Identification of immunodominant CD4+ T cell responses to SARS-CoV-2 spike protein
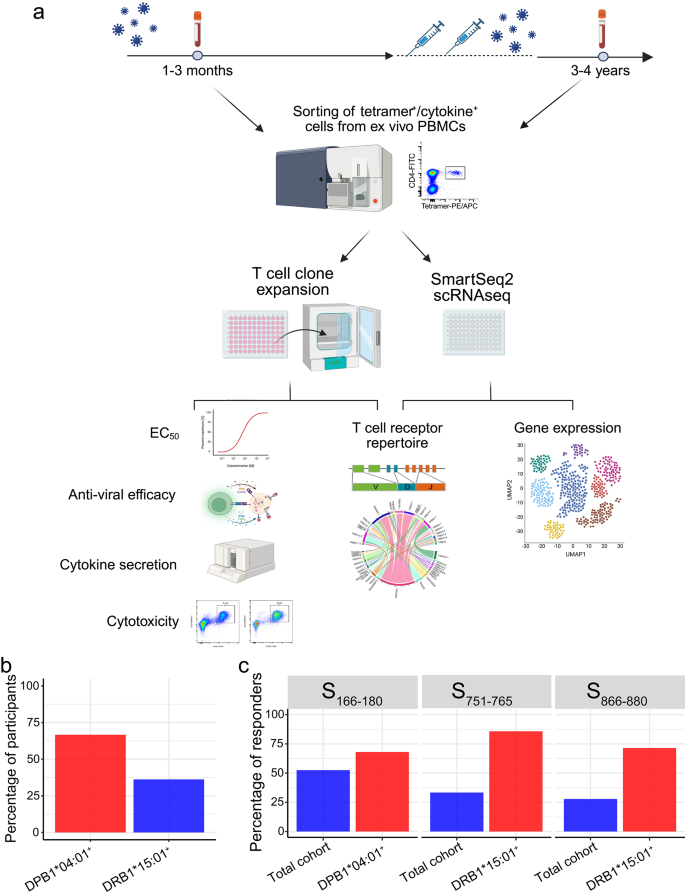
We and others previously identified three dominant SARS-CoV-2 spike protein (S) CD4+ T cell epitopes: S166–180 (CTFEYVSQPFLMDLE)13,14,16, S751–765 (NLLLQYGSFCTQLNR)13,15 and S866–880 (TDEMIAQYTSALLAG)13,34,35. The HLA-restriction of these epitopes was defined using IFN-γ ELISPOT or peptide-MHC-Class II tetramer staining (Supplementary Fig. 1a–c). S166–180-specific T cells are restricted by HLA-DPB1*04:01, while S751–765– and S866–880-specific T cells are restricted by HLA-DRB1*15:01. An overview of the study design can be seen in Fig. 1a. Our cohort comprised 48 individuals who had recovered from COVID-19 (Supplementary Data 1), with 30 (66.7% of 45) being HLA-DPB1*04:01 positive; and 17 (36.2% of 47) carrying HLA-DRB1*15:01 (Fig. 1b). Ex vivo IFN-γ ELISpot analysis using convalescent PBMC samples showed that 68% (17/25) of DPB1*04:01 individuals responded to S166–180, while 85.7% (12/14) and 71.4% (10/14) of HLA-DRB1*15:01 positive patients showed responses to S751–765 and S866–880 respectively (Fig. 1c), confirming the immunodominance of these epitopes.
a Overview of sample collection and study design. Created in BioRender. Dong, T. (2025) https://BioRender.com/r48m616. b Proportion of patients with HLA-DPB1*04:01 (n = 30/45) and DRB1*15:01 (n = 17/47) in overall cohort. c Frequency of convalescent COVID-19 patients with T cells responding to S166–180 (n = 21/40), S751–765 (n = 12/37) or S866–880 (n = 10/36) peptide stimulation.
To further characterise the immune response to these immunodominant epitopes, we generated 50 S166–180-specific T cell clones from four participants, 54 S751–765-specific T cell clones from four participants and 49 S866–880-specific T cell clones from three participants. All those clones were established from 1–3-month convalescent samples, and the TCR clonotype was evaluated. Purity of the T cell clones was confirmed with tetramer staining after each round of expansion, and functional assays were only performed when purity was >95% (Supplementary Fig. 1d–f). To assess antigen-sensitivity of the clones, T cells were co-cultured with B cell lines loaded with titrated peptide and cytokine expression was measured by intracellular cytokine staining (ICS). Antigen-sensitivity of 32 S166–180-specific, 45 S751–765-specific and 48 S866–880-speciifc T cell clones was evaluated. All clones expressed TNF-α, IFN-γ and IL-2 following antigen stimulation (Supplementary Fig. 1g). S866–880-specific T cells showed the highest antigen-sensitivity, with the lowest half maximal effective concentration (EC50) calculated from TNF-α, IFN-γ and IL-2 production.
Dominant TCRα but broad TCRβ clonotypes amongst spike-specific CD4+ T cells
We carried out ex vivo SmartSeq2 to analyse the gene expression and the TCR of spike-specific CD4+ T cells from individuals at both 1–3 months and 3–4 years after infection (Supplementary Table 1). In total, our dataset comprised 702 tetramer-sorted cells from six patients during 1–3 months of convalescence (68 S166–180-specific, 293 S751–765-specific and 341 S866–880-specific CD4+ T cells) and 1735 tetramer-sorted cells from ten patients at the 3–4-years follow-up (1135 S166–180-specific, 266 S751–765-specific and 334 S866–80-specific CD4+ T cells). A further 379 cytokine-sorted S166–180-specific T cells from five patients at 1–3 months of convalescence were analysed for their TCR repertoire.
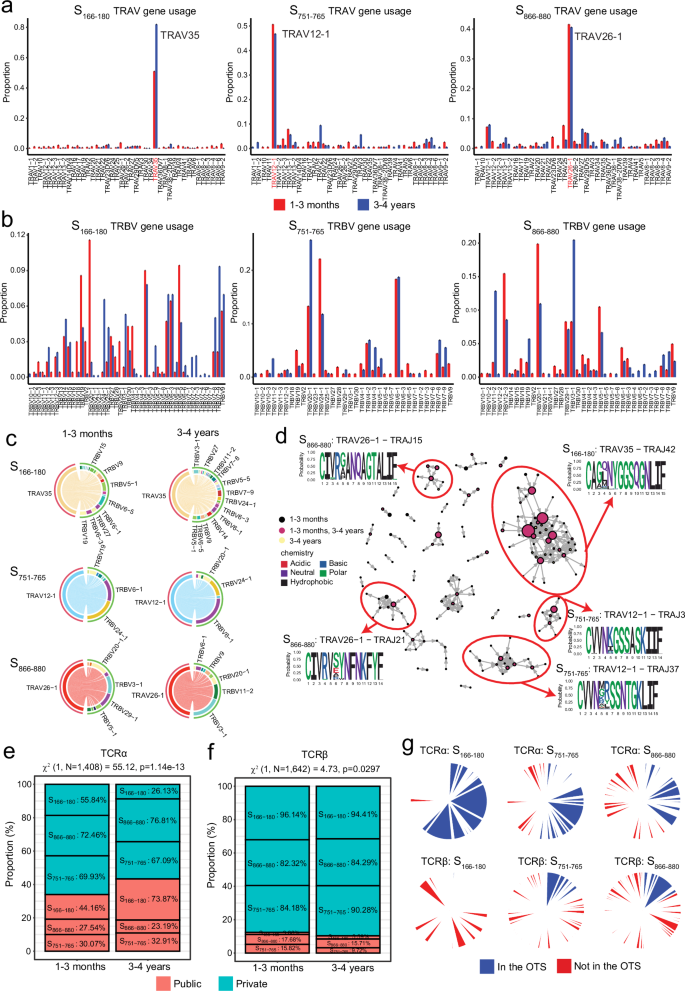
Consistent with previous reports16,29,36,37, we observed a high diversity in V gene usage for both the alpha and beta chains, with a similar broad repertoire at both timepoints (Fig. 2a, b). Interestingly, despite a relatively broad usage of TRAV genes, we identified a bias of TRAV35 used by S166–180-specific cells (66.1% of all S166–180-specific cells), TRAV12-1 used by S751–765-specific cells (45.5% of all S751–765-specific cells) and TRAV26-1 used by S866–880-specific cells (41.5% of all S866–880-specific cells) (Fig. 2a). Unlike the alpha chain, TRBV gene usage appears more diverse across all three epitopes, showing no specific bias (Fig. 2b). Investigating the beta chain pairing with the dominant TRAV genes shows a broad beta gene pairing for S166–180-specific cells with negligible differences between timepoints (Fig. 2c). However, dominant TRAV genes for S751–765– and S866–880-specific T cells show a different preference in beta gene pairing between the timepoints: for S751–765-specific cells, TRBV24-1 is preferential at 1–3 months whereas TRBV6-1 is dominant at 3–4 years; for S866–880-specific T cells, TRBV20-1 pairs dominantly with TRAV26-1 at convalescence, while TRBV3-1 is preferred at 3–4 years (Fig. 2c, d). This finding was similar across all the participants and was not impacted by a particular individual. Overall, we found no difference in TCR clonotype diversity between the two timepoints other than TCRα clonotype diversity of S166–180-sepcific T cells decreased at 3–4 years (Supplementary Fig. 2a, b).
a, b TRAV (a) and TRBV (b) gene usage of S166–180-, S751–765– and S866–880-specific T cells at 1–3 months (red bars) and 3–4 years (blue bars) post primary infection. c TRBV genes pairing with the dominant TRAV gene for S166–180– (TRAV35), S751–765– (TRAV12-1) and S866–880– (TRAV26-1) specific cells at 1–3 months (left) and 3–4 years (right). d TCRα clonotype similarity network. Each vertex corresponds to an individual TCR clonotype, with edges connecting vertices if the CDR3 amino acid sequences show a normalised edit distance >0.9 (scRepertoire). The size of the vertex corresponds to the TCR clonotype frequency, and colour represents the timepoint at which they are found. Cluster motifs were generated using ggseqlogo and amino acid colours based on their biochemical properties. e The proportion of TCRα clonotypes classified as public (coral) or private (cyan) in our dataset. Percentages denote the proportion of cells of a particular epitope specificity classed as public or private. χ2 test of independence was used to compare proportions between 1–3 month and 3–4 years, with two-sided p-values calculated. f Proportion of TCRβ clonotypes classified as public (coral) or private (cyan) in our dataset. Percentages denote the proportion of cells of a particular epitope specificity classed as public or private. χ2 test of independence was used to compare proportions between 1–3 months and 3–4 years, with two-sided p values calculated. g Pie charts denoting the proportion of TCRα (top) and TCRβ (bottom) clonotypes found in the Observed TCR space (OTS).
An earlier study reported that TCRs recognising the same antigen have similar TCR sequences38. We therefore analysed similarity networks for the spike-specific TCR CDR3 alpha clonotypes and identified clusters of highly similar clones (Fig. 2d). A cluster defined as TRAV35-CAGXNYGGSQGNLIF, specific to S166–180, is the largest alpha clonotype. This clonotype corresponded to the dominant TRAV gene used by the S166–180-specific T cells, and is highly public, reported previously16,36. S751–765-specific (TRAV12-1-CVVNXXSSNTGKLIF) and S866–880-specific (TRAV26-1-CIVRXANQAGTALIF) clusters were also identified, corresponding to the dominant TRAV genes used by S751–765– and S866–880-specific T cells, respectively. Similarity network analysis of spike-specific TCR CDR3 beta clonotypes identified smaller clusters compared to the alpha clonotypes (Supplementary Fig. 2c), and these were used for downstream analysis.
We next sought to identify public TCR clonotypes, which are unique clonotypes shared among more than one unrelated individual. We identified 30 public alpha chains specific to S166–180, 12 to S751–765 and 14 to S866–880, as well as 13 public beta chains specific to S166–180, 8 to S751–765 and 8 to S866–880 (Supplementary Data 2). Comparing the proportion of public clonotypes at different timepoints, we observed an increase in the proportion of cells with a public TCRα clonotype at 3–4 years compared to 1–3 months (χ2 = 55.12, p = 1.14 × 10−13 [Fig. 2e]), with 73.87% of S166–180-specific T cells having a public TCRα clonotype at 3–4 years. We did not observe a significant correlation between the number of vaccine doses and the proportion of cells with a public TCRα clonotype at 3–4 years (p = 0.9049). Moreover, we found no evidence of preferential retention of public TCRα clonotypes in specific individuals; all participants exhibited cells with public TCRα clonotypes. In contrast, the TCRβ clonotype showed a much lower overall proportion of cells with a public TCRβ clonotype (Fig. 2f), although there is a slightly greater proportion of cells with a public TCRβ clonotype at 1–3 months compared to 3–4 years (χ2 = 4.73, p = 0.03 [Fig. 2f]). To confirm the public nature of these clonotypes and eliminate the potential risk of cross-contamination, we used the observed TCR space (OTS) database39, which includes 3,185,982 paired TCRS from 892 individuals reported in 13 independent COVID-19 studies. 84.1% of the alpha clonotypes and 62.1% of the beta clonotypes classified as public in our dataset were also identified in the OTS. Using the OTS, we identified a total of 230 unique public alpha clonotypes and 75 unique public beta clonotypes (Fig. 2g).
Dominant TCRα clonotypes are associated with mild COVID-19 disease during primary infection
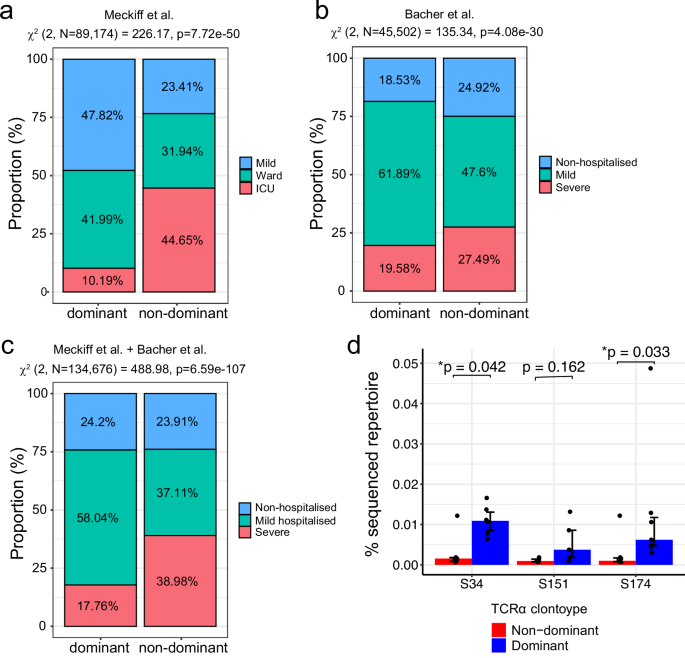
To investigate whether these public dominant TCRα clonotypes are associated with COVID-19 disease severity, we utilised two publicly available datasets of SARS-CoV-2-reactive CD4+ T cells with available TCR information, identified based on the upregulation of CD154/CD137/CD69 expression following peptide pool stimulation, for 52 COVID-19 patients29,37 (Supplementary Table 2). Cells were classified as having a dominant TCRα clonotype if their CDR3α matches the identified clonotypes, TRAV35 (S166–180), TRAV12-1(S751–765) and TRAV26-1(S866–880) shown in Fig. 2d, whereas the rest were considered as non-dominant. For both the ref. 29 and the ref. 37 datasets, we identified a significant association between having the dominant TCRα clonotype and mild COVID-19 disease (χ2 = 226.17, p = 7.72 × 10−50 [Fig. 3a] and χ2 = 135.34, p = 4.08 × 10−30 [Fig. 3b] respectively). Furthermore, since these two datasets used the same method to identify SARS-CoV-2-reactive CD4+ T cells, combining them allowed us to find a significant association of the dominant TCRα clonotype to the mild disease outcome (χ2 = 488.98, p = 6.59 × 10−107, Fig. 3c). To confirm this association was not driven either HLA-DPB1*04:01 or HLA-DRB1*15:01-specific associations, we split the analysis to compare the proportion of cells with the dominant S166–180-specific TCRα clonotypes compared to non-dominant in HLA-DPB1*04:01+ individuals, and compared the proportion of cells with the dominant S751–765-specific or S866–880-specific TCRα clonotypes compared to non-dominant in HLA-DRB1*15:01+ individuals (Supplementary Fig. 3). The dominant S166–180-specific TCRα clonotypes in HLA-DPB1*04:01+ individuals are associated with mild COVID-19 disease (Supplementary Fig. 3a), while the dominant S751–765-specific or S866–880-specific TCRα clonotypes in HLA-DRB1*15:01+ participants are associated with non-hospitalised COVID-19 (Supplementary Fig. 3b). Therefore, the difference in disease severity proportion is likely not driven by any particular epitopes. In summary, these data suggest that these dominant TCRα clonotypes play an important role in protecting individuals from developing severe disease.
a, b Frequency of cells from the Meckiff et al. dataset (a) and the Bacher et al. dataset (b) with dominant and non-dominant TCRα clonotypes split by COVID-19 disease severity of the donor. χ2 test of independence was used to compare proportions between dominant and non-dominant TCRα clonotypes. χ2 test of independence was used to compare proportions between dominant and non-dominant and two-sided p values calculated. c Frequency of cells from the Meckiff et al. and Bacher et al. datasets combined with dominant and non-dominant TCRα clonotypes split by COVID-19 disease severity of the donor. χ2 test of independence was used to compare proportions between dominant and non-dominant TCRα clonotypes and two-sided p values calculated. Non-hospitalised COVID = Meckiff Mild + Bacher non-hospital; Mild hospitalised = Meckiff Ward + Bacher mild; Severe = Meckiff ICU + Bacher severe. d Percentage of the total TCR repertoire of dominant (blue bars) and non-dominant (red bars) TCRα clonotypes in pre-pandemic individuals (n = 6) from the Spindler et al. dataset. Plotted at median ± IQR. The Wilcoxon signed-rank test was used to compare between groups and two-tailed p values calculated.
Given the highly expanded and public nature of these dominant TCRα clonotypes, we explored the possibility that these TCRs may arise from sequences that are present at a higher-than-average frequency in a naïve, pre-pandemic repertoire. Spindler et al.40 carried out paired TCR sequencing of six healthy individuals prior to 2020 and identified 513,963 unique TCRα clonotypes, with an average of 85,660 clonotypes per individual. From this dataset, we calculated the frequency of the dominant and non-dominant TCRα clonotypes. The dominant S166–180-specific and S866–880-specific TCRα clonotypes appeared at a significantly higher frequency in the pre-pandemic repertoires compared to the non-dominant clonotypes (p = 0.042 and p = 0.033 for S166–180 and S866–880 respectively, Fig. 3d). The dominant S751–765-specific TCRα clonotype was detected at a higher frequency than non-dominant clonotypes in the pre-pandemic repertoires, although this did not reach statistical significance.
As SARS-CoV-2-specific responses have been reported to possibly originate from cross-reactive CMV-specific T cell41 and pre-existing cross-reactive CD4+ T cell responses37, we cross checked our TCR clonotypes against those reported in the Immune Epitope Database (IEDB) and VDJ database (VDJdb). None of the public TCRs identified in this study were previously reported to be reactive against other viruses on an HLA-matched background.
TCRα clonotypes maintained at 3-4 years show distinctive cytokine secretion
Given the longitudinal nature of our dataset, we examined whether clonotypes present during the convalescent period persisted for 3–4 years, or whether they are lost and replaced by new clonotypes, due to repeated vaccinations and infection with SARS-CoV-2 variants.
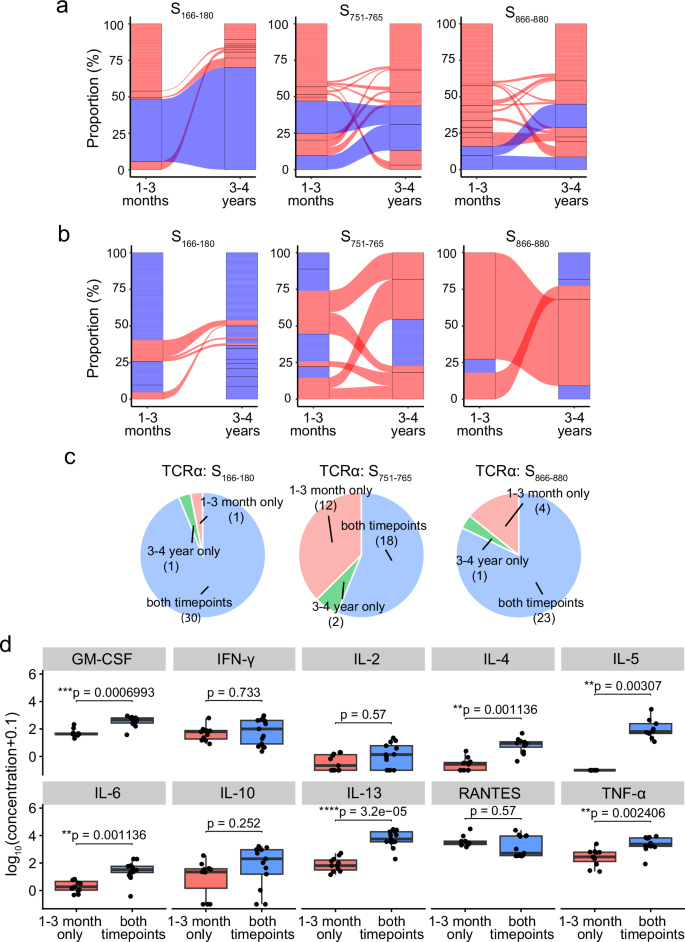
At 3–4 years, 80% of the S166–180-specific cells retain a TCRα clonotype present at 1–3 months, predominantly the expanded and dominant public TCRα clonotype of TRAV35-CAGXNYGGSQGNLIF (Fig. 4a left); 46.2% of the S751–765-specific cells have a TCRα clonotype found at 1–3 months, of which 65.8% are the dominant TCRα clonotypes of TRAV12-1-CVVNXXSSNTGKLIF and TRAV12-1-CVVNXGSSASKIIF (Fig. 4a middle); and 49.3% of the S866–880-specific cells have a TCRα clonotype found at the 1–3-month timepoint, of which 50% are the dominant TCRα clonotypes of TRAV26-1-CIVRXANQAGTALIF and TRAV26-1-CIVRX[Y/W]NFNKFYF (Fig. 4a right).
a Alluvial plots highlighting the TCRα clonotypes at 1–3 months and 3–4 years, with links between columns denoting clonotypes found at both timepoints for S166–180– (left), S751–765– (middle) and S866–880-specific (right) T cells. Blue links are the previously identified dominant TCRα clonotypes. b Alluvial plots highlighting the TCRβ clonotypes that pair with the previously identified dominant TCRα clonotypes at 1–3 months and 3–4 years, with links between the columns denoting clonotypes found at both timepoints. Red clonotypes are clonotypes found at both timepoints, whereas blue blocks are clonotypes found only at one timepoint. c Pie charts of the available clonal functional data based on whether the clone TCRα clonotype is found at both timepoints (blue), 1–3 month only (pink) or 3–4 year only (green) in the single cell data for S166–180-specific (top), S751–765-specific (middle) and S866–880-specific (bottom) clones. d Luminex assay results showing the expression of ten cytokines released by S751–765-specific clones split by whether the clones have a TCRα clonotype found at both timepoints (n = 13) or only found at 1–3 months (n = 11). Y-axis corresponds to log10[concentration (pg/ml) + 0.1]. Boxplots represent the 25th and 75th percentiles with the median marked with whiskers at ±1.5*IQR. The Wilcoxon signed-rank test was used for comparison between groups, and two-sided p values were calculated.
In contrast to the TCRα clonotypes, at 3–4 years, a lower frequency (10%) of S166–180-specific cells have TCRβ clonotypes present at 1–3 months when compared to two other epitope-specific T cells, with 44.4% of the S751–765-specific cells and 45.7% of S866–880-specific T cells with a TCRβ clonotype also found at 1–3 months (Supplementary Fig. 4a). We next looked at the TCRβ clonotypes pairing with the previously identified dominant TCRα clonotypes (Fig. 4b). Similarly, TCRβ clonotypes pairing with the dominant S166–180-specific TCRα clonotype show very little sharing between 1–3 months and 3–4 years, compared to S751–765 and S866–880-specific T cells, which show expanded TCRβ clonotypes paring with the dominant S751–765– and S866–880-specific TCRα clonotypes, maintained at 3–4 years.
As we identified clonotypes that were either maintained between the two timepoints sampled or not detected at the later timepoint, we interrogated our in vitro functional data of T cell clones isolated from 1–3-month samples to investigate whether the T cell clones with clonotypes maintained at the later timepoint showed differences in overall function. T cell clones were annotated as a maintained clonotype if their clonotype matched a TCR clonotype present in the single-cell TCR-seq dataset at both timepoints. Otherwise, T cell clones were categorised as either ‘1–3 month only’ or ‘3–4 year only’ if their TCR clonotype was found exclusively at one timepoint in the single-cell TCR-seq (Fig. 4c). Compared to the clones classified as ‘1–3 months only’, cytokine expression of S751–765-specific T cell clones with maintained clonotypes showed significantly higher secretion of GM-CSF (adjusted p = 0.0014), IL-4 (adjusted p = 0.0014), IL-5 (adjusted p = 0.0042), IL-6 (adjusted p = 0.0042), IL-13 (adjusted p = 1.04 × 10−5) and TNF-α (adjusted p = 0.005) (Fig. 4d). However, when investigating the functional avidity of the S751–765-specific T cell clones, no significant differences were observed in the IL-2 EC50, TNF-α EC50 or IFN-γ EC50 between the two groups (Supplementary Fig. 4b, adjusted p > 0.05), suggesting that the clonal persistence may not be driven by functional avidity.
Taken together, these data suggest that spike-specific T cells with TCRα clonotypes detectable at both timepoints show differentiated cytokine profiles with higher expression of Th2 cytokines, compared to the T cells with TCRα clonotypes that were not detected at the 3–4-year timepoint.
Spike-specific CD4+ T cells show distinct transcriptional profiles over time
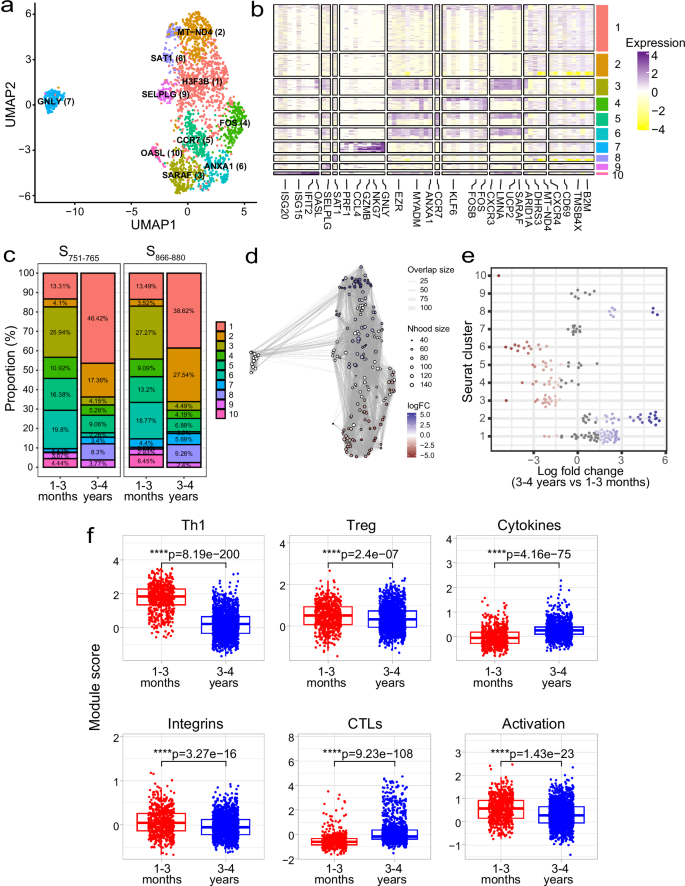
To investigate potential transcriptomic differences in the spike-specific T cells between the two timepoints, we analysed our scRNA-seq dataset that comprised 2213 total cells collected from two timepoints (1–3 months = 634 cells, 3–4 years = 1579 cells) across three epitope specificities (980 S166–180-specific, 558 S751–765-specific and 675 S866–880-specific cells) from 13 individuals. Spike-specific T cells from the different individuals were integrated, and unsupervised clustering revealed ten distinct clusters based solely on their gene expression profiles (Fig. 5a, b). Interestingly, a unique cluster of cytotoxic T cells (CTL, cluster 7) exhibited high expression of cytotoxicity-related genes (e.g. GNLY, GZMH, NKG7 and GZMB). In addition, cells in this cluster showed elevated expression of the apolipoprotein B mRNA editing enzyme, catalytic polypeptide (APOBEC) genes APOBEC3C and APOBEC3G. Other clusters that were identified included an interferon-stimulated genes (ISGs) positive cell cluster (cluster 10, marker genes include OASL, IFIT2 and ISG15), two central memory clusters (clusters 5 and 1, with markers genes including CCR7 [cluster 5] and CXCR4 and IL7R [cluster 1]), and two clusters showing increased activation markers (clusters 4 and 6, with marker genes including FOS, DUSP1 and CD69 [cluster 4] and ANXA1 and LMNA [cluster 6]). Clusters were generally comprised of cells from all patients and all sequencing runs (Supplementary Fig. 5a, b), indicating the clustering analysis does not represent patient-specific subpopulations or batch effects.
a Uniform manifold approximation and projection (UMAP) visualisation of 2213 cells profiled ex vivo from PBMC samples. Cells are coloured based on their cluster occupancy. b Heatmap of the expression levels of the top differentially expressed genes for each cluster. c Stacked bar plots of the proportion of cells in each cluster split by timepoint. Shown here for S751–765– and S866–880-specific cells but not S166–180-specific cells due to the lack of S166–180-specific cells profiled at 1–3 months convalescence. d Milo analysis of differentially abundant cell clusters between 1–3 months and 3–4 years convalescence. Plot represents the embedding of the Milo differential abundance, where each node is a neighbourhood and node size is proportional to the number of cells in that neighbourhood. Colours represent the level of differential abundance. e Beeswarm plot showing the cell abundance changes between 3–4 years and 1–3 months convalescence. Neighbourhoods overlapping the same Seurat cluster identified in (a) are grouped together and neighbourhoods exhibiting significant differential abundance are coloured in red (higher at 1–3 months) or blue (higher at 3–4 years). f Boxplots comparing module scores for cells from 1–3 months (n = 634 cells) and 3–4 years (n = 1579 cells) convalescence. Boxplots represent the 25th and 75th percentiles with the median marked with whiskers at ±1.5*IQR. Wilcoxon signed-rank test was used to compare between groups and two-sided p values calculated.
As scRNA-seq of S166–180-specific cells at 1–3 months was conducted using cytokine-sorted cells and these cells were excluded from the longitudinal transcriptional analysis. We investigated whether there were any proportional differences in the number of cells from each timepoint across different clusters of S751–765-specific and S866–880-specific T cells. Although no differences were evident between these two epitope-specific cells, differences were observed when comparing 1–3-month and 3–4-year samples (Fig. 5c). At 3–4 years, the proportion of cells in clusters 1, 2 and 8 increased, while the cell populations at clusters 3, 4 and 6 declined compared to 1–3 months. This was confirmed by differential abundance testing using the miloR package (Fig. 5d, e).
To further investigate differences between memory cells from 1–3 months and 3–4 years, we generated module scores using gene lists compiled from the literature (Supplementary Table 3). Cells from 1–3-months exhibited a greater expression of genes involved in the positive regulation of Th1 differentiation (ANXA1, CCR7 and IRF1), a greater level of integrin expression (SELPLG, ITGAE and ITGA4), and genes involved in T cell activation (CD44, CD27 and TUBA1B) (p 5f). In contrast, cells from 3–4-year exhibited a greater cytokine gene expression (IL16, IL32 and CCL4) and cytotoxicity (GZMA and GNLY) (p 5f). Moreover, this appears to be independent of epitope specificity (Supplementary Fig. 6a). Comparing the module scores for the S751–765 and S866–880 specific cells showed consistent differences, with increased Th1, Treg, integrin expression and activation signatures at 1–3 months and an increased cytotoxicity and cytokine signature at 3–4 years.
Spike-specific CD4+ T cells at 3-4 years exhibit increased cytotoxicity signatures driven by GZMA expression, correlating with viral suppression in vitro
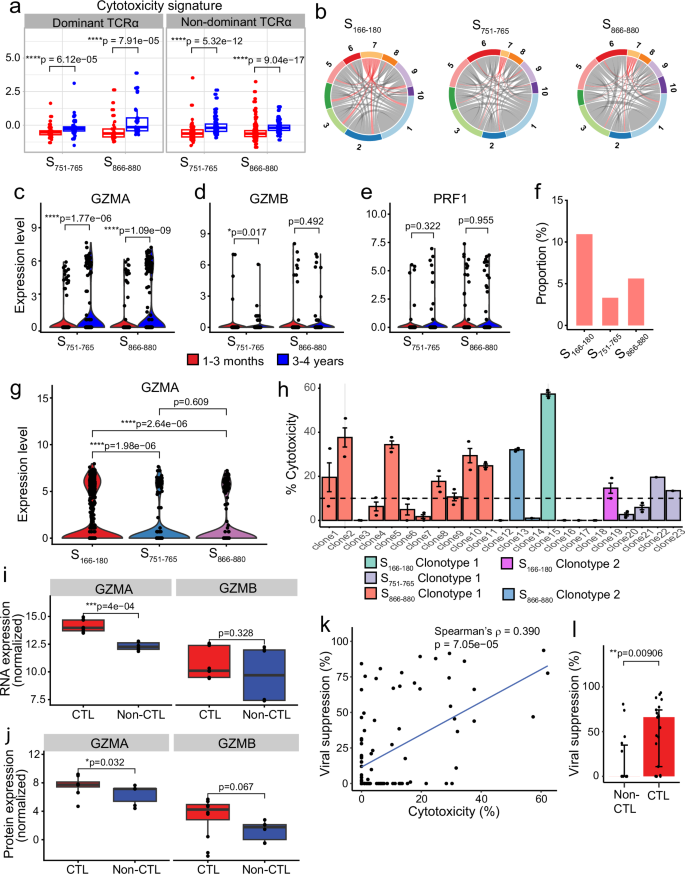
Given the increased cytotoxicity signature observed at 3–4 years post infection, we further investigated whether this difference is driven by TCR clonotype. We categorised the cells into dominant or non-dominant TCRα groups. S751–765– and S866–880-specific CD4+ T cells at 3–4 years consistently showed a greater cytotoxicity signature compared to cells at 1–3 months, regardless of whether the cells had a dominant or non-dominant TCRα clonotype (Fig. 6a). We confirmed this by examining clonotype sharing between cluster 7 (the CD4+ CTL cluster) and the remaining clusters. Our analysis revealed that TCRα clonotypes are not restricted to cluster 7 but shared amongst all clusters (Fig. 6b), indicating that the difference in cytotoxicity is likely to be independent of TCR.
a Boxplots comparing the cytotoxicity signature of cells at 1–3 months (red) and 3–4 years (blue) convalescence for S751–765– and S866–880-specific cells, split by whether cells have a dominant TCRα clonotype or not (n = 49 [1–3 months] and 48 [3–4 years] cells for S751–765-specific cells with dominant TCRα; n = 33 [1–3 months] and 34 [3–4 years] cells for S866–880-specific cells with dominant TCRα; n = 101 [1–3 months] and 110 [3–4 years] cells for S751–765-specific cells with non-dominant TCRα; n = 166 [1–3 months] and 104 [3–4 years] cells for S866–880-specific cells with non-dominant TCRα). b Circos plots highlighting the clusters that share TCRα clonotypes found in cluster 7 (CD4+ CTL cluster). c–e Violin plots comparing the expression of GZMA (c), GZMB (d) and PRF1 (e) in cells between 1–3 months (red) and 3–4 years (blue). f Proportion of 3–4-year epitope-specific cells in the cytotoxic T cell cluster (CTL) compared to all other clusters (non-CTL). g Violin plots comparing the expression of GZMA in cells at 3–4 years between the three epitope-specific cells. h Cytotoxicity of S166–180-, S751–765– and S866–880-specific T cell clones with the different TCR clonotypes. Each bar represents an individual T cell clone, and different coloured bars represent different clonotypes (3T cell clones with S166-180 clonotype 1 and 2 clones with clonotype 2; 4 clones with S751–765 clonotype 1; 12 clones with S866–880 clonotype 1, and 2 clones with clonotype 2), plotted as median±IQR. i, j Boxplots comparing the expression of GZMA and GZMB in cytotoxic (n = 3 clones in triplicates) and non-cytotoxic (n = 2 clones in triplicates) CD4+ T cell clones using bulk RNAseq (i) and bulk proteomics (j). k Correlation of virus suppression with T cell clone cytotoxicity (n = 98, including 33 S166–180-, 30 S751–765– and 35 S866–880-specific CD4+ T cell clones). l Comparison of virus suppression between S866–880-specific CD4+ cytotoxic (n = 21) and non-cytotoxic clones (n = 14), plotted as median ± IQR. The Wilcoxon signed-rank test was used to compare between groups (a, c–g, l), while a paired Wilcoxon signed-rank test was used for the bulk RNAseq and proteomic analysis in (i, j). Two-sided p values were calculated for all Wilcoxon signed-rank tests. Correlation analysis was carried out using Spearman’s correlation. All boxplots (a, j, i) represent the 25th and 75th percentiles with the median marked with whiskers at ±1.5*IQR.
We then investigated individual genes driving the cytotoxicity signature at 3–4 years, and found that the expression of GZMA is significantly upregulated at 3–4 years (Fig. 6c, p = 0.011 and p = 7.5 × 10−8 for S751–765– and S866–880-specific cells, respectively), whereas other cytotoxic molecules, such as GZMB (Fig. 6d, p > 0.05) and PRF1 (Fig. 6e, p > 0.05), showed no significant differences. This suggests that the cytotoxicity signature at 3–4 years is primarily driven by GZMA. Furthermore, at 3–4 years post infection, a greater proportion of cells in the cytotoxic cluster were found in S166–180-specific T cells (CTL, χ2 = 20.007, p = 4.52 × 10−5 [Fig. 6f]), with 11.02% of cells being cytotoxic T cells, compared to 3.4% for S751–765 and 5.68% for S866–880. Amongst these cells, S166–180-specific CD4+ T cells express the highest level of GZMA at 3–4 years, compared to S751–765– and S866–880-specific cells (Fig. 6g, adjusted p = 1.98 × 10−6 and 2.64 × 10−6 for S166–180 vs S751–765 and S166–180 vs S866–880, respectively). Given that our dataset contained three times as many S166–180-specific cells as S751–765-/S866–880-specific cells at 3–4 years, we randomly down-sampled the S166–180-specific cells to 300 cells (matching the number of S751–765– and S866–880-specific cells), and still observed a higher level of GZMA expression in S166–180-specific cells (Supplementary Fig. 6b).
This TCR-independence and GZMA-driven cytotoxicity was further validated in in vitro T cell clones specific for the three epitopes. We assessed the cytotoxicity of 98 CD4+ T cell clones (33 S166–180– specific CD4+ T cell clones from four individuals, 30 S751–765-specific clones from four individuals and 35 S866–880-specific CD4+ T cell clones from three individuals and all clones derived from 1–3-month samples). Clones were categorised as cytotoxic T cell clones if their killing capacity was >10% (see Methods for further details). Their killing of target cells was confirmed to be MHC class II-dependent by using an HLA-DR blocking antibody (Supplementary Fig. 6c). T cell clones with the same TCR clonotype exhibited distinct cytotoxic capability for all three epitopes, with some clones being cytotoxic and others not, despite the same TCR clonotype (Fig. 6h). Bulk RNAseq and proteomic analysis of three cytotoxic and three non-cytotoxic S866–880-specific CD4+ T cell clones showed the cytotoxic CD4+ clones expressed higher GZMA at both RNA level and protein level, compared to non-cytotoxic clones (Fig. 6i, j, p = 4 × 10−4 and p = 0.032, respectively), with no significant differences in the GZMB expression. Importantly, cytotoxicity of the T cell clones associated with their suppression of SARS-CoV-2 replication (Fig. 6k, ρ = 0.390, p = 7.05 × 10−5), with cytotoxic T cell clones showing an overall greater level of viral suppression compared to non-cytotoxic CD4+ T cell clones (Fig. 6l, p = 0.00906). These results suggest that spike-specific memory CD4+ T cells at 3–4 years after infection retain more of a cytotoxic T cell memory, driven by GZMA, capable of controlling virus replication efficiently.