Cohort description
22 PLWH (11 Off-ART and 11 On-ART) were included in the first part of this study. Clinical characteristics are depicted in Table S1. At enrollment, all Off-ART participants had detectable viral loads (VL, median = 5.2log10 HIV RNA copies/mL) and all On-ART participants had undetectable VL. The On-ART group showed significantly higher CD4 counts and CD4/CD8 ratios. Additionally, samples from 31 On-ART participants were obtained for functional assays (Table S2).
GITR is expressed in T-cells from PLWH off and On-ART and correlates with parameters related to viral control
PBMCs were cultured for 3 (Off-ART) or 14 (On-ART) days with Nef or p24 peptide pools (or a cytomegalovirus, Epstein-Barr and influenza virus peptide pool (CEF) as control) and GITR expression was evaluated in bulk CD8TCs, HIV-specific CD8TCs, conventional CD4TCs (Tconv) and Treg (Fig. S1). The longer culture time for the On-ART group was necessary for the enrichment of HIV-specific cells for analysis, based on the notion that the frequency of this population decays significantly following ART initiation.
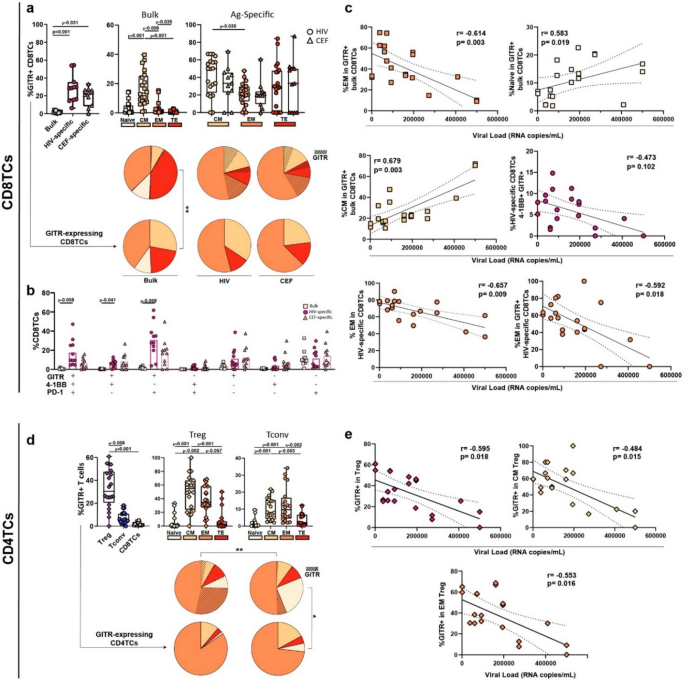
In the Off-ART group, post-stimulation GITR expression within HIV- and CEF-specific CD8TC subpopulations was significantly higher compared to CD8TC bulk compartment (Fig. 1a, upper left). Within bulk CD8TCs, the central memory (CM) subset was the most enriched in GITR (median = 13.50%, IQ25-75 = 7.56–25.48) (Fig. 1a, upper center). The median frequencies of virus-specific CD8TCs were HIV = 0.375 and CEF = 0.13. The mean frequencies of each memory subset within HIV-specific CD8TCs were CM = 20.91, EM = 67.59, TE = 11.48 while within CEF-specific these were CM = 17.15, EM = 72.25, TE = 10.59. Within antigen-specific cells, the percentage of GITR-expressing HIV-specific CM cells was higher than EM cells. No significant differences in GITR expression were found across memory subsets for CEF-specific CD8TCs (Fig. 1a, upper right), nor in global memory distribution between HIV-specific and CEF-specific CD8TCs (Fig. 1a upper pies).
We then analyzed the distribution of memory/effector subpopulations within GITR-expressing CD8TCs (Fig. 1a, lower pies). This distribution was significantly different to the memory distribution of global CD8TCs (upper pie), with a higher contribution of the CM subset. The effector memory (EM) subset dominated in bulk GITR-expressing CD8TCs, reaching a mean = 40.07%, and was even higher in the HIV-specific and CEF-specific compartments (54.01% and 57.79%).
Next, we evaluated the single-expression and coexpression of GITR, PD-1 and 4-1BB (costimulatory receptor, member of TNFRSF). A significantly higher post-stimulation coexpression of GITR+/4-1BB+/PD-1+ was found in HIV-specific CD8TCs compared to bulk. GITR+/4-1BB+ and GITR+/PD-1+ coexpression was also higher in HIV-specific cells than in bulk CD8TCs (Fig. 1b).
Correlation analyses (Fig. 1c and supplementary Table S4) depicted that %EM in bulk GITR + CD8TCs inversely correlated with VL. Conversely, %Naive and %CM in bulk GITR + CD8TCs positively correlated with VL. Within HIV-specific CD8TCs, %GITR+/4-1BB+ cells negatively correlated with VL and %EM in GITR-expressing cells also correlated negatively with VL, although this association could be driven by the overall negative correlation between %EM and VL (Supplementary Table S5, Box 8, 10 and 11). Finally, %GITR in terminal effector (TE) and %TE in GITR + positively correlated with CD4/CD8 and CD4TC counts, respectively (Supplementary Table S5, Box 8 and 12).
In the CD4TC compartment, GITR expression was significantly higher in Treg compared with Tconv and bulk CD8TCs (Fig. 1d left panel). Within Tregs, the CM compartment was the most enriched in GITR (54.55%, IQ25-75 = 23.88–66.70), followed by EM (34.85%, IQ25-75 = 27.23–58.30) (Fig. 1d, center). In Tconv, GITR enrichment was higher in CM (8.09%, IQ25-75 = 4.12–15.43) and EM (7.42%, IQ25-75 = 4.15–14.15) compared with Naïve and TE (Fig. 1d right). The global distribution of subsets was significantly different between Treg and Tconv, with a higher contribution of GITR + EM within Treg (Fig. 1d upper pies). Within GITR-expressing CD4TCs, most Treg and Tconv had an EM phenotype (81.23% and 73.24%) (Fig. 1d, lower pies) and global distribution of bulk Tconv (upper pie) was significantly different compared to GITR-expressing Tconv (lower pie), with a contraction of the Naive subset in the latter.
Correlation analyses depicted negative correlations between VL and %GITR-expressing cells in bulk, CM and EM Tregs (Fig. 1e and supplementary Table S6). No other relevant significant associations were found (Supplementary Figs. S3a, S4a–c).
GITR expression on bulk and HIV-specific T-cells from PLWH Off-ART. (a) Box plots represent the proportion of GITR + events within bulk and antigen-specific (Ag-specific) CD8TCs (left) and within each memory subset (bulk: center and Ag-specific: right). The color code shown at the bottom mirrors the one shown in the pie charts. Upper pie graphs depict the CD8TC memory distribution and contribution of GITR + events on bulk and Ag-specific cells. Lower pie graphs depict the memory distribution of the bulk and Ag-specific GITR + CD8TCs. (b) Bar plot represents the GITR, 4-1BB and PD-1 single and coexpression pattern in bulk, HIV- and CEF-specific CD8TCs. (c) Correlation analysis between GITR expression across memory subsets in bulk/HIV-specific CD8TCs and viral load. (d) Box plots represent the proportion of GITR + events within regulatory (Treg), conventional (Tconv) CD4TCs and bulk CD8TCs (left); and within each memory subset of Treg and Tconv CD4TCs (center and right, respectively). Upper pie graphs depict the CD4TC memory distribution and contribution of GITR + events on Treg and Tconv cells. Lower pie graphs depict the memory distribution of the GITR + Treg and GITR + Tconv. (e) Correlation analyses between GITR expression across different memory subsets in Treg and viral load. CM: central memory, EM: effector memory, TE: terminal effector. N = 11 individuals, 20 HIV responses (10 Nef + 10 p24) and 11 CEF responses. In box plots, boxes extend from Q1 to Q3, horizontal bars within boxes represent median values and whiskers extend from min to max. In bar plots, bar height represents mean. When analyzing differences between CEF and HIV, the Nef and p24 responses were averaged and grouped under the HIV response to obtain the same number of responses as CEF and apply the appropriate statistical test. When analyzing differences within a single response, the full dataset was used. P-values were calculated using the Friedman test for paired samples, followed by Dunn’s post-hoc correction for multiple comparisons. When analyzing global distribution (pies) permutation tests were applied using SPICE software. In all plots, squares represent bulk CD8TCs, circles represent HIV-specific CD8TCs, triangles represent CEF-specific CD8TCs, diamonds represent Treg and hexagons represent Tconv. r and p values were computed by Spearman´s correlation and p-values were adjusted for multiple comparisons using a hypothesis-driven false discovery rate (FDR) procedure, according to the Benjamini and Hochberg method.
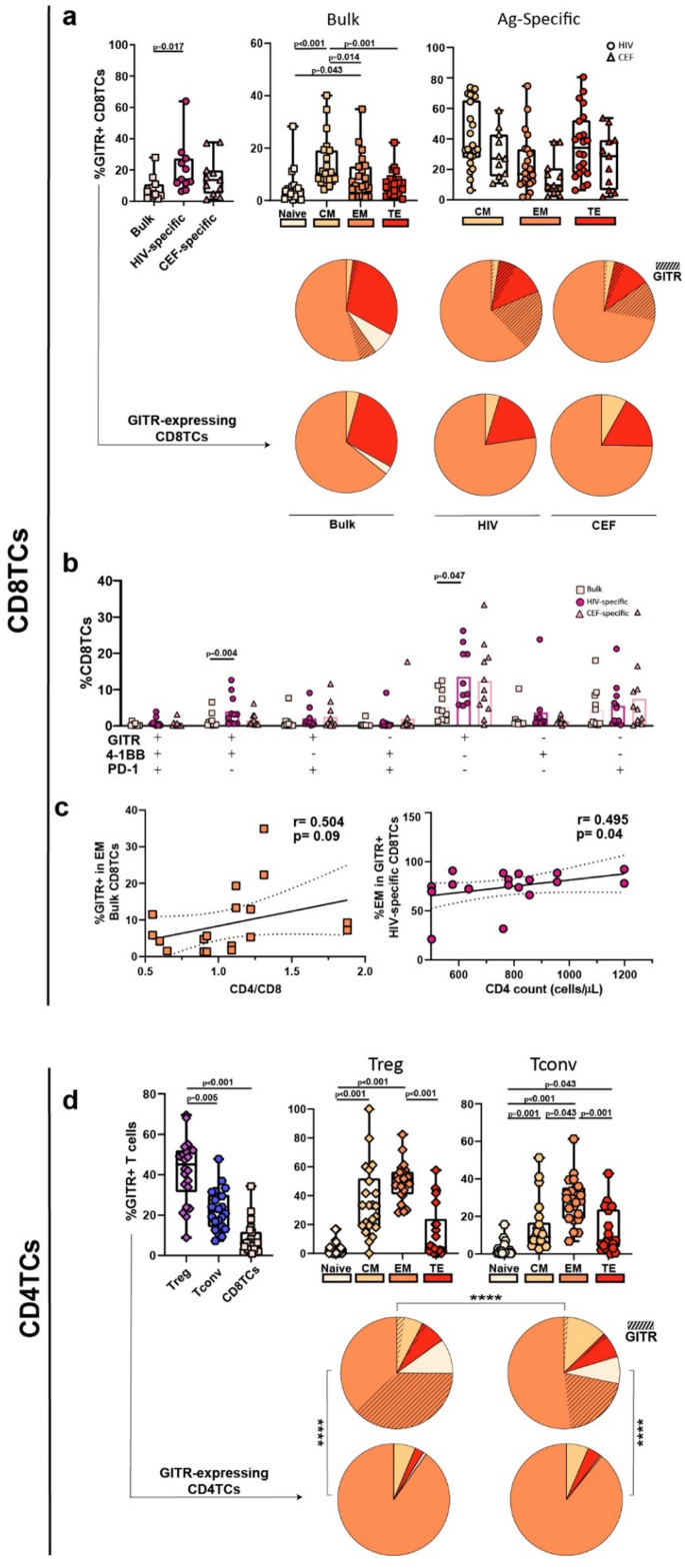
After exploring GITR expression on T-cells from PLWH Off-ART, we extended the analysis to PLWH On-ART. Similar to the Off-ART group, the percentage of GITR + cells was higher among HIV-specific CD8TCs compared with bulk CD8TCs (Fig. 2a upper left). When analyzing GITR expression across memory/effector subsets within bulk CD8TCs, the CM subset showed the greatest GITR enrichment (10.25% IQ25-75 = 7.83–19.18) (Fig. 2a upper center). The median of frequencies of virus-specific CD8TCs were HIV = 19.7, CEF = 9.85. The mean frequencies of each memory subset within HIV-specific CD8TCs were CM = 2.54, EM = 81.35, TE = 16.09, while within CEF-specific these were CM = 3.35, EM = 84.79, TE = 11.86. Within post-stimulated HIV-specific CD8TCs, the TE and CM phenotypes were the most enriched in GITR (34.10% IQ25-75 = 17.38–52.25 and 33.25% IQ25-75 = 27.55–65.28, respectively), followed by EM (16.85% IQ25-75 = 10.63–33.20) (Fig. 2a upper right). The same hierarchy (TE = CM > EM) was observed in CEF-specific CD8TCs. There was no difference between global memory distribution between HIV-specific and CEF-specific CD8TCs (Fig. 2a, upper pie). Among GITR-expressing cells (Fig. 2a, lower pie), the EM phenotype dominated in bulk CD8TCs, with a mean of 71.16%, and was also higher in the HIV- and CEF-specific compartments (74.05% and 75.36%, respectively), with no differences in the global memory distribution between GITR-expressing HIV-specific and CEF-specific cells.
The analysis of the post-stimulation single-expression and coexpression of GITR, 4-1BB and PD-1 revealed a significantly higher percentage of GITR+ and GITR+/4-1BB+ CD8TCs in HIV-specific compartment compared with bulk CD8TCs (Fig. 2b). Correlation analyses showed that the percentage of GITR + cells within EM CD8TCs positively correlated with CD4/CD8 ratios and, within GITR + HIV-specific CD8TCs, the %EM cells directly correlated with CD4TC counts (Fig. 2c; supplementary Tables S7 and S8). No other relevant significant associations were found (Supplementary Table S9, supplementary Figs. S3b and S4d).
On-ART CD4TC analyses also revealed significantly higher GITR expression in Tregs versus Tconv and bulk CD8TCs (Fig. 2d left panel). In both Treg and Tconv, the EM phenotype was the most enriched in GITR (51.50% IQ25-75 = 40.93–56.55 and 28.75% IQ25-75 = 18.75–35.40, respectively), followed by CM (33% IQ25-75 = 18.93–51.98 and 9.03% IQ25-75 = 5.14–16.70, respectively) (Fig. 2d, center and right box charts). Memory global distribution was significantly different between Treg and Tconv, with a higher contribution of GITR + EM within Treg (Fig. 2d upper pies). In both GITR-expressing Tregs and Tconv, most cells (90.14% and 89.18%, respectively) exhibited an EM phenotype (Fig. 2d, lower pies), with significant differences in global memory distribution compared to bulk subsets (upper pies).
GITR expression on bulk and HIV-specific T-cells from PLWH On-ART. (a) Box plots represent the percentage of GITR + events within bulk and antigen-specific (Ag-specific) CD8TCs (left) and within each memory subset (bulk, center and Ag-specific, right). The color code shown at the bottom mirrors the one shown in the pie charts. Upper pie graphs depict the CD8TC memory distribution and contribution of GITR + events on bulk and Ag-specific cells. Lower pie graphs depict the memory distribution of the bulk and Ag-specific GITR + CD8TCs. (b) Bar plots represent the GITR, 4-1BB and PD-1 single and coexpression pattern in bulk, HIV- and CEF-specific CD8TCs. (c) Correlation analysis between percentage of GITR + in EM bulk CD8TCs and %EM in GITR + HIV-specific CD8TCs with CD4/CD8 ratio and CD4TC counts. (d) Box plots represent the proportion of GITR + events within regulatory (Treg), conventional (Tconv) CD4TCs and bulk CD8TCs (left), and within each memory subset of Treg and Tconv CD4TCs (center and right, respectively). Upper pie graphs depict the CD4TC memory distribution and contribution of GITR + events on Treg and Tconv cells. Lower pie graphs depict the memory distribution of the GITR + Treg and GITR + Tconv. CM: central memory, EM: effector memory, TE: terminal effector. N = 11 individuals. For bar charts: 22 HIV responses (11 Nef + 11 p24) and 11 CEF responses. For correlation analyses: 20 HIV responses (10 Nef + 10 p24). In box plots, boxes extend from Q1 to Q3, horizontal bars within boxes represent median values and whiskers extend from min to max. In bar plots, bar height represents mean. When analyzing differences between CEF and HIV, the Nef and p24 responses were averaged and grouped under the HIV response to obtain the same number of responses as CEF and apply the appropriate statistical test. When analyzing differences within a single response, the full dataset was used. P-values were calculated using the Friedman test for paired samples, followed by Dunn’s post-hoc correction for multiple comparisons. When analyzing global distribution (pies) permutation tests were applied using SPICE software. In all plots, squares represent bulk CD8TCs, circles represent HIV-specific CD8TCs, triangles represent CEF-specific CD8TCs, diamonds represent Treg, and hexagons represent Tconv. r and p values were computed by Spearman´s correlation and p-values were adjusted for multiple comparisons using a hypothesis-driven false discovery rate (FDR) procedure, according to the Benjamini and Hochberg method.
Overall, we showed that GITR is expressed in T-cells from PLWH Off- and On-ART. Given the potential role of GITR in HIV control and the fact that most GITR-expressing CD8TCs are EM cells, we explored enhancing antiviral function through a hexameric GITR ligand. Additionally, due to the observed GITR enrichment on CM and EM CD4TCs, we evaluated the HIV latency-reversal properties of GITRL.
GITR expression peaks at 96 h post-stimulation on CD8 and CD4TCs from PLWH On-ART and it is specifically recognized by a hexameric GITR ligand
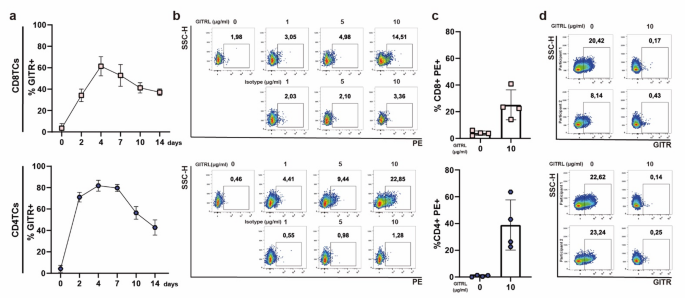
A post-activation kinetic analysis of PBMCs revealed similar GITR expression patterns between CD8 and CD4TCs, with rapid upregulation up to day 4 (Fig. 3a). After establishing GITR peak expression, we assessed the binding capacity of GITRL on CD8 and CD4TCs at that time point. A dose-response relationship between GITRL concentration and fluorescence signal was observed on activated CD8 and CD4TCs (Fig. 3b–c) and an interference assay confirmed GITRL specificity for its receptor (Fig. 3d). A similar binding assay on resting PBMCs in the absence of stimuli for 4 days showed comparable results (Fig. S2a). Finally, using the Flag tag in the GITRL construct, we repeated the binding assay in activated CD4TCs with an anti-Flag mAb M2, observing the same pattern (Fig. S2b).
GITR expression kinetics and GITRL binding on activated CD8 (upper panel) and CD4TCs (lower panel) from PLWH On-ART. (a) GITR kinetic expression. Squares and circles represent the mean percentage of GITR + CD8 and CD4TCs. Whiskers represent the standard deviation. N = 4 participants. (b) GITRL binding on activated CD8 and CD4TCs. Representative flow cytometry dot plot from one participant, showing the proportion of PE + events after GITRL or isotype incubation, developed with αhuman IgG1-biotin + Streptavidin-PE. (c) Pooled data (N = 4 participants) from GITRL binding assay. Percentage of PE + CD4 and CD8TCs determined after T-cell activation and treatment with 0 and 10 ug/mL of GITRL. Bar height represents the mean and whiskers represent the standard deviation. (d) Results from GITR-GITRL interference assay. Flow cytometry dot plots from 2 participants show the percentage of GITR + events within activated CD8 and CD4TCs after 5 h of incubation in the absence (0 µg/mL) or presence (10 µg/mL) of GITRL.
GITRL costimulation enhances IFN-γ production and degranulation in HIV-specific CD8TCs from PLWH On-ART in the short-term, but it has no effect on their long-term cytolytic function
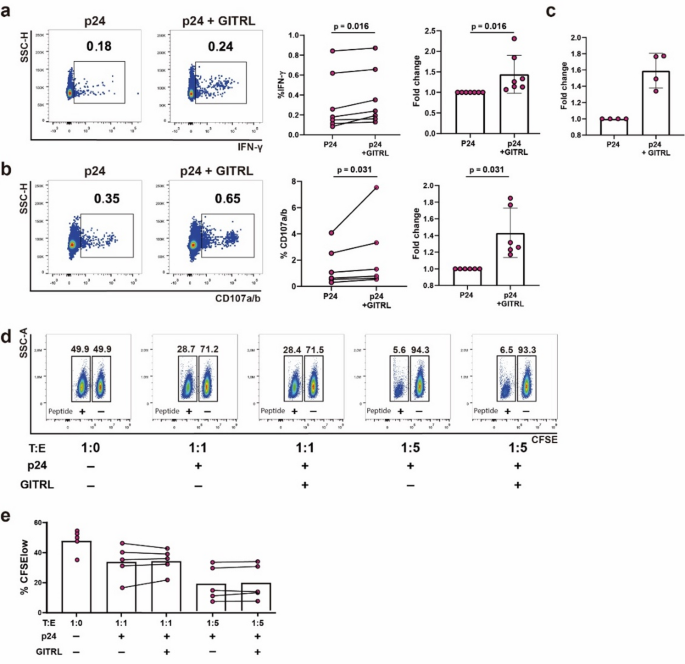
To assess GITRL effect on enhancing HIV-specific responses, PBMCs from 7 On-ART PLWH were treated with a p24 peptide pool plus GITRL for 4 days. After a brief restimulation, the frequency of IFN-γ + and CD107a/b + CD8TCs significantly increased in the GITRL-stimulated condition (Fig. 4a–b). A similar trend was observed for IFN-γ levels measured by ELISA in culture supernatants (Fig. 4c).
To evaluate if this improvement in cytokine production and degranulation had an effect on CD8TC cytolytic capacity, a VITAL killing assay was performed, following our group’s established protocol21. Figure 4d shows flow-cytometry dot-plots obtained in one representative assay, which indicates a specific progressive loss of antigen-pulsed CFSElow target cells as effectors are added at increasing ratios. Treatment of effector cells with GITRL during the expansion with p24 peptides had no effect on the cytotoxic function of CD8TCs in either effector: target condition (Fig. 4e). Moreover, a CFSE-based proliferation assay on PBMCs stimulated with p24 peptides showed no significant difference in the percentage of proliferating CD8TCs between conditions with or without GITRL after 7 days of stimulation (Supplementary Fig. 5).
Effect of GITRL costimulation on HIV-specific CD8TCs from PLWH On-ART. Effect of GITRL costimulation on (a) IFN-γ production and (b) degranulation. Within panels A and B, dot plots depict the results from a representative donor. Scatter plots depict pooled data from 7 donors. Left scatter plot with lines shows percentages of IFN-γ + or CD107a/b + events while right scatter plot with bars shows the fold change between conditions. p values were calculated using Wilcoxon test. (c) Scatter plot with bar graph of pooled data from 4 participants showing the fold change of IFN-γ concentration from supernatants of PBMCs after 4 days of p24 peptide stimulation, in the absence or presence of 10 µg/mL GITRL. In all scatter plots bar height represents mean value and whiskers represent standard deviation. (d) Dot plots obtained in one representative VITAL assay. At the 1:0 target-to-effector (T: E) ratio (far left panel), an even distribution of peptide-loaded (CFSElow) targets and non-loaded targets (CFSEhigh) is observed. From left to right, increasing ratios of effectors CD8TCs (1:1 and 1:5) expanded in the absence or presence of GITRL are shown. (e) Scatter plot with bar graph of pooled data from 5 donors showing percentages of peptide-loaded target (CFSElow) at each condition, where bar height represents the mean value.
GITRL costimulation promotes CD4TC proliferation but it has no effect on the HIV reservoir size
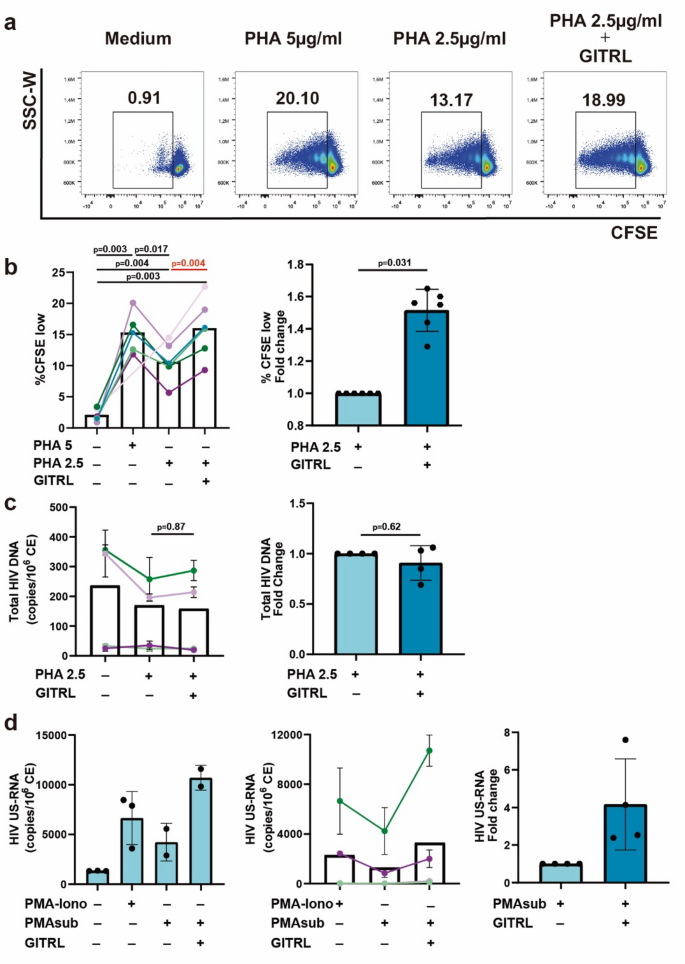
We examined GITRL effect on the proliferation of purified CD4TCs from 6 On-ART donors. After 7 days of co-culture with suboptimal concentrations of PHA plus GITRL (10 µg/mL), a significant increase in proliferating CD4TCs (1.5x) was observed compared to PHA alone (Fig. 5a–b). To evaluate if GITRL costimulation had an impact on the viral reservoir size, we measured cell-associated total HIV DNA across unstimulated, suboptimal PHA and suboptimal PHA + GITRL conditions from 4 donors (Fig. 5c). Both PHA alone and PHA + GITRL conditions showed a non-significant decline in HIV DNA, probably due to replication-mediated death of cells harboring viral DNA or to a higher duplication rate of non-infected cells. Interestingly, we observed no significant differences between conditions with or without GITRL, suggesting that the effect on proliferation is comparable between HIV-infected and non-infected cells and that GITRL costimulation does not impact the size of the HIV reservoir.
GITRL costimulation reverts HIV latency
HIV transcription is dependent on NF-κB activation22. Given GITR signaling relevance in this process23,24 we hypothesized that GITR costimulation could have an effect on HIV latency reversion. To test this, we performed an ex vivo latency reversal assay. Purified CD4TCs from 4 On-ART participants were treated for 16 h with a suboptimal PMA concentration (2ng/mL) plus GITRL (10 µg/mL) or PMA (50ng/mL) and Ionomicyn (1 µg/mL) as a positive control. While PMA alone did not induce the same level of viral transcription observed in the PMA-Ionomycin condition (Fig. 5d), co-stimulation with GITRL led to a 4.16-fold increase in HIV US-RNA, suggesting a role of GITR costimulation in viral reactivation and positioning this pathway as an interesting target for the development of new latency reversal agents.
GITRL-mediated effect on CD4TC proliferation, HIV reservoir size and HIV latency reversion. (a) Representative flow cytometry dot plot showing the frequency of proliferating CD4TCs after 7 days of costimulation with optimal (5 µg/mL) and suboptimal (2.5 µg/mL) concentrations of PHA plus 10 µg/mL of GITRL. Proliferating cells were identified as CFSElow. (b) Pooled data from 6 donors. Percentages of proliferating CD4TCs for each condition (left) and fold change of suboptimal PHA condition plus GITRL (right). Bar height stands for mean values and whiskers represent standard deviation. p-values were calculated using a mixed-effects model for repeated measures with missing values, applying Tukey’s correction for multiple comparisons (left) and one sample t test (right). (c) Total HIV DNA evaluation in proliferating CD4TCs after 7 days of costimulation with suboptimal concentrations of PHA in the absence or presence of GITRL. Left: Total HIV DNA levels were reported as the number of HIV copies per 106 cell equivalents (CE). Each point represents the mean of three experimental replicates. Right: fold change of suboptimal PHA condition plus GITRL. Bar height stands for mean values and whiskers represent standard deviation. p values were calculated with Friedman test followed by Dunn´s post-hoc test (left) or one sample Wilcoxon test (right). (d) Left: scatter plot with bar graph from a representative donor showing the levels of HIV US-RNA after 16 h of costimulation with optimal (PMA = 50 ng/mL + Ionomicyn 1 µg/mL; PMA-iono) and suboptimal (PMA = 2 ng/mL; PMAsub) policlonal stimuli, in the absence or presence of GITRL. Each condition was evaluated in triplicates. Replicates positive for the no reverse transcriptase control condition were excluded. HIV US-RNA levels were reported as the number of RNA copies per 106 cell equivalents (CE). Center: pooled data from 4 donors. Each point represents the mean value of experimental replicates. Right: Fold change of suboptimal PMA condition plus GITRL. Bar height stands for mean values and whiskers represent standard deviation.