendrogram (i.e., from the highest to the lowest resolution) and dashed red lines illustrate how the congruence is related with the dataset’s phylogenetic structure (dendrogram obtained with Bionumerics and visualized in auspice.us141). b Zoom-in in the high resolution level highlighted in the orange square of (a). c Bi-directional corresponding points (gray lines) connecting thresholds providing similar clustering in the two pipelines exemplified in (a). d Illustrative linear trend lines expected for the corresponding points with a slope deviation of 10% and 20% to be used as scale reference for the boxplots. Boxplots present the slope distribution for allele vs. allele (orange, n = 58) and SNP. vs. SNP (blue, n = 22) pipeline comparisons for the linear trend lines with a r2 ≥ 0.99, illustrated in Supplementary Data 2 and detailed in Supplementary Data 6 (“n” refers to the number of comparisons with r2 ≥ 0.99 over the total number of comparisons). The boxplot of the allele vs. SNP scenario is not presented due to the low number of comparisons with r2 ≥ 0.99 (Supplementary Data 6). e Density of the distance thresholds required for the identification of clusters detected by at least one allele-based pipeline at 7 ADs. Only clusters having the same composition in all allele-based pipelines were included (n = 316). f Distribution of the difference between the minimum and maximum AD threshold needed to detect the same clusters across allele-based pipelines, using the clusters of (e) (n = 316). g Overlap between the genetic clusters detected at 7 ADs. h Overlap between the genetic clusters detected by one pipeline at 7 ADs and those detected by the others at ≤ 9 ADs. Boxplots in (d) and (f) show the interquartile range and median, and whiskers extend 1.5 times the range, with outliers plotted separately. Source data are provided as a Source Data file.
A similar pairwise comparison was performed between SNP-based pipelines for the top-represented STs in the L. monocytogenes dataset (ST1, ST5, ST6, ST8, and ST121) with a focus on the clustering obtained at up to a 100 SNPs threshold (detailed in Sections 3 and 4 of Supplementary Data 2). We observed an overall concordant clustering regardless of the ST under analysis, as supported by a similar maximum number of possible distance thresholds and number of clusters throughout most levels of resolution (Supplementary Data 2). In most comparisons, this high concordance is also illustrated by the nearly symmetric CS matrices with high scores mainly falling within the diagonal (Section 3 of Supplementary Data 2 and Supplementary Data 6) and by a linear trend of the inter-pipeline corresponding points with a slope very close to 1 (Fig. 3d). A notable exception included an intermediate level of resolution with low congruence between CSI Phylogeny and snippySnake/WGSBAC for ST6 and ST8, despite the good concordance at outbreak level (Section 3 in Supplementary Data 2).
On the contrary, when comparing allele- and SNP-based pipelines, in most situations, we observed (slightly) asymmetric matrices (see heatmaps in Section 4 of Supplementary Data 2), with similar clustering (assessed as high CS) often obtained at higher SNP threshold levels than ADs. These results indicate that SNP-based pipelines, when using the same ST reference for read mapping (a strategy in place in some laboratories), tend to provide a higher discriminatory power than cgMLST pipelines, even though this might not be applicable for all STs. For instance, this asymmetric trend was not so evident for STs 6 and 8 (Section 4 in Supplementary Data 2), as similar cluster composition was observed at similar SNP and AD threshold levels. In general, we found a low number of corresponding points in the pairwise comparisons (assessed at up to 100 ADs/SNPs), which rarely (42/248) yielded a linear trend (i.e., with r2 ≥ 0.99, Supplementary Data 6), thus challenging the overall comparison of the discriminatory power through this approach. Still, there was a high concordance at outbreak level in most situations, as seen in the heatmaps (Section 4 in Supplementary Data 2), showing that a more detailed analysis for outbreak detection is more informative about pipeline performance when comparing allele- and SNP-based pipelines (next section).
As a complementary exercise, a SNP-based pipeline (CSI Phylogeny62) was also applied in a dataset combining the five STs assessed individually using the reference of ST6. As expected, the discriminatory power dropped, reaching a level even lower than the one provided by allele-based pipelines (Sections 3 and 4 of Supplementary Data 2), showcasing that read mapping against a single reference for multiple STs does not provide enough resolution for routine surveillance and outbreak investigation.
Concordance for outbreak detection
Allele-based approaches are the most commonly applied for L. monocytogenes outbreak detection, and the distance threshold corresponding to 7 ADs is conventionally used to determine potential outbreak-related samples60,63. As such, we used this threshold to identify the potential outbreak-related clusters determined by each allele-based pipeline for the L. monocytogenes dataset. Each pipeline detected between 310 to 340 clusters at 7 ADs, from which ~94.2% had similar composition in at least two pipelines and 5.8% were exclusively detected by a single pipeline (Supplementary Data 7). Only ~50% of the clusters detected by a given pipeline was also detected with the exact same composition in all pipelines, but this value is highly influenced by the diversity of the studied pipelines and by the use of a static cut-off. For instance, this value would increase to ~72%, if the most discrepant pipeline was removed (MentaLiST), and to almost 90%, if only same-schema pipelines were compared (Supplementary Data 7).
As these results are impacted by the use of a static threshold, we identified the minimum threshold (ADs or SNPs) at which each 7 AD cluster would be detected by the other pipelines (see the Methods section for details) (Supplementary Data 8). This analysis yielded 316 clusters that, once detected at 7 ADs by at least one pipeline, were detected by all pipelines regardless of the threshold (Supplementary Data 8). As expected, most of these clusters were detected at ≤ 7 ADs in all pipelines, or at higher threshold levels close to 7 ADs (Fig. 3e). The difference between the AD thresholds required by the different allele-based pipelines to detect each outbreak-level cluster had a median of 2 ADs (1 AD, if MentaLiST is excluded), with a minimum of 0 and maximum of 24 ADs (Fig. 3f). Without MentaLiST, the pairwise comparisons of the remaining pipelines showed that the overlap of clusters detected at 7 ADs with the exact same composition was 84.5%, on average, a value that increased to 93.0%, when applying a flexible threshold of up to 2 ADs above (Fig. 3g and h and Supplementary Data 9). The cluster congruence at this level of resolution is influenced by the cgMLST schema used, with pipelines using the same schema yielding more similar results (Fig. 3g and h). This is showcased through the analysis of the threshold flexibilization, in which the overlap increases to 95.2% and 97.5% when only Moura or Ruppitsch pipelines are compared (Fig. 3h), respectively. In a case scenario where the recommended static thresholds for each schema would be applied, i.e., 7 ADs for Moura60 and 10 ADs for Ruppitsch61, the overlap of outbreak signals between pipelines running different schemas would be considerably lower than the one obtained with a flexible approach (Supplementary Data 9). We also tested the application of a more stringent threshold (4 ADs) to identify isolates with more compelling evidence of being part of the same outbreak, followed by the application of a higher cut-off (7 ADs) for identifying probable cases, as previously proposed63. This exercise showed that clusters defined at 4 ADs by a given pipeline are very often captured with the same composition by any other pipeline with a threshold of up to 7 ADs, with the exception of MentaLiST (Supplementary Data 9).
When looking at the genomic diversity (SNPs/ADs) within the cgMLST outbreak-level clusters (7 ADs), our results showed that the maximum allele/SNP distances increase with the size of the cluster and are larger when looking at SNPs (Supplementary Fig. 1). These results are consistent with the previous observation that higher SNP thresholds (which increase alongside with AD thresholds) are needed to identify cgMLST clusters with the exact same composition (Supplementary Fig. 1), while suggesting that, in general, SNP-based pipelines run per ST leverage good resolution to discriminate strains from the same outbreak.
Salmonella enterica
Salmonella enterica dataset (Fig. 4a) was analyzed with seven allele-based and four SNP-based pipelines (Table 1 and Tables S1.1 and S1.2 in Supplementary Data 10).
a Composition of the S. enterica dataset used in this study in terms of serotype and in comparison with datasets of previous studies (INNUENDO22 and BioProject PRJEB20997142), and the Enterobase database, as of November 202164. A GrapeTree59 visualization of the MST obtained with the INNUENDO-like-INNUENDO99 pipeline is shown. Nodes (i.e., samples) are collapsed at the threshold with highest congruence with serotype (1514 ADs for this pipeline) and colored according to the ST classification. b Number of partitions obtained by each pipeline at each possible distance threshold. c Clustering stability regions determined for each pipeline. To better distinguish each region (represented by separated rectangle blocks), the different blocks are vertically phased, starting in a different line. Distance thresholds (x axis) are presented in log2 scale. d Barplot (top) with the number of samples of the top represented serotypes (≥50 samples) in S. enterica dataset, with a swarmplot (bottom) indicating the AD threshold at which each pipeline clusters together all samples of each serotype. Source data are provided as a Source Data file.
Evaluation of allele-based clustering and comparison of stability regions
Following QC of the initial 2974 S. enterica isolates (Supplementary Data 1), a maximum of 2% of the isolates were filtered out by each pipeline (Table S1.1 in Supplementary Data 10). Despite the intrinsic differences between the pipelines, they generally provided very similar clustering patterns in terms of number of clusters across all possible partitions, with the exception of MentaLiST (Fig. 4b). Given the outlier behavior of MentaLiST that, as seen for L. monocytogenes, has a considerable negative impact in pipeline comparisons and interpretation of the results, we decided to remove this tool from all downstream analyses, including for E. coli and C. jejuni.
Independently of the schema (Enterobase64 or INNUENDO22) and clustering algorithm (GT or HC), the pipelines consistently revealed low stability in the region of high resolution spanning the outbreak level, but several regions of high stability could be identified with good correspondence between pipelines, likely reflecting the pathogen population structure and dataset diversity (Fig. 4c). The largest stability region (covering ~690 subsequent AD thresholds) was similar between pipelines, occurred between ~1000 and ~1700 ADs and corresponded to the largest stability region detected in a previous study65.
Assessment of allele-based clustering congruence with traditional typing
Our results revealed a good correspondence between the largest stability region detected in all allele-based pipelines and S. enterica serotype classification (Fig. 4c and d), with the highest congruence point being identified between 1261 and 1663 ADs (CS~2.3) (Table S1.1 in Supplementary Data 10). From the 91 serotypes with at least two samples, between 44% to 68% are grouped in the same cluster at the highest congruence point. This observation is aligned with the results of a large study in which 70.1% of the analyzed serotypes mapped to a single cgMLST cluster in an equivalent stability region65. Still, when focusing on those serotypes having a one-to-one cluster correspondence (i.e., the whole cluster corresponds to all samples of a serotype) in our study, this number decreased to about 30% of the serotypes, regardless of the pipeline. Remarkably, the one-to-one correspondence was detected for the majority of the most prevalent serotypes, although the lowest threshold needed to collapse all samples was quite diverse across serotypes (Fig. 4d). Our analysis also revealed that, at the identified highest congruence point, between 8% to 25% of all serotypes are split into three or more clusters, suggesting the existence of polyphyletic serotypes. Among these, we highlight the Thompson and Newport serotypes (Supplementary Data 11), for which a threshold of more than 2400 ADs was required to collapse all the respective samples, which is in accordance with their previously reported multi-lineage nature65,66,67,68.
Regarding the congruence with the ST, the highest congruence point was identified between 205 and 310 ADs (CS~2.6), always falling within a pipeline stability region (Fig. 4c, Table S1.1 in Supplementary Data 10). From the 112 STs with more than two samples, depending on the pipeline, between 73% to 85% (82 to 95 STs) were grouped in a single cluster at the highest congruence point. On average, 59% of the STs exactly corresponded to a single cluster and a small proportion (between 4% and 9%) were split into three or more clusters. When looking at the earliest threshold to merge all samples of a given ST, some STs clustered considerably below the highest CS (e.g., ST34 and ST26) while others revealed high intra-ST heterogeneity (e.g., ST11 and ST15) (Supplementary Data 12 and Fig. S1.1 in Supplementary Data 10).
In this dataset, Enteritidis serotype is mainly composed of ST11 samples, and our results revealed a good concordance between the thresholds required to merge either Enteritidis or ST11 samples, with ST11 requiring a slightly higher threshold due to few samples not predicted as Enteritidis (Fig. 4d and Fig. S1.1 in Supplementary Data 10). Regarding the samples classified in sílico as Typhimurium, they were segregated into three main STs (ST19, ST34 and ST36) with different levels of intra-ST diversity. This likely justifies why all Typhimurium samples were only collapsed in a single cluster at a high threshold (Fig. 4d and Fig. S1.1 in Supplementary Data 10). Still, we cannot discard that this value is overestimated due to ST34 samples that were classified as Typhimurium instead of its most common classification within the Typhimurium monophasic variant 4,[5],12:i:- (here treated as an independent serotype). Finally, Infantis serotype has a high diversity and a potential polyphyletic signature, which contrasts with its dominant ST (ST32). This is due to a few non-ST32 Infantis samples present in this dataset (Fig. 4d and Fig. S1.1 in Supplementary Data 10).
Evaluation of cluster congruence between different pipelines at all threshold levels
Our in-depth pairwise congruence analysis showed a general high concordance between all allele-based pipelines (as exemplified for a pairwise comparison in Fig. 5a and b, and detailed in Section 2 of Supplementary Data 10). Indeed, the AD threshold points with highest concordance (assumed as CS ≥ 2.85) between every two pipelines (corresponding points) were observed across all levels of resolution and followed a linear trend (r2 ≥ 0.99) in all comparisons (Fig. 5c and d, Supplementary Data 10, 13 and 14). Despite the good inter-pipeline concordance (even at low threshold levels), differences in the discriminatory power were still observed, as shown by deviations from a y = x scenario (Fig. 5d). A fine-tuned analysis of pipeline performance and comparability at the outbreak level is presented below (next section).
a Heatmap with the CS of two pipelines (details on each pairwise comparison are in Supplementary Data 10, with Bionumerics vs. chewieSnake using the HC algorithm being presented here as an example). The inverted dendrogram (i.e., from the highest to the lowest resolution) and dashed red lines illustrate how the congruence is related to the dataset’s phylogenetic structure (dendrogram obtained with chewieSnake and visualized in auspice.us141). b Zoom-in in on the high resolution level highlighted in orange in (a). c Bi-directional corresponding points (gray lines) connecting thresholds providing similar clustering in the two pipelines exemplified in (a). d Illustrative linear trend lines expected for the corresponding points with a slope deviation of 10% and 20% to be used as scale reference for the boxplots. The boxplot presents the slope distribution for allele vs. allele (orange, n = 90) pipeline comparisons for the linear trend lines with r2 ≥ 0.99, illustrated in Supplementary Data 10 and detailed in Supplementary Data 14 (“n” refers to the number of comparisons with r2 ≥ 0.99 over the total number of comparisons). The boxplots of the SNP vs. SNP and allele vs. SNP scenarios are not presented due to the low number of comparison with r2 ≥ 0.99 (Supplementary Data 14). e Density of the distance thresholds required for the identification of clusters detected by at least one allele-based pipeline at 14 ADs. Only clusters having the same composition in all allele-based pipelines were included (n = 255). f Distribution of the difference between the minimum and maximum AD threshold needed to detect the same clusters across allele-based pipelines, using the clusters of (e) (n = 255). g Overlap between the genetic clusters detected at 14 ADs. h Overlap between the genetic clusters detected by one pipeline at 14 ADs and those detected by the others at ≤16 ADs. Boxplots in (d) and (f) show the interquartile range and median, and whiskers extend 1.5 times the range, with outliers plotted separately. Source data is provided as a Source Data file.
Regarding the SNP-based pipelines, the analysis was conducted with a focus on the clustering obtained at up to a 100 SNPs threshold for the top-represented serotypes, namely Enteritidis, Typhimurium, and Infantis, in all pipelines, except for SnapperDB, as the partner institute could only run it for Typhimurium and Infantis subdatasets. As SnippySnake and WGSBAC yielded matching partitions at all levels, only SnippySnake results are presented. SNP-based pipelines revealed considerable differences in the maximum number of thresholds (detailed in Sections 3 and 4 of Supplementary Data 10), which are more pronounced between SnapperDB and the other pipelines. SnapperDB excluded samples that, although being in sílico predicted as Typhimurium or Infantis, were phylogenetically distant from the remaining ones, reducing the number of informative sites in the core alignment and, consequently, leading to a lower maximum number of partitions for these serotypes in this pipeline but a higher resolution power. This variability in sample inclusion/exclusion challenged the assessment of the congruence at all levels between pipelines, as illustrated by the asymmetry of the heatmaps (Section 3 in Supplementary Data 10). Still, most pairwise comparisons were informative, as revealed by the observation of concordant clustering results, especially at lower threshold levels.
When comparing allele- and SNP-based pipelines, we observed (slightly) asymmetric matrices (see heatmaps in Section 4 of Supplementary Data 10), often deviating towards high SNP thresholds (i.e., high CSs are observed when the SNP threshold is higher than the corresponding AD threshold) (e.g., Fig. S4.1.6 in Section 4 of Supplementary Data 10). As such, the SNP-based pipelines, when using the same serotype reference for read mapping, tend to provide a higher discriminatory power than cg/wgMLST pipelines, even though this might not be applicable for all serotypes and pipelines. For instance, this trend was inverted for Enteritidis serotype when using SnippySnake pipeline (e.g., Fig. S4.3.2 in Section 4 of Supplementary Data 10). In general, we found a low number of corresponding points in the pairwise comparisons (assessed at up to 100 ADs/SNPs) and the few identified points did not follow a linear trend (i.e., with r2 ≥ 0.99, Supplementary Data 14), thus hampering a broad assessment of the discriminatory power. Still, the observed concordance trends at the outbreak level showed that a more detailed analysis for outbreak detection is more informative about pipeline performance when comparing allele- and SNP-based pipelines (as addressed in next section).
Concordance for outbreak detection
Allele-based approaches are the most commonly applied for S. enterica outbreak detection, but the method and distance threshold used to determine a possible outbreak-related cluster usually vary between laboratories, and the inclusion criteria for the outbreak is usually set during investigation. The INNUENDO project proposed a dynamic threshold of 0.43% of the cgMLST schema, corresponding to 14 ADs in the INNUENDO cgMLST schema, due to its good concordance with clusters of epidemiologically verified isolates22. We used this 0.43% threshold (which translates into 14 ADs in all pipelines) to start exploring the pipeline congruence at the potential outbreak level.
Each pipeline detected between 216 to 254 clusters at 14 ADs, from which, on average, 95.9% had similar composition in at least two pipelines and 4.1% were exclusively detected by a single pipeline (Supplementary Data 15). On average, 62.6% of the clusters detected by a given pipeline was also detected with the exact same composition by all remaining pipelines. However, this value is highly influenced by the diversity of the studied pipelines and by the use of a static cut-off. Indeed, this value would increase to almost 75%, if only same-schema pipelines are compared (Supplementary Data 15). We further evaluated the minimum threshold level (ADs or SNPs) at which each 14 AD cluster would be detected by the other pipelines. This analysis yielded a total of 255 clusters that, once detected at 14 ADs by at least one pipeline, were detected by all pipelines regardless of the threshold (Supplementary Data 16). As expected, most of these clusters were detected at a threshold ≤ 14 ADs in all pipelines, or at higher threshold levels close to 14 ADs. At SNP level, two different profiles were observed (Fig. 5e). While SnippySnake showed a density profile similar to allele-based pipelines (i.e., a threshold of 14 SNPs would be enough to capture most of the clusters determined at 14 ADs), the other two SNP-based pipelines (SnapperDB and CSI Phylogeny) very often required higher SNP thresholds to merge isolates belonging to the same cluster (Fig. 5e). The difference between the AD thresholds required by the different allele-based pipelines to detect each outbreak-level cluster had a median of 2 ADs, with a minimum of 0 and maximum of 14 ADs. This trend is less influenced by the clustering algorithm rather than the cg/wgMLST schema, as a median of only 1 AD difference is observed when comparing same-schema pipelines (Fig. 5f). Looking at pairwise comparisons between all pipelines, our results showed that the overlap of clusters detected at 14 ADs with the exact same composition was 79.0%, on average, a value that increased to 89.8% when applying a flexible threshold of up to 2 ADs above (Fig. 5g and h, Supplementary Data 17). Importantly, the overall pairwise congruence only slightly decreased when testing thresholds with higher resolution, namely 85.0% for ≤10 ADs, as commonly defined34,69, or 81.6% for ≤ 5 ADs, a strict threshold that has recently been used for case definition in multi-country outbreaks70,71 (Supplementary Data 17).
When looking at the genomic diversity (SNPs/ADs) within the cg/wgMLST clusters at 14 ADs, our results showed that allele-based pipelines behave similarly, with the maximum intra-cluster distance increasing with the size of the cluster, as expected (Supplementary Fig. 2a). Although this trend is also seen at SNP level, SnapperDB and CSI Phylogeny required higher SNP thresholds than SnippySnake to identify cgMLST clusters with the exact same composition (Fig. 5e), and yielded a higher SNP diversity within the clusters (Supplementary Fig. 2b). The evaluation of intra-cluster diversity across incremental distance thresholds also shows that SNP-based pipelines capture a higher diversity than allele-based pipelines. For example, 95% of the clusters detected at 5 ADs by at least one allele-based pipeline were composed of strains that diverge by no more than 11 alleles, a value that increases to 17 when assessed in terms of SNPs (Supplementary Fig. 2c).
Finally, we conducted an additional exercise with the pipeline running an wgMLST schema (INNUENDO-like-INNUENDO99) to explore the potential gain in resolution to discriminate potential outbreak isolates (as assessed by cgMLST) when increasing the number of wgMLST loci under comparison, aligned with a previously explored rationale22,58. Regardless of the clustering algorithm, this approach resulted in an average increase of 6 ADs in the maximum pairwise distances observed between the isolates of the same original cgMLST cluster (Supplementary Data 18), demonstrating the clear increase in resolution provided by the dynamic extension of the cgMLST schema with wgMLST loci shared by the same-outbreak isolates.
Escherichia coli
Escherichia coli dataset (Fig. 6a) was analyzed with seven allele-based and two SNP-based pipelines (Table 1 and Tables S1.1 and S1.2 in Supplementary Data 19).
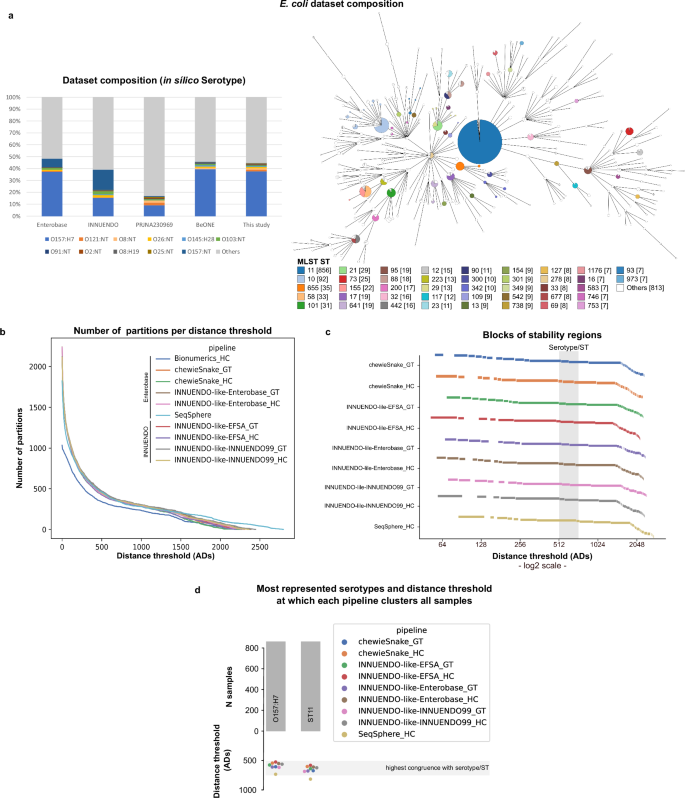
a Composition of the E. coli dataset used in this study in terms of serotype in comparison with the composition of the datasets of previous studies (INNUENDO22 and BioProject PRJNA230969143,144), and the Enterobase database, as of November 202164. A GrapeTree59 visualization of the MST obtained with the INNUENDO-like-INNUENDO99 pipeline is shown. Nodes (i.e., samples) are collapsed at the threshold with the highest congruence with serotype (620 ADs for this pipeline) and colored according to the ST classification. b Number of partitions obtained by each pipeline at each possible distance threshold. c Clustering stability regions are determined for each pipeline. To better distinguish each region (represented by separate rectangular blocks), the different blocks are vertically phased, starting in a different line. Distance thresholds (x axis) are presented in log2 scale. d Barplot (top) with the number of samples of the most represented serotype (O157:H7) and ST (ST11) in E. coli dataset, with a swarmplot (bottom) indicating the AD threshold at which each pipeline clusters together all samples of each of them. Source data are provided as a Source Data file.
Evaluation of allele-based clustering and comparison of stability regions
Following QC of the initial 2307 E. coli isolates (Supplementary Data 1), all pipelines retained more than 99% of the samples, with the exception of SeqSphere and Bionumerics, which only retained 89.99% and 46.25%, respectively (Table S1.1 in Supplementary Data 19). As all pipelines used an inclusion criterion of at least 95% cgMLST loci called, this result was most likely linked to the allele caller than to the schema. Indeed, all pipelines relying on chewBBACA72 retained >99% of the samples, even when using the same schema as SeqSphere and Bionumerics (the Enterobase schema64). Given the very low number of samples that passed the QC of Bionumerics, this pipeline was excluded from the analysis of the E. coli dataset. Despite the intrinsic differences between the remaining pipelines, they generally provided very similar clustering patterns in terms of a number of clusters across all possible thresholds, with the exception of SeqSphere (Fig. 6b), which presented a deviating pattern possibly due to the removal of ~10% of the samples.
Independently of the schema (Enterobase64 or INNUENDO22) and clustering algorithm (GT or HC), the pipelines consistently revealed low stability in the region of high resolution spanning the outbreak level. Beyond this region, multiple regions of high stability could be identified across all levels, including a first high-resolution region (around 60 to 120 ADs), likely reflecting the dataset diversity, as discussed below (Fig. 6c).
Evaluation of allele-based clustering congruence with traditional typing and WGS-derived pathogen main lineages
This assessment is highly influenced by the often incompleteness in the inference of O and H antigens and, specially, by the dominance of serotype O157:H7 strains (almost all from ST11) in the dataset, which reflects the bias towards this pathogenic E. coli in public databases. As consequence, for all pipelines, the highest congruence point was the same for serotype (CS~2.7) and ST (CS~2.9) (Table S1.1 in Supplementary Data 19) and corresponded to the minimum threshold needed to collapse all O157:H7 and ST11 strains, ranging between 545 and 738 ADs (Fig. 6d). This point revealed a good correspondence with one of the largest stability regions (Fig. 6c and d), but, due to the E. coli diversity bias, this result should be eyed with caution, if intended to inform nomenclature design.
Regarding the less abundant serotypes, we observed very different profiles, with those needing a higher threshold to collapse all samples being also the ones with the highest ST diversity. For instance, among the ones with ≥10 isolates, O145:H28 (including ST32 and ST137) was collapsed at around 350 ADs, while O8:H19 (including ST88, ST90, ST201 and ST3233) required around 2250 ADs to merge all same-serotype isolates (Fig. 6d, Supplementary Data 20 and 21). Also, the lack of one-to-one correspondence was also illustrated by the detection of some STs (e.g., ST88 and ST90) comprising isolates from different serotypes.
Finally, we assessed the congruence between cg/wgMLST clustering and WGS-derived pathogen main lineages inferred through PopPUNK73. For this E. coli dataset, PopPUNK clustering had a high congruence (CS > 2.97) at allelic distance thresholds (723 to 1002 ADs) above the level with the highest congruence with serotype and ST (Table S1.1 in Supplementary Data 19).
Evaluation of cluster congruence between different pipelines at all threshold levels
Our in-depth pairwise congruence analysis showed a high concordance between all allele-based pipelines (as exemplified for a pairwise comparison in Fig. 7a and b, and detailed in Section 2 of Supplementary Data 19). Indeed, the AD threshold points with highest concordance (assumed as CS ≥ 2.85) between every two pipelines (corresponding points) were observed across all levels of resolution and followed a linear trend (r2 ≥ 0.99) in all comparisons (Fig. 7c, Supplementary Data 19, 22 and 23). Despite the good inter-pipeline concordance (even at low threshold levels), differences in the discriminatory power were observed, as shown by deviations from a y = x scenario (Fig. 7d). In particular, most differences were seen when SeqSphere was involved, as it revealed a higher resolution across all threshold levels. A fine-tuned analysis about pipeline performance and comparability at outbreak level is presented below (next section).
a Heatmap with the CS of two pipelines (details on each pairwise comparison are in Supplementary Data 19, with INNUENDO-like-Enterobase vs. INNUENDO-like-INNUENDO99 using the HC algorithm being presented here as an example). The inverted dendrogram (i.e., from the highest to the lowest resolution) and dashed red lines illustrate how the congruence is related with the dataset’s phylogenetic structure (dendrogram obtained with INNUENDO-like-INNUENDO99 and visualized in auspice.us141). b Zoom-in in the high resolution level highlighted in orange in (a). c Bi-directional corresponding points (gray lines) connecting thresholds providing similar clustering in the two pipelines exemplified in (a). d Illustrative linear trend lines expected for the corresponding points with a slope deviation of 10% and 20% to be used as scale reference for the boxplots. The boxplot presents the slope distribution for allele vs. allele (orange, n = 68) pipeline comparisons for the linear trend lines with r2 ≥ 0.99, illustrated in Supplementary Data 19 and detailed in Supplementary Data 23 (“n” refers to the number of comparisons with r2 ≥ 0.99 over the total number of comparisons). The boxplot of the allele vs. SNP scenario is not presented due to the low number of comparisons with r2 ≥ 0.99 (Supplementary Data 23). e Density of the distance thresholds required for the identification of clusters detected by at least one allele-based pipeline at 9 ADs. Only clusters having the same composition in all allele-based pipelines were included (n = 185). f Distribution of the difference between the minimum and maximum AD threshold needed to detect the same clusters across allele-based pipelines, using the clusters of (e) (n = 185). g Overlap between the genetic clusters detected at 9 ADs. h Overlap between the genetic clusters detected by one pipeline at 9 ADs and those detected by the others at ≤ 12 ADs. Boxplots in (d) and (f) show the interquartile range and median, and whiskers extend 1.5 times the range, with outliers plotted separately. Source data are provided as a Source Data file.
Regarding the SNP-based pipelines, the analysis was conducted with a focus on the clustering obtained at up to a 100 SNPs threshold for the O157:H7 serotype. In summary, CSI Phylogeny provided a higher resolution than SnippySnake (Section 3 of Supplementary Data 19), but this observation should not be extrapolated without taking into consideration that the former excluded almost 20% of the sequences. When comparing allele- and SNP-based pipelines (see heatmaps in Section 4 of Supplementary Data 19), we again observed a bias towards the higher resolution of CSI Phylogeny, while SnippySnake provided a similar resolution power as the allele-based pipelines.
Concordance for outbreak detection
Allele-based approaches are the most commonly applied for E. coli outbreak detection, but the method and distance threshold used to determine a possible outbreak-related cluster usually varies between laboratories, and the inclusion criteria for the outbreak is usually set during investigation. The INNUENDO project proposed a dynamic threshold of 0.34% of the cgMLST schema, corresponding to 8 ADs in the INNUENDO cgMLST schema, due to its good concordance with clusters of epidemiologically verified isolates22. We used this 0.34% threshold (which translates into 9 ADs in all pipelines) to start exploring the pipeline congruence at the potential outbreak level.
Each pipeline detected between 169 to 182 clusters at 9 ADs, from which, on average, 96.6% had similar composition in at least two pipelines and 3.2% were exclusively detected by a single pipeline (Supplementary Data 24). On average, 70.4% of the clusters detected by a given pipeline were also detected with the exact same composition by all remaining pipelines. We further evaluated the minimum threshold level (ADs or SNPs) at which each 9 AD cluster would be detected by the other pipelines. This analysis yielded a total of 185 clusters that, once detected at 9 ADs threshold by at least one pipeline, were detected by all pipelines regardless of the threshold (Supplementary Data 25), with SeqSphere and INNUENDO-like-ENTEROBASE requiring slightly higher thresholds to yield the same clusters as the other pipelines. Regardless of this observation, most of the outbreak clusters were detected at a threshold ≤ 9 ADs in all allele-based pipelines, or at higher threshold levels close to 9 ADs (Fig. 7e). At SNP level, the assessment was restricted to those outbreak-level clusters corresponding to O157:H7 (94 out of the 185 clusters) and to the SnippySnake pipeline, because of the CSI Phylogeny behavior noticed above. As anticipated above, SnippySnake revealed a density profile similar to allele-based pipelines (Fig. 7e), thus demonstrating that core SNP and cg/wgMLST analyses have equivalent performance for O157:H7 outbreak detection, in line with a previous observation74. The difference between the AD thresholds required by the different allele-based pipelines to detect each outbreak-level cluster had a median of 3 ADs, with a minimum of 0 and a maximum of 15 ADs. This trend is less influenced by the clustering algorithm rather than the cg/wgMLST schema (Fig. 7f). A slightly higher diversity was observed between those pipelines relying on the Enterobase schema (median of 2 ADs) than between the ones relying on INNUENDO (median of 1 AD), possibly due to the fact that the Enterobase schema was run with different allele callers (and also different versions of the same allele caller), while all pipelines relying on the INNUENDO schema used chewBBACA (the allele caller used for its refinement)72,75. A side observation was that the direct use of chewBBACA on the Enterobase schema systematically led to around 2% of loci not called in each sample, even though this did not substantially affect the congruence and outbreak detection performance (Supplementary Data 19). Looking at pairwise comparisons between all pipelines, our results showed that the overlap of clusters detected at 9 ADs with the exact same composition was 83.7%, on average, a value that increased to 94.9% when applying a flexible threshold according to the median estimation above (i.e., 3 ADs above, Fig. 7g and h, Supplementary Data 26). This exercise further corroborated the outlier behavior of SeqSphere and INNUENDO-like-ENTEROBASE, which needed higher thresholds to yield the same clusters as the other pipelines (Fig. 7g, Supplementary Data 25 and 26).
When looking at the genomic diversity (SNPs/ADs) within the cg/wgMLST outbreak-level clusters (9 ADs), the INNUENDO-like-ENTEROBASE consolidated its outlier behavior by capturing a higher genomic diversity within the clusters and showing higher intra-cluster maximum distances with incremental cluster sizes than the other pipelines (Supplementary Figs. 3a). The other outlier pipeline, SeqSphere, could not be used for this particular analysis, due to the unavailability of suitable pairwise distance input within the BeONE project. SnippySnake again provided similar clustering and outbreak resolution power as the allele-based pipelines Supplementary Figs. 3a and 3b). For example, 95% of the clusters detected at 5 ADs by at least one allele-based pipeline were composed by strains that diverge by no more than 9 alleles, a value that only slightly increased to 12 when assessed in terms of SNPs (Supplementary Fig. 3c).
Finally, we conducted an additional exercise with the pipeline running an wgMLST schema (INNUENDO-like-INNUENDO99) to explore the potential gain in resolution to discriminate potential outbreak isolates (as assessed by cgMLST) when increasing the number of wgMLST loci under comparison, aligned with a previously explored rationale22,58. Regardless of the clustering algorithm, this approach resulted in an average increase of 5 ADs in the maximum pairwise distances observed between the isolates of the same original cgMLST cluster (Supplementary Data 27), demonstrating the clear increase in resolution provided by the dynamic extension of the cgMLST schema with wgMLST loci shared by the same-outbreak isolates.
Campylobacter jejuni
Campylobacter jejuni dataset (Fig. 8a) was analyzed with seven allele-based and one SNP-based pipeline (Table 1 and Tables S1.1 and S1.2 in Supplementary Data 28).
a Composition of the C. jejuni dataset used in this study in terms of CC and in comparison with the composition of the INNUENDO22 dataset and the PubMLST database, as of November 202123. A GrapeTree59 visualization of the MST obtained with the INNUENDO-like-PubMLST pipeline is shown. Nodes (i.e., samples) are collapsed at the threshold with highest congruence with CC (839 ADs for this pipeline) and colored according to the ST classification. b Number of partitions obtained by each pipeline at each possible distance threshold. c Clustering stability regions determined for each pipeline. To better distinguish each region (represented by separated rectangle blocks), different blocks are vertically phased, starting in a different line. Distance thresholds (x axis) are presented in log2 scale. d Barplot (top) with the number of samples of the top represented CCs (≥50 samples) in C. jejuni dataset, with a swarmplot (bottom) indicating the AD threshold at which each pipeline clusters together all samples of each CC. Source data are provided as a Source Data file.
Evaluation of allele-based clustering and comparison of stability regions
Following QC of the initial 3686 C. jejuni isolates (Supplementary Data 1), the pipelines using the large and static cgMLST schema from PubMLST (1343 loci)76 retained ~95% of the samples, while the ones with shorter cgMLST schemas retained ~98.5% (Table S1.1 in Supplementary Data 28). The latter either perform cgMLST on a set of core loci dynamically extracted from an wgMLST schema (either 899/2754 core loci of the INNUENDO schema22 or 860/1595 core loci of the Ridom schema77) for this dataset, or run a short and static cgMLST (678/2754 core loci of the INNUENDO schema22) (Table S1.1 in Supplementary Data 28). These three pipeline groups, relying on different schema sizes, revealed distinct clustering patterns in terms of a number of clusters across all possible thresholds (Fig. 8b), with the pipelines using PubMLST schema displaying a higher resolution, i.e., a higher number of clusters for the same threshold.
Independent of the schema and clustering algorithm (GT or HC), no pipeline revealed regions of high stability until 55 ADs, far above the common thresholds for outbreak detection. After this point, although multiple regions of high stability have been identified across all levels, they were quite small, hampering the identification of stability plateaus common to all pipelines (Fig. 8c). This scenario contrasts with the other species analyzed in this study and likely reflects the high genetic diversity, extensive mosaicism and polyclonal population structure of C. jejuni78,79,80.
Evaluation of allele-based clustering congruence with traditional typing and WGS-derived pathogen main lineages
Regarding the cg/wgMLST clustering congruence with CC classification, the pipelines relying on PubMLST schema presented the highest congruence point between 644 and 839 ADs (CS~2.7), while this threshold ranged between 315 and 522 (CS~2.7) for the pipelines with shorter schemas (Table S1.1 in Supplementary Data 28). Still, the clustering of each pipeline at this point yielded a similar number of partitions, which varied between 161 and 229. When assessing the distribution of the 39 CCs across these partitions, only 25% had all the respective samples integrated in the same cluster. In another perspective, almost two-thirds of the CCs have strains dispersed across three or more clusters at this level. Despite the fact that the likelihood of two samples of the same CC cluster together at this cgMLST level is still high (AWC > 0.8 for all pipelines), our results show that the CC classification only slightly mimics C. jejuni clustering at the genome scale. Indeed, the thresholds needed to cluster all samples of the same CC are quite high across all pipelines, almost reaching the size of the respective cgMLST schema, as observed for most of the dominant CCs in this dataset (Fig. 8d and Supplementary Data 29). The comparison between cgMLST clustering and ST classification revealed a more informative scenario, as 78.2% of the 227 STs present in this dataset have all their samples grouped into a single cluster at the highest congruence point with cgMLST (Table S1.1 in Supplementary Data 28), and only 4 to 7% of the STs were split into three or more clusters at this level. Moreover, it allowed the identification of STs with very different genetic heterogeneity in the dataset. For instance, when examining the most prevalent STs in the dataset (Fig. 8d), some (e.g., ST45, ST48 and ST50) exhibited significant intra-ST diversity, which escalates with the cgMLST schema size (as seen in the CC evaluation). In contrast, others displayed less diversity, requiring similar threshold levels for merging all samples from the same ST, regardless of the pipeline used (Supplementary Data 30 and Figure S1.1 in Supplementary Data 28).
Finally, we assessed the congruence between cg/wgMLST clustering and WGS-derived pathogen main lineages inferred through PopPUNK73. For this C. jejuni dataset, PopPUNK clustering had the highest congruence with cg/wgMLST typing (CS~2.6) at allelic distance thresholds consistently falling in between the points of highest congruence with ST and CC (Table S1.1 in Supplementary Data 28).
Evaluation of cluster congruence between different pipelines at all threshold levels
Our in-depth pairwise congruence analysis showed a high concordance between all allele-based pipelines (as exemplified for a pairwise comparison in Fig. 9a and b, and detailed in Section 2 of Supplementary Data 28). Indeed, the AD threshold points with the highest concordance (assumed as CS ≥ 2.85) between every two pipelines (corresponding points) followed a linear trend (r2 ≥ 0.988) in all comparisons (Fig. 9c and d, Supplementary Data 28, 31 and 32). Not unexpectedly, due to the differences in cgMLST schema size between pipelines, the discriminatory power was consistently higher in the PubMLST schema pipelines, as shown by deviations from a y = x scenario (Fig. 9d). Moreover, the pairwise comparisons between these pipelines revealed many corresponding points even within the outbreak region (Supplementary Data 31), which was not observed in the comparisons involving the pipelines with shorter schemas. A fine-tuned analysis about pipeline performance and comparability at the outbreak level is presented below (next section). The pairwise comparisons between allele-based pipelines and the available SNP-based pipeline (CSI Phylogeny) were conducted with a focus on the clustering obtained at up to a 100 SNPs threshold for the top-represented STs, namely ST21, ST50, ST45, ST48 and ST257. Our results showed that CSI Phylogeny, with an ST-specific reference, generally offered superior resolution compared to allele-based pipelines using short schemas, although less frequently than when comparing with those employing PubMLST schemas (Supplementary Data 32).
jejuni. a Heatmap with the CS of two pipelines (details on each pairwise comparison are in Supplementary Data 28, with Bionumerics vs. INNUENDO-like-INNUENDO99 using the HC algorithm being presented here as an example). The inverted dendrogram (i.e., from the highest to the lowest resolution) and dashed red lines illustrate how the congruence is related with the dataset’s phylogenetic structure (dendrogram obtained with INNUENDO-like-INNUENDO99 and visualized in auspice.us141). b Zoom-in in the high resolution level highlighted in orange in (a). c Bi-directional corresponding points (gray lines) connecting thresholds providing similar clustering results in the two pipelines exemplified in (a). d Illustrative linear trend lines expected for the corresponding points with a slope deviation of 10% and 20% to be used as scale reference for the boxplots. The boxplot presents the slope distribution for allele vs. allele (orange, n = 104) pipeline comparisons for the linear trend lines with r2 ≥ 0.99, illustrated in Supplementary Data 28 and detailed in Supplementary Data 32 (“n” refers to the number of comparisons with r2 ≥ 0.99 over the total number of comparisons). The boxplots of the SNP vs. SNP and allele vs. SNP scenarios are not presented due to the low number of comparisons with r2 ≥ 0.99 (Supplementary Data 32). e Density of the distance thresholds required for the identification of clusters detected by at least one allele-based pipeline at 4 ADs. Only clusters having the same composition in all allele-based pipelines were included (n = 430). f Distribution of the difference between the minimum and maximum AD threshold needed to detect the same clusters across allele-based pipelines, using the clusters of (e) (n = 430). g Overlap between the genetic clusters detected at 4 ADs. h Overlap between the genetic clusters detected by one pipeline at 4 ADs and those detected by the others at ≤4 ADs. Boxplots in (d) and (f) show the interquartile range and median, and whiskers extend 1.5 times the range, with outliers plotted separately. Source data are provided as a Source Data file.
Concordance for outbreak detection
Allele-based approaches are expected to become the standard method for C. jejuni outbreak detection, but this is not yet been conducted routinely in most countries. Despite the existence of limited data about the cgMLST resolution levels with good epidemiological concordance, a threshold of 4 ADs has often been pointed out as a proxy for generating outbreaks signals81, with proven application in real investigations81,82. As such, in the present study, in order to start exploring the pipeline congruence at the potential outbreak level, we applied a 4 AD threshold for all pipelines.
Each pipeline detected between 271 to 388 clusters at 4 ADs, from which, on average, 97.1% had similar composition in at least two pipelines and 2.9% were exclusively detected by a single pipeline (Supplementary Data 33). However, as anticipated above, due to the substantially different sizes of the studied cgMLST schemas, the clustering congruence at a 4 AD threshold was considerably higher between pipelines with similar schema sizes (Fig. 9e and f). On average, only 36.6% of the clusters detected by a given pipeline were detected with the exact same composition in all pipelines, but this value considerably increased when this comparison is restricted to pipelines with similar cgMLST schema size (81.7% for pipelines running the PubMLST schema and 58.6% for the others) (Supplementary Data 33). We subsequently sought to evaluate the minimum threshold level (ADs or SNPs) at which each 4 AD cluster would be detected by the other pipelines. This analysis yielded a total of 430 clusters that, once detected at 4 ADs threshold by at least one pipeline, were detected by all pipelines regardless of the threshold (Supplementary Data 34). The cg/wgMLST pipelines with the larger schema very often needed thresholds two or three times higher to provide the same clusters, which reflects their higher resolution power (Supplementary Data 34). When looking at this threshold dispersion in terms of SNPs (with the CSI Phylogeny), most of the cgMLST clusters at 4 ADs were detected at a threshold ≤4 SNPs or at higher threshold levels close to 4 SNPs (Fig. 9e). The difference between the AD thresholds required by the different allele-based pipelines to detect each outbreak-level cluster had a median of 3 ADs, with a minimum of 0 and maximum of 24 ADs. This trend is less influenced by the clustering algorithm than the cg/wgMLST schema, as a median of only 1 AD difference is observed when comparing pipelines running schemas with similar sizes (Fig. 9f). Looking at pairwise comparisons between all pipelines, our results showed that the overlap of clusters detected at 4 ADs with the exact same composition was, on average, 88.6% between pipelines using the PubMLST schema and 75.0% between the pipelines running the shorter schemas (Fig. 9g and Supplementary Data 35). Importantly, given the significant difference in resolution at outbreak level, one would need to decrease the threshold applied in the shorter schema pipelines to a threshold as low as 1 AD to have a proxy of the potential outbreak clusters obtained with the PubMLST schema at the commonly used 4 ADs threshold (Fig. 9h and Supplementary Data 35). This hampers the application of a single static threshold across different pipelines, highlighting the challenge of applying shorter schemas for outbreak detection. Given these results, we exclusively measured the genomic diversity within potential outbreak-level clusters (i.e, 4 ADs) for the pipelines using the PubMLST cgMLST schema. The maximum distance between same-cluster samples per pipeline had an average of 2 ADs, a value that increases to 3 SNPs when mapping to an ST-representative reference (Supplementary Fig. 4).
Finally, we conducted an additional exercise to evaluate the resolution gain obtained when applying a previously explored rationale22,58 in which the cgMLST is dynamically increased to the maximum number of wgMLST shared loci for a given cgMLST cluster. This exercise was conducted for the wgMLST INNUENDO schema (n = 2754 loci), starting either from a static core of 678 loci (INNUENDO-like-INNUENDOcgMLST) or from a core of 899 loci shared by 99% of the studied dataset (INNUENDO-like-INNUENDO99), as well as for the SeqSphere wgMLST schema (n = 1595 loci), starting from a core of 860 loci shared by 99% of the studied dataset (SeqSphere-wgMLST). In all cases, there was a clear gain in the discriminatory power, with an increase of about 5 ADs in the maximum pairwise distances observed between the isolates of the same original cgMLST cluster (Supplementary Data 36). These clusters often enroll isolates with considerably high genetic distances (Supplementary Data 36), thus consolidating the observation that a threshold of 4 ADs is high for pipelines with original short cgMLST schemas, as it would likely overestimate the size of a real outbreak. The application of a dynamic wgMLST approach as the one tested here might mitigate this risk, providing a layer of extra resolution towards a more reliable detection of outbreak signals.
Impact of applying a different assembly pipeline and updating the allele-caller: proof-of-concept study
Given the availability of an alternative assembly pipeline within the Consortium (INNUca22,83), we sought to investigate if its usage in place of AQUAMIS would impact the study outcomes. Additionally, taking into account that the main allele-caller tested in this study (chewBBACA72) was significantly updated while we were conducting our analyses, we extended this proof-of-concept investigation by also comparing the clustering results obtained when two different versions of this software are used, namely the stable chewBBACA v2.8.5 and the recent (at the time of this analysis) chewBBACA v3.3.5. In summary, for each species, we took the INNUENDO-like pipeline (exactly as applied throughout the whole study, i.e., providing the AQUAMIS assemblies as input and using chewBBACA v2.8.5 – details in the Methods section) and compared its clustering results with those obtained when INNUca assemblies or chewBBACA v3.3.5 are used. Similar to the species-specific sections, these exercises were conducted for the two clustering methods used in this study (i.e., single-linkage and MSTreeV2). Our results revealed a high cluster congruence at all threshold levels for all species, with the usage of either INNUca or chewBBACA v3.3.5 yielding barely no deviation from the theoretical y = x scenario (slope varying between 0.984 and 1.003), indicating that the clustering at one level with the original pipeline is highly concordant with the clustering at the exact same level in each of the two alternative workflows (Supplementary Data 37). We further performed an in-depth comparison at the outbreak level. Although our pairwise comparison strategy is quite strict, i.e., only considers a cluster as detected by two pipelines if it has exactly the same composition at the same AD level, the alternative pipelines were able to detect more than 94% and 97% of the clusters yielded by the original pipeline, when INNUca assemblies or chewBBACA v3.3.5 are used, respectively (Supplementary Data 37). These values are considerably higher than those obtained in the inter-pipeline comparisons presented above for each species. Altogether, this exercise shows that the species-specific results and conclusions of this study would be maintained if other assemblies had been provided or if the partners using chewBBACA had updated their respective pipelines to the newest version of this software. In another perspective, it underscored the added value of the tools developed in this study to comprehensively and rapidly evaluate the impact of software updates and changes of components, which is pivotal to ensure the long-term robustness and sustainability of routine genomics workflows.