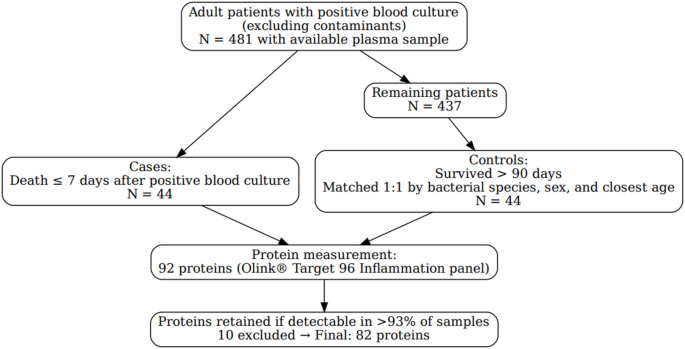
The 44 patients selected as cases in our study represented all patients who died within 7 days after bacteremia in our original bacteremia cohort of 481 patients6. A flowchart illustrating inclusions and exclusions is provided in Fig. 1.
The controls were stratified according to bacterial species, sex, and age and therefore showed no statistical difference between cases and controls (Table 1).
Additionally, there were no statistically significant differences in underlying diseases or infection foci. The severity of the disease (septic shock, use of vasopressors, and admission to the intensive care unit) was significantly worse in the cases than in the controls (P ≤ 0.001).
The protein panel’s internal quality control measurements raised a warning for one case-control pair and one control sample. The corresponding case sample from the latter was also excluded, resulting in the exclusion of two case and two control samples from the analyses. Out of the 92 proteins analyzed using the proximity extension assay, the levels of 82 proteins were further compared between the case and the control samples, whereas 10 proteins (Beta-NGF, IL-33, IL-13, IL-1 A, IL-2, IL-5, IL-4, IL-22RA1, TSLP, and IL-2RB) were excluded from the analyses due to a high number of samples falling below the limit of detection for the corresponding protein.
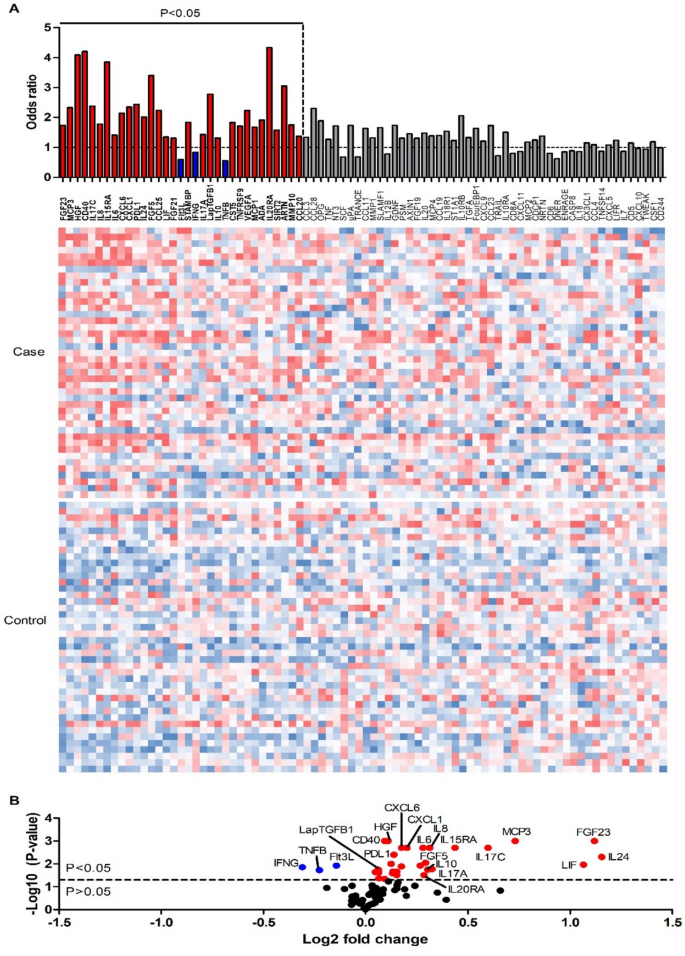
Figure 2 shows the odds ratios, the relative protein levels and the average fold changes together with the P-values, whereas Supplementary Table 2 shows missing relative values, medians, and inter quartile ranges.
Bacteremia cases have an altered inflammatory response in bacteremia. (A) Heatmap of protein expression levels in 84 bacteremia cases, ordered by statistical significance. Columns representing proteins with significantly higher odds ratios in cases compared to controls are marked in red, while those with significantly lower odds ratios are marked in blue. Proteins without statistically significant differences are marked in grey. Red indicates high relative expression, and blue indicates low relative expression for each protein. Color scales have been normalized separately for each protein, with the range adjusted between the lowest and highest expression values. (B) Volcano plot representing the p-values (–log10 scale) and average fold changes (log2 scale) of protein expression levels. Upregulated proteins (red) with the lowest p-values, highest fold changes, and highest odds ratios, as well as all downregulated proteins (blue), are indicated in the figure. The dotted line represents the p-value threshold of 0.05 (approximately 1.30). Note that ARTN is not included in the graph due to a negative average expression value in the control group, which results in a complex number for the log2 fold change.
Of the 82 proteins, 33 had statistically significant odds ratios (ORs) between cases and controls. In majority of these (n = 30), cases had higher expression level than controls (OR > 1). In other words, the high levels of these 30 proteins in plasma were associated with death within 7 days. IL-20RA, CD40, HGF, IL-15RA, FGF-5, and ARTN showed the highest ORs (4.34, 4.21, 4.08, 3.85, 3.40, 3.05, respectively). Conversely, lower abundance of IFNG, Flt3L and TNFB (OR 0.84, 0.59, and 0.56, respectively) were observed in the case group, indicating that reduced levels of these proteins in plasma are associated with poor prognosis in patients with bacteremia. All ORs, 95% confidence interval (CIs), and p-values are shown in Supplementary Table 3. The Benjamini–Hochberg False Discovery Rate (FDR) was also tested (Supplementary Table 4). An FDR threshold of 0.12 corresponded to the same biomarker selection as a p-value threshold of 0.05 in this study.
To explore the signaling pathways associated with prognosis, we next analyzed the differentially expressed proteins between cases and controls (30 up-regulated and 3 down-regulated) using the Panther overrepresentation test15. To exclude possible selection bias caused by the pre-selected protein panel (Olink Target 96 Inflammation panel), we also analyzed the enriched pathways in the background list of proteins (49 analytes) that showed no differences in the expression levels between cases and controls. Additionally, analysis was performed to all proteins (82 analytes) for comparison. We identified 253 biological processes that were enriched within the upregulated list of proteins (Supplementary Fig. 1, Supplementary Table 5). Specifically, we identified several pro-inflammatory cytokine signaling pathways, such as “regulation of interleukin-6 production” (Gene Ontology ID (GO):0045408, P = 2.69 × 10− 6), “regulation of interleukin-23 production” (GO:0045396, P = 9.18 × 10− 5) and “positive regulation of interleukin-17 production” (GO:0032740, P = 7.05 × 10− 4). The downregulated proteins (IFNG, Flt3L, TNFB) did not reveal statistically significant pathway enrichment. We also verified the pathways obtained with the upregulated proteins using Reactome PathwayBrowser (v. 3.7.) (Supplementary Table 6). While the analysis identified some previously described pathways in the target group (e.g. “Interleukin-6 family signaling”), background group (e.g. “Interleukin-1 family signaling”) and target/background group (e.g. “Interleukin-10 signaling”), certain pathways such as “regulation of IL-17 production” and “regulation of IL-23 production” were not found, indicating that the pathway analyses do not have complete overlap.
To identify the key analytes within the groups (highest/lowest ORs, IL-17/Th17 cytokines), we next performed logistic regression analyses. In the proteins with highest odds ratio HGF remained statistically significant (OR 4.11, 95% CI 1.72–9.79, P = 0.001). When the logistic regression was done on the differentially expressed IL-17/Th17 associated proteins (IL-17 C, IL-17 A, LAPTGFbeta1, IL-8, IL-6, and CXCL-1), only IL-17 C remained statistically significant (OR 2.38, 95% CI 1.38–4.11, P = 0.002). Finally, in the logistic regression analysis of the downregulated proteins, only Flt3L remained statistically significant (OR 0.59, 95% CI 0.39–0.89, P = 0.012).