Extreme clonal expansion and inbreeding in a region under massive drug selection
A total of 5,014 dried blood spot (DBS) samples with geographic references from 413 MPs were collected between November 2015 and August 2020 as part of the METF malaria elimination programme in eastern Myanmar, led by the SMRU (Fig. 1 and Extended Data Fig. 1). We processed and analysed genome-wide sequencing data of 2,270 DBS samples collected from 283 MPs (Supplementary Table 1). After filtration of low-coverage samples and low-quality genotypes, the final Kayin State dataset contains 1,927 P. falciparum samples with a set of 25,461 high-quality single-nucleotide polymorphisms (SNPs). Collectively, 92.4% (1,781/1,927) of the samples were from single-genotype infections (Fws ≥ 0.95; Extended Data Fig. 2 and Supplementary Table 2).
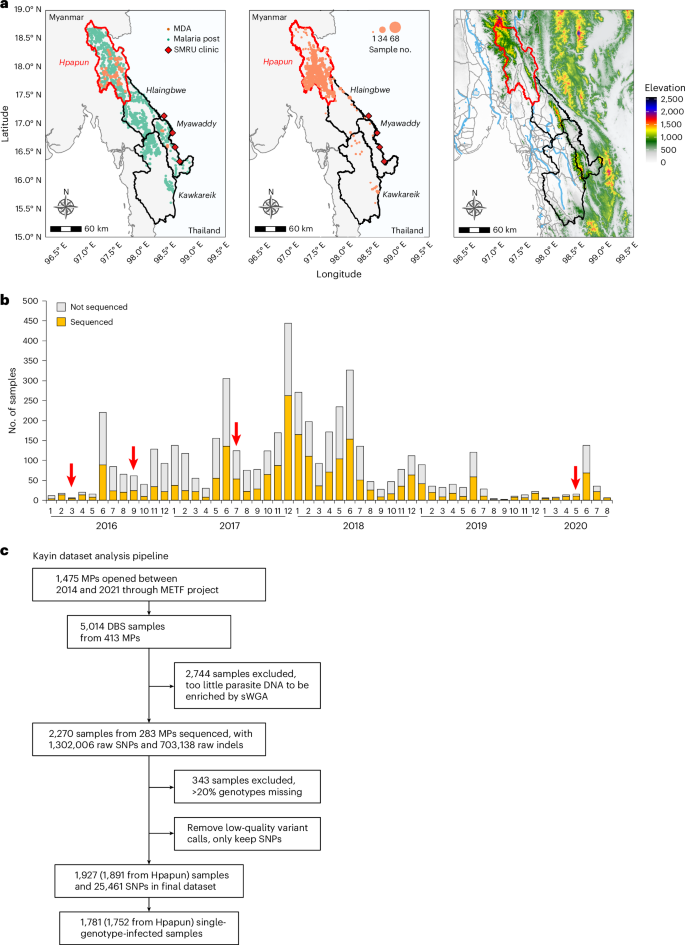
a, Physical geography of the METF intervention region. The METF project was performed at four townships (Myawaddy, Kawkareik, Hlaingbwe and Hpapun) of Kayin State, Myanmar. Left: the distribution of MPs (green dots). Vermillion dots indicate locations where MDAs were applied. Middle: location of sequenced samples. Over 96% of the sequenced sample were from Hpapun township, the northern part of Kayin State. Right: elevation map. Elevation data were downloaded from the US Geological Survey (https://earthexplorer.usgs.gov/). The locations of roads (grey) and rivers (light blue) were obtained from Myanmar Information Management Unit (https://themimu.info/). b, Temporal distribution of samples collected through the METF project. Red arrows indicate the time when MDAs were applied. c, Analysis pipeline for samples collected from Kayin State.
Pairwise genetic relatedness (r, proportion of genome that was identity by descent (IBD)) was used to identify highly related parasites, and samples sharing ≥90% genome IBD (r ≥ 0.90) were grouped into clonal clusters (Fig. 2a and Extended Data Fig. 3). From the 1,781 single-genotype Kayin samples, we identified 93 IBD clusters (2–229 samples per cluster) and 73 singletons, giving 166 unique genomes. We then calculated genetic richness (RG, number of unique genomes divided by the total number of parasites) to quantify the proportion of genetically distinct parasites within each population. The Kayin population exhibited very low RG values (0.10–0.30 across 2015–2020; 166 unique genomes among 1,781 single-genotype infections), reflecting extensive clonal expansion, including 14 large clonal clusters (>30 samples each; Fig. 2a). RG values have declined over time in SE Asian parasite populations (Fig. 2b and Supplementary Tables 3 and 4). The Kayin population showed comparable RG to population samples collected during the same time period (2015–2020) from Vietnam and Laos but were lower than those observed in nearby SMRU clinics (2001–2014) or in other Myanmar regions (excluding Kayin, 2007–2018). For comparison, African P. falciparum populations showed consistently high RG values (0.54–1.00) (Fig. 2b).
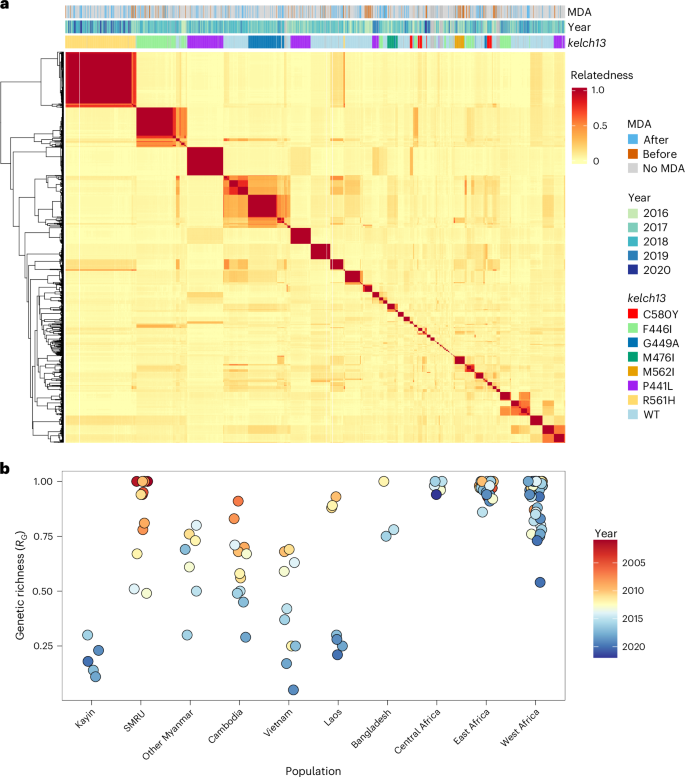
a, Heat map showing relatedness among Kayin samples. Pairwise parasite relatedness (r) was measured as the proportion of the genome with shared IBD between pairs of samples. Samples with r ≥ 0.9 are considered as IBD and share the same unique genome. Colour bars at the top of the heat map indicate information for each sample: MDA, if the sample came from an MP with (orange) or without (blue) MDA; year, the year of sampling; and kelch13, the genotype of kelch13—only alleles with frequency >2% in at least one sampling year were coloured. b, Level of clonal transmission. Genetic richness (RG) was defined as the number of unique parasite genotypes divided by the total number of parasite genotypes.
The ten largest IBD clusters account for 51.6% of the population, indicating low parasite diversity in Kayin State. In total, 151 out of 166 (91.0%) unique genomes were genetically related (r ≥ 0.25) to at least one other genome, indicating high levels of inbreeding (Fig. 3a). Nine closely related families account for 110/166 unique genomes and 66.9% (1,192/1,781) of single-genotype infections, with members sharing r ≥ 0.45. Analysis of chromosomal IBD segments revealed genealogical relationships within families (Fig. 3b and Extended Data Fig. 4). For two families (family 1 and 7), both parents and F1 progeny were identified, indicating an extremely small parasite population size in Kayin State.
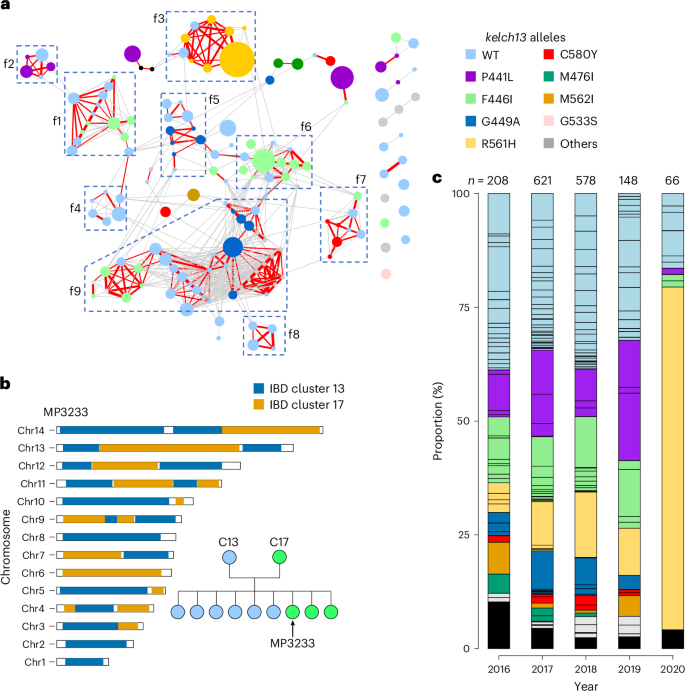
a, IBD network of unique genomes from Kayin population. Nodes: each circle indicates one unique genome and is colour-coded on the basis of its kelch13 alleles; circle size indicates sample size (ranged from 1 to 229). Edges: connections with relatedness (r) ≥0.25; thicker lines indicate higher relatedness; red lines are connections with r ≥ 0.45. Parasites from closely related families (f1 to f9) are labelled using boxes. In total, 152 of 166 unique genomes were included in the network, representing 98.6% of single-genotype-infected samples. b, Pedigree tree of parasites from f1 and chromosome plot for an estimated progeny (MP3233). See Extended Data Fig. 7 for chromosome plots for all progeny. We infer that the parents of f1 are C13 (IBD cluster 13, kelch13-WT) and C17 (IBD cluster 17, kelch13-F446I). c, Proportion of unique genomes across time. Each segment within a bar represents one unique genome, which is coloured on the basis of its kelch13 allele. Black blocks indicate number of unique genomes that were recovered only once (‘singletons’). A clonal expansion of IBD cluster 1 (kelch13-R561H) parasites was detected in 2020.
Localized transmission and regional stability of haplotypes
Patterns of genetic relatedness across space reveal localized transmission dynamics of malaria parasites. Spatial groups of IBD clusters reflect direct or indirect transmission chains and show clear geographic structure (Fig. 4a and Supplementary Fig. 1). For example, 86% of samples from the largest IBD cluster (carrying kelch13 R561H) were collected to the north of Hpapun Township (where >96% sequenced samples were collected; Fig. 1a). The second-largest IBD cluster (carrying kelch13 F446I) was found mainly in the centre of Hpapun Township, while the third-largest IBD cluster (carrying kelch13 P441L) was found in the west of the same township. Similarly, different IBD clusters carrying kelch13 wild type (WT) alleles show localized distribution. Spatial correlograms confirm that parasite relatedness is positively correlated at distances ≤20 km (Fig. 4b), consistent with local transmission of IBD clusters. We also observed significant negative correlations in relatedness between 27.5 km and 90 km, reflecting boundaries between distinct genetic clusters or transmission foci. The negative correlations further indicate the spatial pattern of parasite relatedness. Not only are parasites that are geographically close more likely to be related—those that are geographically distant (27.5–90 km) are less likely to be related.
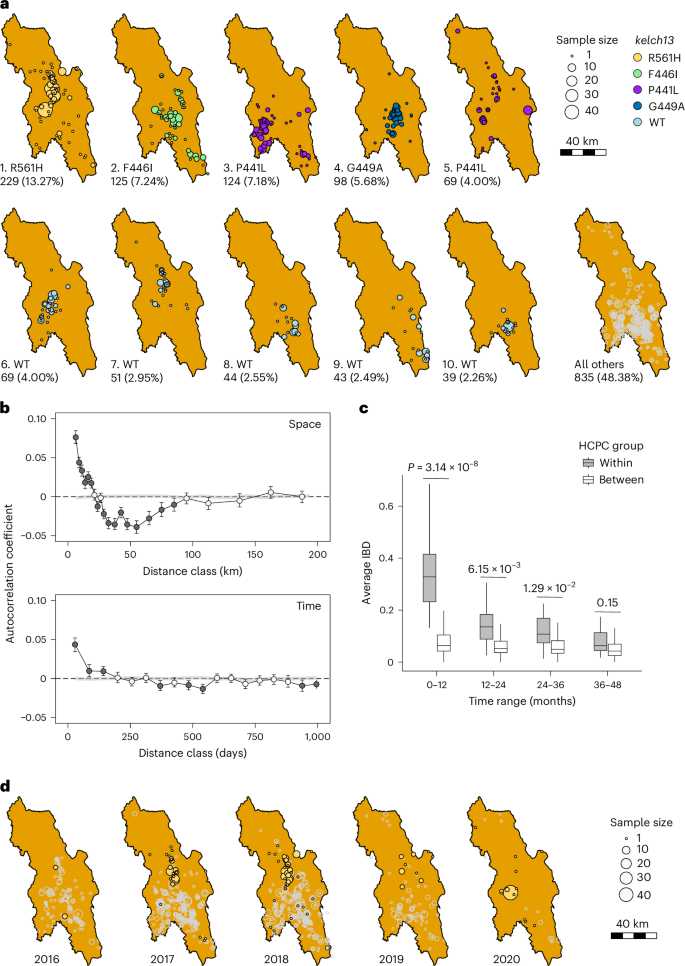
a, Spatial distribution of different IBD clusters. Each circle represents samples collected from an individual MP. IBD cluster IDs and corresponding kelch13 alleles are indicated at the bottom left of each panel; for example, 1_R561H indicates IBD cluster number 1 carrying the kelch13 R561H allele. b, Correlogram analysis of pairwise parasite relatedness across space and time. A total of 1,718 samples with available location and collection time were included in the Mantel correlogram analysis. Bars indicate 95% confidence intervals from 500 bootstrap replicates. c, Comparison of relatedness between within-group and between-group HCPC region pairs. The x axis indicates time intervals in months. Relatedness was measured as average IBD among samples within each HCPC group or between pairs of the 29 groups (more than 10 samples per group). Boxes show interquartile ranges, centre lines indicate medians, whiskers extend to 1.5× the interquartile range, and outliers are not shown. P values were calculated using a two-sided Welch’s two-sample t-test; no adjustment for multiple comparisons was applied. d, Temporal dynamics and spatial distribution of the IBD cluster 1 (kelch13 R561H) through the sampling year.
Temporal patterns of genetic relatedness provide insight into the frequency of outbreeding within malaria parasite populations. We detected clonal IBD clusters (r ≥ 0.90, contain two or more samples) that were sampled across the 56-month study period, as well as new genome haplotypes generated through recombination (Extended Data Fig. 5). Of 93 IBD clusters, 9 lasted ≥36 months, while 34 lasted ≤6 months; the mean cluster duration in the Kayin population was 13.8 (s.e.m, 1.4) months. The mean duration of closely related parasite family members (r ≥ 0.45) was 48.7 (s.e.m, 3.0) months, consistent with limited outbreeding. In two families where both parents and progeny were identified, there were only 12 recombination events among the 109 samples (14 unique genomes, 41 months of family duration) in family (f)1 and 6 among 31 samples (8 unique genomes, 48 months of family duration) in f7 (Extended Data Fig. 4), further supporting with low parasite outbreeding frequency.
We used a temporal correlogram to examine the number of days over which correlations in relatedness were observed (Fig. 4b). This revealed positive correlations in relatedness between parasites sampled ≤170 days apart. To evaluate the relative impact of space and time on parasite relatedness, we divided the control region into 50 regions using hierarchical clustering on principal components (HCPC; Extended Data Fig. 6); 29 of these HCPC regions contained 10–161 parasites. We examined relatedness in parasites sampled within and between HCPC regions for parasites sampled 1–12, 13–24, 25–36 and 37–48 months apart (Fig. 4c). We observed significantly greater relatedness among parasites sampled from the same HCPC unit, relative to those sampled from different HCPC units. This remained significant for parasites sampled up to ≤36 months apart, demonstrating spatial stability of parasite populations within Hpapun Township.
Long-distance connectivity of parasite populations in SE Asia
We used parasite relatedness to measure connectivity within west SE Asian populations and between west and east SE Asia. Genetic similarity and population structure revealed two subpopulations: west SE Asia (Kayin State, SMRU clinics and other Myanmar regions) and east SE Asia (Cambodia, Vietnam and Laos) (Extended Data Fig. 7 and Supplementary Fig. 2). Evidence of gene flow was detected between SMRU clinics and Kayin State: 38.3% of Kayin samples had ≥25% IBD (r ≥ 0.25) with at least one sample from SMRU clinics and 18.1% had ≥35% genome IBD (r ≥ 0.35).
We found no evidence for clonal transmission or recent recombination (r ≥ 0.15) between west and east SE Asia. The low connectivity is further supported by the distribution of pfcrt mutations conferring PPQ resistance. Collectively, 51.21% of east SE Asian (Cambodia, Vietnam and Laos) parasites carried PPQ-resistant pfcrt alleles between 2015 and 2020 (MalariaGEN Pf8 (ref. 10)). These included T93S (22.91%), I218F (13.97%)11,12, H97Y (3.38%), F145I (5.91%), G353V (1.33%)13 and G367C (3.70%)14. By contrast, these pfcrt mutations were absent from the Kayin dataset (Supplementary Table 5) and from other west SE Asian regions (other Myanmar regions and west Thailand).
Genomic measures of parasite population size
We evaluated three genetic metrics—proportion of multiple-genotype infections, RG and effective population size (Ne) in both Kayin State and SMRU parasite populations, for assessing how control efforts impact parasite population size (Fig. 5). Malaria incidence declined sharply in both regions: from 273.9 to 2.2 cases per 1,000 person-years at SMRU clinics (2001–2014) and from 58.8 to 5.3 cases per 1,000 person-years in Hpapun Township, northern Kayin State (2016–2020), where >96% of sequenced samples originated (Supplementary Tables 4 and 6). To facilitate direct comparison with Kayin, we stratified the SMRU dataset into high-transmission (2001–2008) and low-transmission (2009–2014) periods using an incidence threshold of 100 cases per 1,000 population per year15.
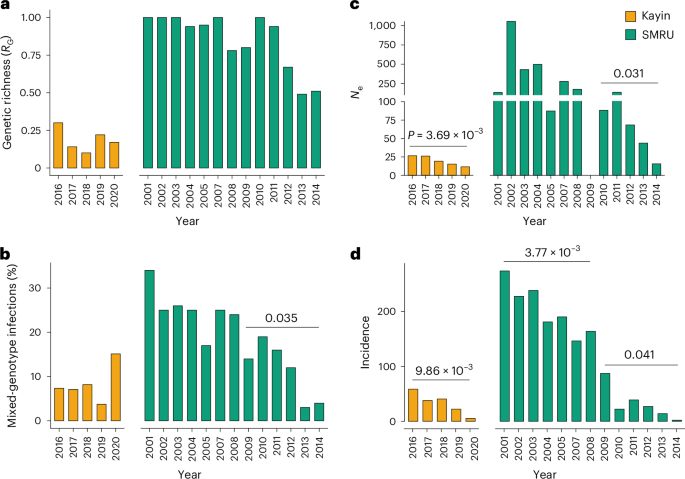
a–c, Population size was estimated using genetic richness (RG) (a), the proportion of samples from mixed-genotype infections (b) and effective population size (Ne) (c). d, Malaria incidence. For Kayin State, data were from Landier et al.8 and Legendre et al.9; the incidence data for SMRU clinics (regions near Mae Sot) was from Nkhoma et al.16. Temporal trends were assessed using linear regression against year; P values are shown only for statistically significant trends; no adjustment for multiple comparisons was applied.
The proportion of multiple genotype infections (7.6%, 146/1927) in Kayin samples was low compared with other P. falciparum populations and remained low throughout the year (range, 3.7–15.1%; Supplementary Table 6). By contrast, SMRU clinics showed a marked decline in multiple genotype infections, from 34.3% in 2001 to 4.3% in 2014, consistent with previous findings16. This decline was not significant during the high-transmission period before 2008 (linear regression, P = 0.23, R2 = 0.27) but was significant during the low-transmission period (P = 0.04, R2 = 0.71). No significant temporal change was observed in Kayin (P = 0.61, R2 = 0.10).
The RG ratio is expected to be negatively correlated with the level of clonal expansion and positively correlated with transmission intensity16. RG ratios were consistently lower in Kayin (0.10–0.30; 2016–2020) than in the SMRU clinics (0.50–1.00; 2001–2014). In SMRU, RG was higher in 2001–2008 (high transmission) than in 2002–2014 (low transmission) (two-sided Welch’s two-sample t-test, P = 0.03). However, RG was not correlated with sampling year in either low- or high-transmission time periods. In Kayin, RG showed little temporal change (P = 0.61, R2 = 0.10).
We calculated single-sample estimates of Ne using unique genomes from each population for each sampling year. Ne estimates were lower in Kayin (11.5–26.6) compared with SMRU clinics (15.5 to infinite). We detected significant reductions in Ne in both Kayin (P = 3.69 × 10−3, R2 = 0.96) and SMRU clinics (P = 0.03, R2 = 0.94) when transmission was low (
The evolution of pfkelch13 alleles
Patterns of drug resistance provide insight into the impact of malaria elimination efforts in Kayin State. Overall, 61.32% of samples from Kayin State carried non-synonymous SNP mutations in kelch13, with P441L (15.19%), F446I (15.01%), R561H (14.02%) and G449A (7.87%) being the most common alleles; only 2.28% carried C580Y (Extended Data Fig. 8). By comparison, at adjacent SMRU clinics, C580Y was the dominant allele, reaching 71.05% frequency in 2014 (Supplementary Table 3). Few samples were collected from SMRU after 2014 owing to the extremely low malaria incidence, limiting temporally matched comparisons. There were 47 IBD clusters (r ≥ 0.90) carrying mutant kelch13 alleles and 46 IBD clusters carrying WT kelch13 in the Kayin samples. Cluster sizes did not differ significantly between mutant and WT kelch13 IBD groups (Extended Data Fig. 9). These results suggest that ART selection was not the main driver for clonal expansion in Kayin State.
Clonal expansion of parasite carrying kelch13-R561H was observed in 2020 (Fig. 3c). Despite strong drug selection, frequencies of mutant kelch13 alleles remained stable between 2016 and 2019 (ref. 6) (Extended Data Fig. 8). However, in 2020, one of the kelch13 alleles—R561H—reached 74.2%. This clonal expansion results from near elimination of parasites from most areas in Kayin, except for northern Hpapun Township where parasites carrying kelch13-R561H predominate (Fig. 4d). In total, 54.8% (40/73) of samples collected between January and August 2020 before the coronavirus disease 2019 pandemic lockdowns were from one single MP (LH-0266B) (Supplementary Table 1).
The changes in kelch13 allele frequencies were reflected by changes in diversity in this gene and its flanking regions. Expected heterozygosity (He) in kelch13 remained high between 2016 and 2019 (He = 0.78) but declined to 0.41 in 2020, with parallel reductions in flanking-region diversity from 0.48 to 0.19 (Extended Data Fig. 8). The sharp decline probably reflects the combined impact of MDA and intensive case management, which together drove a major reduction in malaria transmission. By 2020, infections were detected only in a few remaining villages, indicating a dramatic reduction in parasite population size. This severe bottleneck, and the corresponding drop in Ne, probably led to the marked loss of kelch13 allelic diversity.
Haplotype reconstruction around kelch13 (±100 kb) revealed diverse genetic backgrounds underlying resistance alleles (Extended Data Fig. 10 and Supplementary Fig. 3). Two P441L, one F446I, one G449A and one R561H haplotypes had shared ancestry between Kayin State and SMRU clinics or other Myanmar regions. However, none of these alleles had a high frequency in SMRU clinics or other Myanmar regions. Two F446I and one C580Y haplotypes were uniquely observed in Kayin State, consistent with local origin. For two Kelch13 resistance alleles (G449A and R561H), the WT kelch13 haplotypes on which these resistance mutations arose were sampled in the early 2000s in SMRU clinics. Of the three C580Y haplotypes identified in Kayin, two were also found in SMRU clinics, while one was unique to Kayin State. The two C580Y haplotypes shared with SMRU clinics were only found to the south of Hpapun Township and 60–120 km north of the SMRU clinics (Supplementary Fig. 4). Malaria incidence was extremely low in the SMRU catchment (12 samples sequenced) between 2015 and 2017; hence, direct comparison of contemporaneous parasite genomes from SMRU and Kayin was not possible. None of these haplotypes shared IBD with east SE Asian C580Y haplotypes (Supplementary Fig. 5).
Impact of mass drug use on parasite population structure
We predicted that MDA would reduce relatedness between pre- and post-MDA parasite populations, by clearing local infections and enabling repopulation by new parasite genotypes, leading to founder effects. There were three HCPC units in which sufficient parasites (n ≥ 20) were sequenced both before and after MDA (Fig. 6a). The relatedness between pre- and post-MDA parasites from these three HCPC units was significantly lower than observed between malaria parasites collected during the sampling time period from HCPC regions where MDA was not used (Fig. 6b). Hence, MDA impacted parasite relatedness, consistent with efficient control of pre-MDA parasite genotypes and post-MDA recolonization with unrelated genotypes.
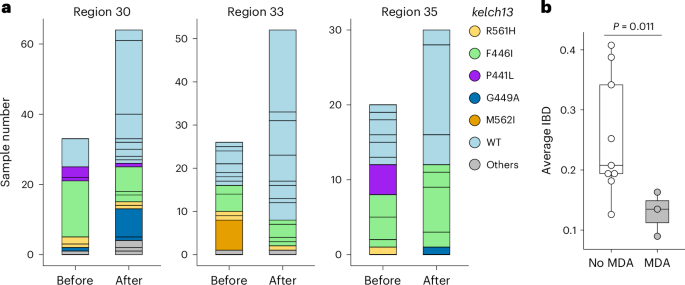
a, Population relatedness before and after MDA. MDAs were implemented between 24 June 2017 and 12 October 2017 in HCPC regions 30, 33 and 35, using DHA–PPQ plus a single dose of primaquine administered monthly for three consecutive months. Bar segments represent unique genomes identified within the 6 months before (12 April 2017 to 12 October 2017) or after the intervention (13 October 2017 to 12 April 2018), colour-coded by kelch13 allele type. b, Comparison of parasite genetic relatedness between HCPC regions with (n = 3) and without (n = 9) MDA intervention. Parasites collected from the same time windows from HCPC regions where MDA was not used provided ‘no MDA’ controls. P values were calculated using a two-sided Welch’s two-sample t-test; no adjustment for multiple comparisons was applied.
We further assessed drug-resistance markers before and after MDA (Supplementary Table 5). No pfcrt SNPs associated with PPQ resistance were detected. Copy number variation (CNV) analysis of plasmepsin II/III (pm2/3) identified three samples with fold-change >1.5 among 281 with sufficient depth; however, manual inspection in Integrative Genomics Viewer showed no clear CNV boundaries, suggesting that these signals were artefacts of selective whole-genome amplification (sWGA). Thus, we found no evidence of pm2/3 amplification. Analysis of mutations in aat1, mdr1, dhfr, dhps and gch1, as well as CNVs in mdr1, revealed no signatures of MDA-associated selection. Genome-wide IBD-based selection scans similarly showed no strong signals of recent selection (Supplementary Fig. 6).






