Architecture
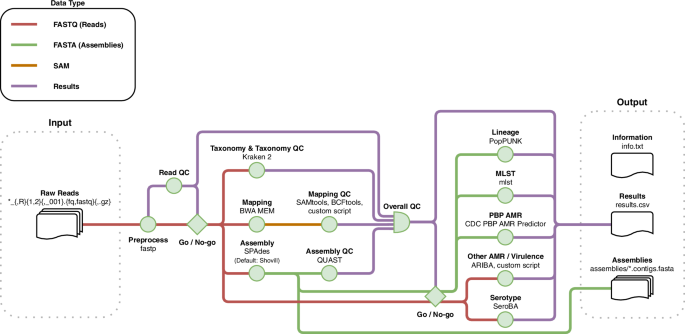
To ensure portability, reproducibility, and easy deployment, the GPS Pipeline (Fig. 4 for schematic overview; Supplementary Fig. 1 for technical flowchart) was designed and implemented in Nextflow v24.10.340 within containerised environments deployed by Docker41 as default or by Singularity42 as an alternative option. The pipeline has a single-entry point for inputting the directory containing raw paired-end short read FASTQ files by Illumina paired-end short read sequencing (Illumina, San Diego, CA, USA) of pneumococcal genomes, then the FASTQ files are automatically processed by multiple Nextflow processes in parallel. The pipeline output includes (1) a single CSV file named results.csv containing all quality control (QC) and in silico typing results, including in silico serotypes, GPSC, MLST, and antimicrobial susceptibilities to 19 common antibiotics in MIC value with CLSI guideline interpretation, (2) de novo assemblies in the format of FASTA files, and (3) a text file named info.txt that includes runtime information and software versions.
Once the user starts the GPS Pipeline with a directory path provided, the pipeline will capture all samples which raw read files match the glob pattern in that specific directory. All captured samples are processed by several bioinformatics tools and have their quality assessed by the quality control (QC) processes, which jointly determine their overall quality based on the metrics of read, taxonomy, mapping and assembly. For samples that have passed the quality control, they are subjected to further analysis by the in silico typing processes, which assign Global Pneumococcal Sequence Cluster (GPSC) lineage, and predict serotype, multilocus sequence typing (MLST) and a wide range of antimicrobial resistance (AMR) and virulence targets. The results for all samples are collated into a single comma-separated values (CSV) file. The pipeline saves the runtime information and version of the software for reproducibility, and the generated assembly of each sample.
The pipeline can be run without requiring installation or administrative privileges on any POSIX-compatible system, provided that Bash 3.2 or above, Java 11 or above, and either Docker or Singularity are pre-installed. Compatible operating systems include Linux, Windows (through WSL), and macOS. It is also compatible with LSF-based HPC clusters and the Seqera Platform (previously known as Nextflow Tower). We tested the GPS Pipeline on Linux (Ubuntu 22.04), Windows (Windows 11 with Ubuntu 22.04 through WSL2), and macOS (Sonoma 14.4) machines with ≥ 16 GB RAM, as well as LSF-based HPC clusters (Wellcome Sanger Institute Farm5 and Farm22 HPC clusters), and generated consistent results. After downloading the pipeline and running its initialisation function, which downloads all required additional files and container images, the pipeline can be used without an internet connection.
The pipeline logic of the GPS Pipeline is programmed in Nextflow Domain-Specific Language 2 (DSL2) and supplemented by custom Apache Groovy functions. The pipeline logic invokes bioinformatics tools for analyses, and executes Shell or Python 3 scripts for data and metadata manipulations.
In this pipeline, we selected bioinformatics tools that are automated, reliable, fast, memory-efficient, and output results in a standardised text-based format that can be parsed programmatically. When possible, we selected existing bioinformatics tools that met these criteria and built custom scripts when necessary.
Quality control
The pipeline starts with a series of quality control checks on the FASTQ files and genomes based on the configurable criteria listed in Supplementary Table 1. The pipeline first checks for corrupted FASTQ files (files that cannot be decompressed or contain incomplete data). Samples with any corrupted files are not further processed. An error message to indicate which file is corrupted is shown in the result file (results.csv).
Intact paired-end FASTQ files are then assessed by fastp v0.23.443. fastp automatically performs adaptor trimming, quality filtering, and other operations to clean up the FASTQ files for downstream processing, as well as acquiring the total base count of the read files. To pass the QC, the total base count should exceed the multiplication of the minimum sequence depth and lower assembly length limit, which is 38 Mbps. Pre-processed samples that pass QC are then processed in parallel through taxonomy, mapping and de novo assembly processes as stated below for further quality control based on these metrics.
To detect any potential contamination, the pipeline runs Kraken 2 v2.1.244 on the FASTQ file pairs against the Minikraken v1 database45 as the default for taxonomy classification. Kraken 2 runs faster and requires less memory when compared to Kraken 1, and it uses a capped-size database available46, enabling it to run on the usually limited system resources available on personal computers (PCs). In contrast, alternatives like GTDB-Tk with its Genome Taxonomy Database (GTDB)47, have memory requirement exceeding most PCs capability (Supplementary Table 2). A QC-passed genome should have ≥ 60% reads mapped to S. pneumoniae while ≤ 2% reads mapped to non-pneumococci species.
To ensure the sequencing completeness and to detect a mixture of two or more pneumococcal isolates in a single sample, the FASTQ file pairs are mapped to the reference genome S. pneumoniae ATCC 700669 (NCBI accession no. FM211187) by default, the standard reference since the beginning of the GPS Project, using the BWA-MEM algorithm of BWA v0.7.1748. Its successor, BWA-MEM2, was not used due to its much higher memory requirement, rendering it unsuitable to run on PCs49. The output SAM files are converted into BAM files and sorted using SAMTools v1.1650, and the reference coverage percentage is then calculated. The sorted BAM files are used to call the SNP sites, which are saved into VCF files using BCFTools v1.1650. Het-SNP sites are then counted by a custom Python script (bin/het_snp_count.py). This script filters out het-SNPs that are within 50-bp proximity to avoid overestimating the het-SNPs. In this step, a QC-passed genome is expected to have a coverage of the reference genome ≥ 60% and a count of het-SNP ≤ 220. The former threshold ensures sequence reads of the sample have sufficient coverage over the genome, while avoiding being over-aggressive to account for the highly diverse genome of the pneumococcus. The later threshold confirms isolates having an acceptable purity, which were established based on analysed data across 29,913 pneumococcal genomes in the GPS project (Supplementary Fig. 2).
To assess the assembled genome quality based on sequencing depth, length and number of contigs, de novo assembly is carried out using Shovill v1.1.0 by default on the FASTQ file pairs. The quality metrics are summarised by QUAST v5.0.251. QC-passed genomes are expected to have a sequence depth of ≥ 20x, a length of assembly between 1.9−2.3 Mbps, and a number of contigs ≤ 500.
At the end of each QC check above, a Shell script (bin/validate_file.sh for FASTQ files integrity, bin/get_read_qc.sh for basic read quality, bin/get_taxonomy_qc.sh for taxonomy-based purity check, bin/get_mapping_qc.sh for mapping-based coverage and purity check, bin/get_assembly_qc.sh for assembly quality) assigns QC pass/fail category based on the QC metrics. Another Shell script, bin/get_overall_qc.sh, then assigns genomes that have passed all QC checks to the overall QC pass category. QC-failed genomes do not proceed downstream for in silico typing.
De novo assembly
Shovill v1.1.0, a de novo assembler with automated optimisations, is set as the default assembler, with the included option of Unicycler v0.5.052 to be used by the pipeline as an alternative assembler, in case Shovill fails to generate optimal assemblies for some samples. Both assemblers are based on SPAdes. A third, Velvet-based assembler VelvetOptimiser v2.2.6, was also evaluated, but not included. The decision was based on results from tested reads from different Illumina short-read sequencing platforms (Table 3), carried out using the default parameters of the assemblers, except instructing them to use all available threads on the machine and discarding contigs with less than 500 bps. Shovill was selected as the most appropriate assembler for this pipeline based on it having the shortest runtime, lowest maximum memory usage, and ability to generate high-quality assemblies. Shovill only takes 25.3% of runtime and 70.5% of maximum memory to generate assemblies of similar quality, as compared to Unicycler, based on the number of contigs and N50 metrics, which is more than sufficient for the downstream tools in the pipeline. At the same time, VelvetOptimiser requires 172.7% of runtime compared to Shovill while generating the worst quality assemblies and is a less stable software, making it unsuitable for the pipeline.
In silico typing
Genomic lineage
We assign Global Pneumococcal Sequence Clusters (GPSCs) to define lineages, which are based on genome-wide variations as compared to the conventional multilocus sequence typing (MLST) scheme, which only takes the sequences of 7 housekeeping genes into account14,53. GPSCs assignment is coherent by nature, allowing comparison of results between different studies. To assign GPSCs, PopPUNK v2.6.3 and the latest PopPUNK GPS database v9 (gps-project.cog.sanger.ac.uk/GPS_v9.tar.gz & gps-project.cog.sanger.ac.uk/GPS_v9_external_clusters.csv) are used.
In silico serotyping
For in silico serotype prediction, SeroBA v2.0.4 was selected. It has a low memory and computational power requirement, yet high prediction accuracy and specificity. These advantages are evident when compared to PneumoCat which is another common in silico pneumococcal serotyping tool54. The current pipeline utilises SeroBA database v2.0.4 as it could identify 102 of 107 known pneumococcal serotypes and 4 groups of non-encapsulated pneumococci. As new serotypes are detected, the pipeline can be easily updated to accommodate the latest version of SeroBA and its database.
MLST
mlst v2.23.0 is used to infer MLST profile and sequence type (ST) from assemblies. The current pipeline uses pneumococcal PubMLST data as of 1st July 2024. As new ST are detected, the pipeline can also be updated to include the latest version of the PubMLST database.
Antimicrobial resistance
The GPS Pipeline predicts antimicrobial susceptibility to 19 commonly used antibiotics using the CDC PBP AMR Predictor and ARIBA v2.14.6, and interprets the MIC predictions based on the 2014 CLSI guideline31. The known resistance genes and mutations are summarised in Table 5.
The CDC PBP AMR Predictor assigns PBP transpeptidase domain protein sequence types to the PBP1a, PBP2b, and PBP2x proteins encoded by draft genome assemblies, collectively referred to as PBP type. It then predicts the MICs against 6 β-lactam antibiotics, including: AMO, CFT, TAX, CFX, MER, and PEN. The machine learning model used for the prediction7 was previously shown to yield high percent essential agreement (MICs agree within ± 1 dilution) > 97% and percent category agreement (interpretive results agree) > 93% in a pneumococcal genome dataset that was not used for building the machine learning model55. The resistance category against each antibiotic is also inferred from the predicted MIC according to the CLSI guidelines.
ARIBA v2.14.6 is used with a pneumococcus-specific database, which we compiled to perform the AMR predictions against 13 non-β-lactam antibiotics by detecting the presence or absence of known resistance genes, mobile elements or mutations listed in Table 5. It was chosen because newly recognised resistance genes or mutations can be easily added to the ARIBA database for detection, enabling the pipeline to quickly adapt to newly discovered AMR mechanisms in the future. It should be noted that the nonsense mutations across the nucleotide range 166-201 (amino acid residue 56–67) of folP, which result in resistance to SMX56,57, cannot be readily represented in an ARIBA database. Therefore, ARIBA is only used to detect and extract any mutations in the folP gene, and a subsequently run Python script (bin/parse_other_resistance.py) detects any disruptive mutation within the nucleotide range.
To avoid spurious detection, the presence of a resistance gene is confirmed by the same Python script (bin/parse_other_resistance.py), which determines if the gene meets the criteria of ≥ 80% coverage and ≥ 20x read depth. For a resistance mutation, a ≥ 10x sequence depth threshold is used for confirmation by this script. The categorical prediction of antimicrobial susceptibility (susceptible, intermediate, resistant) against CHL, CLI, COT, DOX, ERY, ERY and CLI, FQ, KAN, LFX, RIF, SMX, TET, TMP, and VAN is then saved to a report. The MIC range based on the categorical prediction is also added, based on the 2014 CLSI guidelines.
Virulence factor
The GPS Pipeline also detects the presence and absence of virulence factors, including pilus genes of PILI-1, and PILI-2, using ARIBA v2.14.6 with the identical criteria as for detecting resistance genes (≥ 80% coverage and ≥ 20x read depth).
Containerisation
To implement the above functions in containerised environments, we included publicly available Docker images (Supplementary Table 3) developed by a communal effort led by the State Public Health Bioinformatics Group (StaPH-B)58 whenever possible. These images are well maintained and tested to be stable, appropriately versioned (which enables version pinning and simple upgrade), and is open source. During the development of the GPS Pipeline, we have also made contributions toward this effort by updating and adding testing stages to the Dockerfiles for the Docker images of PopPUNK and ARIBA, which were promptly reviewed and added to become part of their releases. In addition, we built Docker images of CDC PBP AMR Predictor and SeroBA, as up-to-date images were not available. To handle the extensive usage of command line utilities and Python scripts in the pipeline, the Network-Multi Tool Docker image by WBITT59 and the Pandas Docker image by Alexander Mancevice60 were chosen, respectively.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.