Construction of a M. tuberculosis pangenome reference graph
We obtained 1363 long-read datasets (ONT or PB) from public repositories and performed quality control for read length and decontamination of human DNA before assembling genomes with Flye18. After also incorporating 176 long-read assemblies from NCBI to enlarge the dataset, all assemblies were evaluated with QUAST19, with 834 remaining after filtering (see Methods; Supplementary Fig. 1). To ensure phylogenetic coverage, we sequenced 25 isolates from underrepresented lineages with ONT, achieving representation of lineages 1–9, including phenotypically drug-resistant isolates without known resistance mutations (Supplementary Table 1). Sub-lineage 2.2.1 (Beijing) was the most frequent of the dataset (23.4%), while lineage 4 and its sub-lineages dominated overall (48%; Supplementary Fig. 2). The final cohort of 859 assemblies, representing both genetic and geographical diversity (Supplementary Data 1 and Supplementary Fig. 3), had a mean N50 of 2.13 Mb (range: 11.2 kb–4.45 Mb) and mean coverage depth of 125× (range: 7 × -859 × ; full metrics in Supplementary Data 2).
To construct the Mtb-PRG, we used minigraph20, which efficiently incorporates sequences that are larger than 49 bp and missing from the reference genome while preserving reference coordinates. Starting with the H37Rv reference genome (accession: NC_000962.3) as the backbone, additional genomes were added to the Mtb-PRG in lexicographic order by sample name. Newly added sequences formed nodes, which represent distinct segments of genomic sequences that are connected by paths, resulting in 9350 nodes in total. Nodes connected to each other but not part of the reference formed 848 bubbles, with the largest spanning 97,162 bp (H37Rv coordinates: 3,386,328–3,483,490; Supplementary Fig. 4).
Genotyping SVs using the Mtb-PRG and miniwalk is more precise than using a traditional short-read SV caller
PRG-involving approaches have been shown to call SVs more accurately from short-read sequencing data compared to traditional single-reference genome approaches16. Since most Mtb genomic data are derived from Illumina short-read sequencing, accurately identifying SVs using such data is valuable. However, no existing method genotypes SVs directly from graphs generated using minigraph. To address this, we developed miniwalk, a tool that identifies SVs by comparing the graph traversal paths of an assembly or long reads, to a reference genome mapped to a minigraph PRG (Supplementary Methods 1). Miniwalk outputs insertions (INSs), deletions (DELs), duplications (DUPs), and inversions (INVs).
We benchmarked miniwalk using 16 samples, each of which had a high-quality, polished long-read assembly to act as truth. To avoid bias, the 16 samples were removed from the PRG for the benchmarking21. Truth SVs were called using SVIM-ASM22 on assemblies mapped to the H37Rv reference genome, while miniwalk called/queried SVs on assemblies mapped to the Mtb-PRG. Precision and recall, calculated using the miniwalk bench module (Supplementary Methods 2), averaged 0.85 and 0.82, respectively, showing comparable genotyping performance to SVIM-ASM (Fig. 1a). Nevertheless, precision varied across SV types, with DELs of size 500–1000 bp having the lowest precision (0.5; Fig. 1b), though fewer SVs were found in that size range. Further analysis revealed that this drop was largely driven by a single complex SV present in 11 assemblies, consistently detected by miniwalk but missed by SVIM-ASM (Supplementary Fig. 5). We also constructed an additional PRG excluding assemblies with N50 <90,000 bp and coverage <25 × to determine whether a more stringent approach should have been taken. The average precision and recall using the more stringent PRG was 0.84 and 0.83, respectively, indicating that our initial PRG had better precision. To test whether assembly ordering introduced bias, we rebuilt the Mtb-PRG with assemblies ordered by lineage rather than lexicographic filename and re-evaluated SV genotyping performance. Precision and recall did not differ significantly (two-sided t-tests; precision p = 0.96, recall p = 0.29), indicating that graph construction order had no measurable effect on genotyping accuracy (Supplementary Fig. 6). To complement real-data benchmarking, we simulated an assembly by spiking real SVs into the H37Rv reference. Miniwalk achieved 0.92 precision and 0.82 recall, outperforming results on real data (0.78/0.81), highlighting the reduced complexity of simulations.
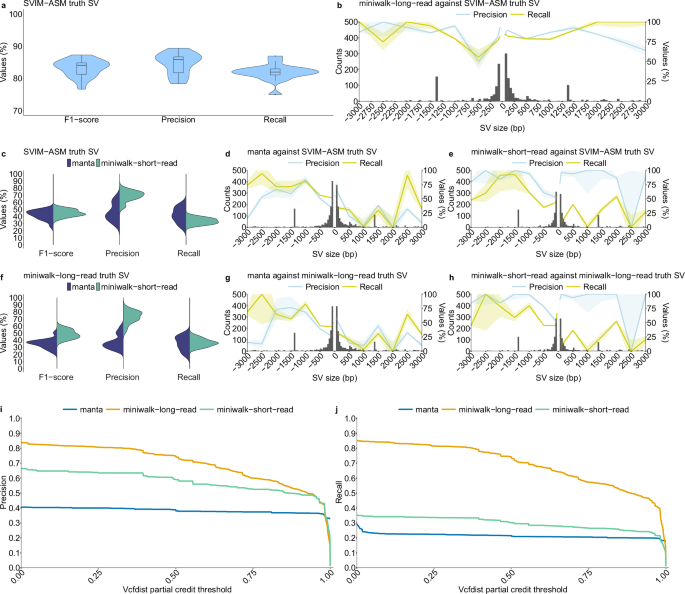
a Results comparing Mtb-PRG and miniwalk SV genotyping against SVIM-ASM truth SV calls using the 16 independent long-read, polished assemblies. Each data point represents an SV call set from one assembly, with assemblies serving as independent biological replicates. No technical replicates were used. No explicit control group was included; instead, distributions reflect genotyping performance across the independent assemblies. Box plots indicate the median (center line), interquartile range (box), and whiskers extending to 1.5 × the interquartile range; outliers are plotted individually. b Histogram with 50 bp bins showing the found SVs in (a), with two segments with 50 bp bin from SVs of size 50–100 bp, 400 bp bin for SVs of size 100–500 bp, 500 bp bin for SVs of size 500–3000 bp and SVs larger than 3000 bp grouped together. The two segments show the precision and recall means across different SV sizes. Shaded regions represent 95% confidence intervals (mean ± 1.96 × standard error). c Results for genotyping SVs from Illumina data mapped to H37Rv and calling with manta and mapping to the Mtb-PRG and calling with miniwalk against the SVIM-ASM truth SVs in (a). d Average SV precision, recall and frequency of short-read data stratified by length from manta SVs against the SVIM-ASM truth SVs. Bin sizes as in (b). Shaded regions represent 95% confidence intervals (mean ± 1.96 × standard error). e Same plot as (d) for genotyped short-read SVs using the Mtb-PRG and miniwalk against the SVIM-ASM truth SVs. f Results for genotyping SVs from Illumina data mapped to H37Rv and calling with manta and mapping to the Mtb-PRG and calling with miniwalk-long-read SVs as truth (a). g Same plot as (d) with miniwalk-long-read SVs as truth. h Same plot as (e) with miniwalk-long-read as truth SVs. i Precision values across different vcfdist partial credit thresholds for miniwalk-long-read (long-read assemblies mapped to the graph, genotyped with miniwalk), manta (short reads mapped to H37Rv, genotyped with manta) and miniwalk-short-read (short-read assemblies mapped to the graph, genotyped with miniwalk) SVs against the SVIM-ASM truth SVs. j Similar to (i), for recall values.
To assess short-read data performance, Illumina reads used for polishing the 16 assemblies were analyzed using manta23, a leading short-read SV caller24. Reads were also assembled with shovill25, mapped to the Mtb-PRG, and SVs genotyped with miniwalk. To assess precision and recall of SV genotyping using these two approaches (linear H37Rv/manta and Mtb-PRG/miniwalk), we compared both to SVIM-ASM truth SVs as in Fig. 1a. We found miniwalk demonstrated significantly higher precision (0.7 vs. 0.46 for manta; two-sided t-test, p = 1e-6) at the cost of slightly lower recall (0.34 vs. 0.42 for manta; two-sided t-test, p = 0.007; Fig. 1c). Breaking down by SV type, manta achieved a precision and recall of 0.42 and 0.63, respectively, for DELs and 0.42 and 0.36, respectively, for INSs (Fig. 1d). Meanwhile, miniwalk achieved a precision and recall of 0.64 and 0.54, respectively, for DELs, and 0.86 and 0.19, respectively, for INSs. DELs sized >1000 bp had precision exceeding 0.8 (Fig. 1e). Considering SVs genotyped with miniwalk from the long-read assemblies mapped to the Mtb-PRG as the truth (SVs from Fig. 1a, b; miniwalk-long-read truth SV), precision and recall decreased for Illumina data mapped to the H37Rv linear reference (Fig. 1f, g) but improved slightly for short-read assemblies mapped to the Mtb-PRG (Fig. 1f, h). These results highlight miniwalk’s precision advantage when using a PRG compared to a linear genome-based method. To explore whether recall could be improved, we created another Mtb-PRG, retaining SVs ≥20bp, and evaluated it against the short-read assemblies. We observed, on average, a 2% increase in recall at the cost of a 2% drop in precision (Supplementary Fig. 7).
To further validate the use of miniwalk, we used partial credit from vcfdist26 to evaluate the similarity between our variant calls and the SVIM-ASM truth set. Vcfdist assigns each pair of variants a partial credit score ranging from 0 to 1, based on edit distance (sequence content and position), where 1 means identical variants and 0 means no overlap. We applied growing partial credit (ct) thresholds starting from 0.001. Lower ct thresholds allow called SVs with partial similarity to the truth to be considered true positives, while higher ct thresholds impose stricter criteria, requiring close matches in SV type, position, and sequence content. This evaluation included the previous long- and short-read assemblies mapped to the Mtb-PRG as well as the variants called from manta. For miniwalk-long-read SVs, precision and recall showed a gradual decline as the threshold increased, reflecting differences in sequence content (e.g., long-read sequencing errors, SNPs) and positions between called and truth SVs. Nonetheless, precision (0.75) and recall (0.74) remained moderately high for long-read assemblies mapped to the Mtb-PRG at a ct threshold of 0.5, demonstrating the accuracy of SVs called using our graph-based approach. For short-read data, miniwalk outperformed manta across all ct thresholds up to 0.985 for both precision and recall (Fig. 1i, j).
Further analysis revealed that SVs in complex regions were the ones driving the decrease in precision and recall in short-read data (Supplementary Note 1).
These results suggest using a genome graph as a superior alternative to a linear reference genome for genotyping SVs. With this in mind, we proceeded to learn about SVs in Mtb using the Mtb-PRG and miniwalk.
Landscape of structural variation across the M. tuberculosis genome and phylogeny
We sought to improve the characterization of SVs across diverse Mtb lineages using high-quality long-read data. Assemblies used to construct the Mtb-PRG were mapped back to the PRG. The 38 most fragmented assemblies were excluded from this analysis as they could not be mapped using minigraph’s default parameters (see Methods). SVs in the remaining assemblies (821) were genotyped against the H37Rv reference genome using miniwalk. To further define the diversity of SVs in the Mtb genome, DELs and DUPs in close proximity were grouped as copy number variants (CNVs). SVs within ±25bp proximity, ±50 bp size, and of the same type were considered identical across isolates.
A total of 3077 unique SVs were identified relative to H37Rv, broken down into 1127 INSs, 1694 DELs, 59 DUPs, 58 INVs, and 139 CNVs. The largest SV was a 66,582bp region (H37Rv coordinates: 2,901,855–2,968,437) that was inverted and translocated to a novel genomic position (H37Rv coordinates: 415,370) in lineage 9 (Supplementary Fig. 8). The translocation implicated 70 genes and truncated Rv2577 and Rv0345. This rearrangement highlights the complexity of SV patterns observed in Mtb, though most isolates presented small DELs, CNVs and numerous IS6110 elements (~1358bp; Fig. 2a). We found various INS-DEL pairs of similar size (≥90%) and high identity (≥90%) across each genome (range of 0–3 per genome), likely stemming from IS6110 transposition. A principal component analysis (PCA) performed with the SV genotypes revealed that SVs contribute to population structure, especially along PC2 which separates the ancient (L1, L5–9) from the modern (L2–4) lineages (Fig. 2b). Notably, the “TbD1” SV at position 1,761,789 explains 2% of the variability in PC2 (Supplementary Fig. 9). The allele frequency spectrum of SVs revealed that over half (1617/3077) were singletons (Supplementary Fig. 10a). Importantly, singleton SVs were not enriched in fragmented assemblies, suggesting they represent genuine rare variation rather than artifacts (Supplementary Fig. 10b).
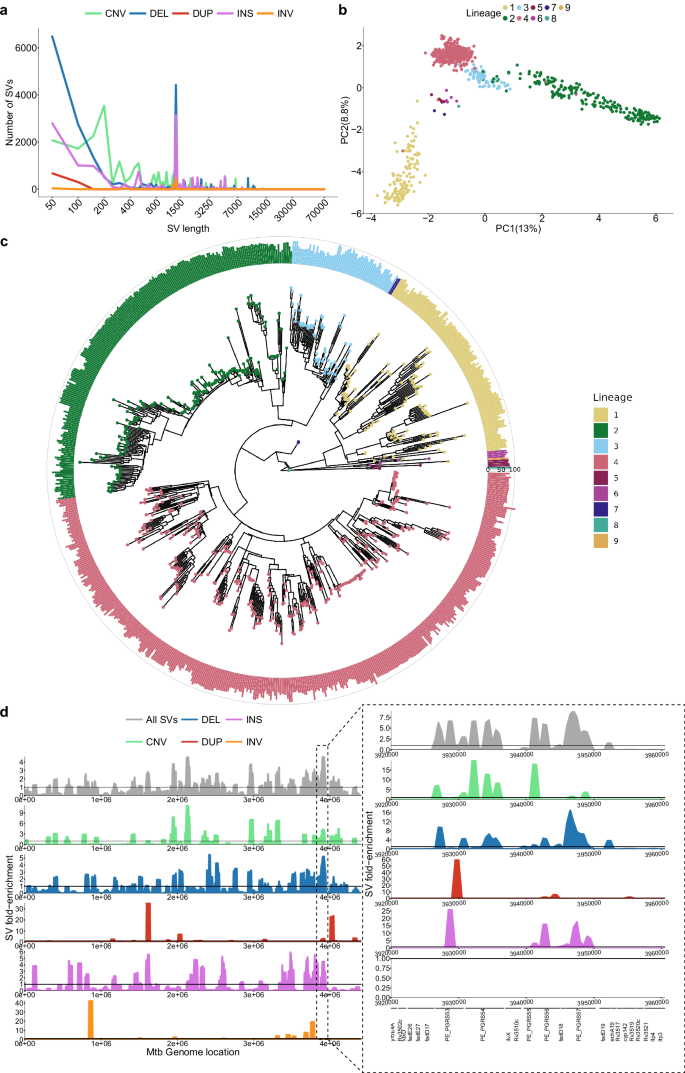
a Distribution of SVs in isolates by size and type. The x-axis is in log10 scale. b PCA based on the isolates’ SV genotypes. c Phylogenetic tree filtering out 75 isolates with disconcordant SNP genotypes and lineage calls due to lower quality long-read SNP calls. Barplots represent the number of SVs compared to the ancestral reference sequence MTBC0. Rings, inner to outer, represent 0, 50, and 100 SVs, respectively. d Distribution of SVs along the genome by their fold-enrichment and type across all isolates. Horizontal lines represent no fold change of a specific position from the rest of the genome. The panel on the right represents the zoomed-in region of 3,200,000–3,600,000 with its own fold-enrichment values and genes along the region. Source data are provided as a Source Data file.
To understand the acquisition of SVs across lineages over time, SVs were genotyped against an ancestrally reconstructed sequence (MTBC027) and mapped to a phylogenetic tree from the genotyped isolates (Fig. 2c). This showed that all isolates have acquired a similar amount of SVs across the phylogeny, the least being L4 with an average of 67 SVs per isolate and the most being L8 with 82 SVs (though 1 isolate was used). Interestingly, Beijing lineage isolates segregated into two branches, one with an average of 86 SVs per isolate and the other with an average of 68 SVs per isolate, indicating distinct evolutionary trajectories of SV accumulation within this sub-lineage. When analyzing L4 sub-lineages, the globally spread generalists L4.1.2 and L4.328 had an average of 60 and 63 SVs per isolate, respectively, whereas the specialists L4.1.3 and L4.6.228 had an average of 74. From these results, generalist lineages have significantly less SVs than specialist lineages (two-sided Wilcoxon test, p value = 0.03). SVs are generally deleterious in nature, so lineages with more SVs could have suffered more deleterious mutations than those with less SVs, hindering their geographical spread28. Alternatively, the higher number of SVs in specialist sub-lineages could reflect the process of host-population adaptation via reductive genomic evolution29.
To identify SV hotspots, we scanned the genome using 50,000 bp windows with 5000 bp steps and calculated fold-enrichment relative to the average SV density. Results indicated an uneven SV distribution, with a prominent hotspot between 3,200,000–3,600,000 bp enriched in PE/PPE genes (Fig. 2d). We found that 75% of SV breakpoints fall in genes. This is lower than expected by chance (mean random breakpoints = 92%, absolute z-score = 6.42, p value = 1e-10), likely due to the deleterious effect of SVs (Supplementary Fig. 11). Moreover, SV breakpoints were significantly associated with GC-rich sequences compared to randomly sampled genome regions (two-sided Wilcoxon test, p = 5e-8; Supplementary Fig. 12). PE/PPE genes are known for their GC-rich repetitive regions, which serve as an SV hotspot. These findings highlight that SVs in Mtb are not randomly distributed but are enriched in specific genomic features and regions.
Pangenomic insights into structural variation in the ESX loci of M. tuberculosis
The Mtb genome encodes five type-7 secretion systems, which are known to play an important role in pathogenesis and virulence, the most studied being ESX-130,31. ESX-5 is important for full virulence32 and has been involved with various functions, such as the secretion of PE/PPE proteins, the activation of the host cell inflammasome response or macrophage death induction31. Its paralogs’ (ESX-5a, ESX-5b, and ESX-5c) functions, however, are less understood. To determine the presence of SVs in the ESX-5 loci and which are more conserved, we analyzed the haplotypes from the 821 long-read assemblies mapped to the Mtb-PRG.
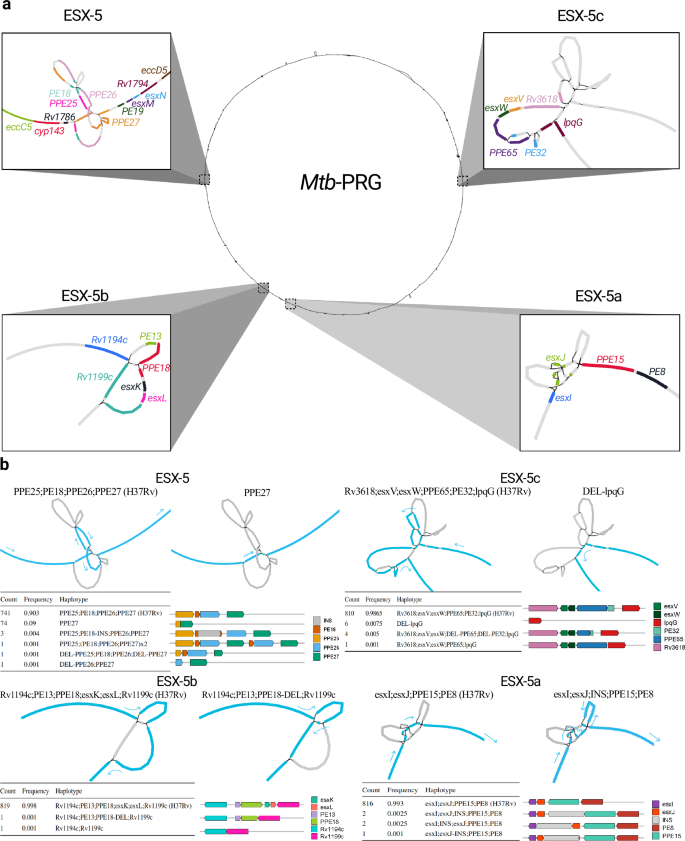
All ESX-5 loci had one SV bubble spanning their locus (Fig. 3a). Most isolates lacked SVs across these loci, with most non-reference haplotypes being present solely in 1–6 isolates. Nevertheless, the most frequent non-reference haplotype was a ppe25–ppe27 deletion, present in 9% of all isolates (Fig. 3b). The deletion was found in all sub-lineage 4.4 isolates and sporadically in other lineages (L4.1, L4.3, L4.5, L1, L3, and L8). The deletion can most likely be attributed to non-allelic homologous recombination, as the 3’ sequences of ppe25 and ppe27 are highly homologous (Supplementary Fig. 13), that is, the deletion spans the ppe25-ppe27 locus and removes ppe25, pe18, and ppe26 while leaving ppe27 intact. The deletion was found to provide higher persistence in mouse models and showed convergent evolution in M. canettii33, which could explain its homoplasic effect across the Mtb phylogeny (Supplementary Fig. 14). A previous study documenting this deletion hypothesized that the loss of immunomodulatory PE/PPE genes could help the bacteria escape the immune system34.
a Locations of ESX-5, ESX-5a, ESX-5b, and ESX-5c in the Mtb-PRG. Different colors in each loci represent different genes. b Structural haplotypes of each ESX-5 locus. The order of the loci is the same as in (a), the graph representation is of the H37Rv reference on the left and of the non-reference most frequent haplotype on the right, for each locus. Nodes traversed through each haplotype are colored in blue with arrows pointing towards the direction of the genome. Tables and gene representations show the frequency of each haplotype and its effect on the gene content. Visualizations done using Bandage98.
We extended the analysis to assess the impact of SVs across all of the ESX genes. To understand the probability of observing SVs in a gene, we calculated the trimmed (10%; removal of outlier genes with high SV counts) mean number of SVs in essential (0) and non-essential (0.76) genes35,36, with 33% of genes being overlapped by an SV. Genes prefixed with the letters “ecc” and “mycP” in the ESX loci encode proteins which form the secretion system responsible for translocating proteins involved in virulence37. Consistent with their essential role in virulence38, we found 0 isolates from our collection of 821 to have SVs impacting structural components of the ESX-3 and ESX-5 loci, and only one with an SV impacting ESX-1 genes eccE1 and mycP1. In contrast, the ESX-2 and ESX-4 loci, which are non-essential and have yet to be ascribed a function38, had between 1 and 16 isolates with SVs overlapping their gene members38 (Supplementary Fig. 15a). Among all genes, espI from ESX-1 had the highest number of SVs, with 125 isolates exhibiting overlaps (Supplementary Fig. 15b). EspI acts as a negative regulator of the ESX-1 secretion system; thus, the high frequency of SVs in this gene may suggest positive selection for maintaining constant expression of the system39. Genes within the ESX loci prefixed with the letters “esx” encode proteins that are secreted through the ESX apparatus and thought to be implicated in host-pathogen interaction. Across our collection of 821 isolates, no esx genes in the ESX-1 and ESX-5 loci were found to have any SVs (Supplementary Fig. 15c), however esx genes in the ESX-2 locus displayed SVs in three isolates. Of the ESX-5 paralogs, ESX-5a was the most strongly conserved (1 isolate with an SV falling in a gene), suggesting deletions of this locus are less well tolerated than deletions in ESX-5b and ESX-5c (Fig. 3b and Supplementary Fig. 15c, d). Interestingly, one instance of a full ESX-5c deletion was found in the geographically restricted African lineages (L6 and L9).
A genome and phylogeny-wide screen of SV positive selection in M. tuberculosis
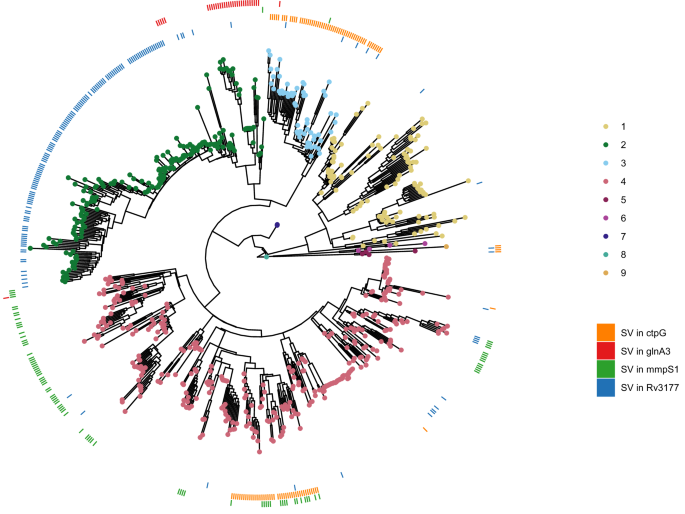
We performed a systematic screen for genes which evolved SVs repeatedly across the phylogeny, following the rationale that this may reveal genes targeted by positive selection40. The criteria for our screen stipulated that the gene had to be affected by an SV in at least ten isolates, belonging to more than one Mtb lineage. We further excluded genes in or ±3000 bp of low-mappability regions10, as those regions are more prone to structural rearrangements, which could be falsely perceived as positive selection. This analysis resulted in 59 homoplasic gene candidates. The gene with the strongest homoplasic effect was Rv3177, with SVs in 158 isolates from L1.2.1, L1.1.1, L2.2.1 (82% of all isolates), L2.2.2, L3.1.2, L3.1.3, various L4 sub-lineages and L5 in 15 different breakpoints across the gene (Fig. 4). Rv3177 is a probable peroxidase involved in detoxification of reactive oxygen species (ROS). In a previous study, variants in Rv3177 were associated with isolates harboring the isoniazid resistance-conferring katG-S315T mutation, suggesting a possible compensatory role in drug resistance41. Another gene that acquired SVs in different branches of the phylogeny was ctpG with six different DEL breakpoints across the gene (Supplementary Fig. 16) in L3, L4.3.4 and L5 (Fig. 4). CtpG is a zinc (Zn) exporter whose deletion has been associated with Zn accumulation42. During host infection, Mtb goes from Zn-rich to Zn-limited environments43. A non-functional Zn exporter could possibly provide bacteria in Zn-limited conditions a selective advantage. Other interesting genes were mmpS1, with SVs in various L4 sub-lineages with a prominence in L4.1 as well as a few isolates in L2 and L3, and glnA3, with SVs in all L2.2.2 isolates and sporadically in L2.2.1, L3, and L4.1 (Fig. 4).
Same phylogenetic tree as in Fig. 2c. The outer rings represent the presence (color) or absence (white) of an SV in a specific gene for each isolate. The patchy distribution of some SVs within clades may be explained by inaccurate placement of isolates in the phylogeny, likely due to higher SNP error rates associated with long-read sequencing, especially in isolates generated using earlier versions of these technologies. Source data are provided as a Source Data file.
We sought to validate these results within a larger cohort. To do so, we downloaded a list of 44,709 high-quality Illumina sequences grouped in a previous study17. After assembly, we ended with 41,134 SV VCF files. We found a few isolates with SVs in Rv3177 and mmpS1, as Rv3177 is in a large bubble, likely hard to genotype using short-read data, and the mmpS1 SVs were all INSs, variants with low genotyping recall using short-read data. We were able to confirm the same distribution of SVs for ctpG and glnA3 (Supplementary Fig. 17). A similar distribution was observed for SVs in ethA, a gene whose deletion is associated with ethionamide (ETO) resistance, with a main cluster located in the Beijing lineage. For pncA, a pyrazinamide (PZA) resistance-associated gene, we found less isolates with SVs and the distribution was not as clustered, though mainly isolates in L2 had SVs in that gene. These results showcase the added benefit of using a pangenome to identify key genes implicated in Mtb evolution, and highlight four candidates for prioritization in future research.
Characterization of a M. tuberculosis lineage 1.2.1-specific copper exporter deletion
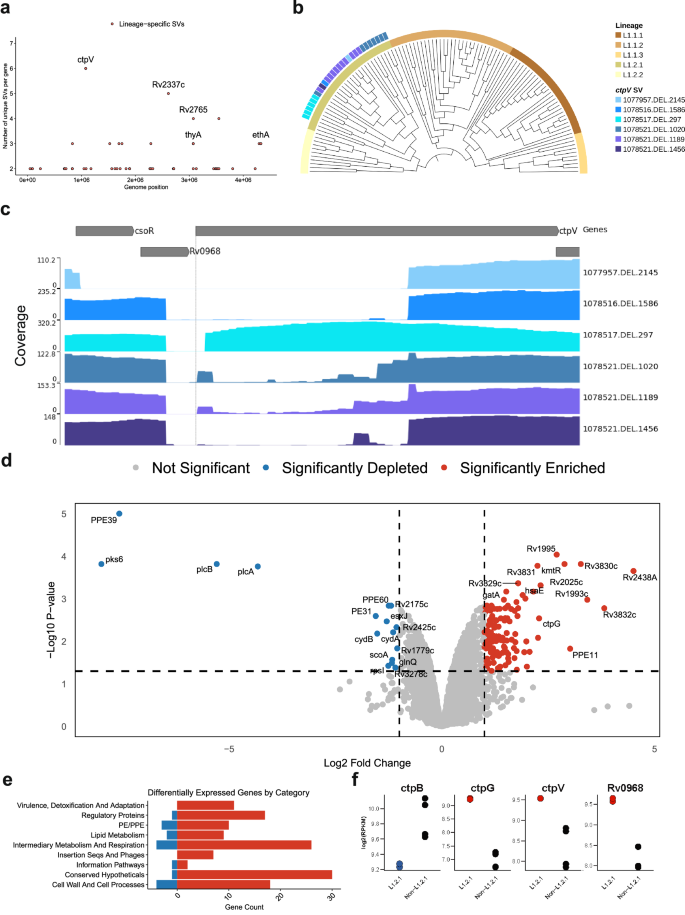
We expanded the previous screen to detect genes with multiple lineage-specific SVs. This analysis revealed diverse deletions of the ctpV gene, unique to the L1.2.1 sub-lineage (excluding its deepest branch split; Fig. 5a and Supplementary Fig. 18). These deletions increased in size further from the branch split (Fig. 5b) and overlapped the gene’s start codon, indicating a loss-of-function mutation (Fig. 5c). SNPs could not be reliably called for all the isolates as long-read data were used, nevertheless, we found no loss-of-function variants in this gene in other L1 sub-lineages. CtpV is a putative P-type ATPase copper (Cu) exporter protein, with the main Cu binding site located in the first 66 amino acids. A multiple sequence alignment of the protein sequence among diverse mycobacteria showed >80% pairwise identity across the Mycobacterium genus (Supplementary Fig. 19), aside from Mycobacterium leprae, highlighting the importance of this protein not only in Mtb.
a All genes across the genome with the number of isolates that have SVs overlapping them on the y-axis. Red-colored points represent those that passed the second positive selection screen; i.e., those genes that have different SVs in the same gene, unique to a lineage. b Phylogenetic tree of all lineage 1 isolates, showing those that present ctpV deletions and/or non-synonymous mutations. SV IDs are formulated as following “{start position of SV}.{SV type}.{size of SV}”. Tree visualized with TreeViewer99. c Read coverage across the csoR operon for each unique deletion spanning ctpV. Y-axis shows the read depth at each genomic position. SV ID is the same as (b). Visualization done using pyGenomeTracks. d Volcano plot summarizing the differential expression results comparing L1.2.1 against L1.1.2 and L1.1.1 isolates. P values are calculated using a two-sided t-test, with Benjamin–Hochberg multiple hypothesis correction. Log fold change on the x-axis. e Functional categories from Mycobrowser of the significantly differentially expressed genes. f Dot plots showing expression patterns for four copper-associated genes in L1.2.1 against L1.1.2 and L1.1.1. In red are genes enriched, and in blue are genes depleted in L1.2.1. Source data are provided as a Source Data file.
L1.2.1 is predominant in Southeast Asia, especially the Philippines, the fourth country in the world with the highest burden of active tuberculosis1, accounting for ~65–80% of all lineages in the country44,45. Moreover, this sub-lineage shows high drug resistance rates, high transmissibility and is the most prevalent L1 sub-lineage globally46,47. Given these characteristics, we hypothesized that this deletion impacts the expression of copper-associated genes, potentially contributing to the unique adaptive traits of this sub-lineage. To determine this, we compared the transcriptional profile of an L1.2.1 isolate (297bp ctpV confirmed deletion) against L1.1.1 and L1.1.2 isolates under normal growth conditions48 (Supplementary Table 2 and Supplementary Fig. 20). This analysis revealed 147 differentially expressed genes (Fig. 5d). Functional categorization revealed that the L1.2.1 isolate showed enrichment of genes involved in virulence, detoxification and adaptation (Fig. 5e). ctpG and Rv0968 were among the differentially expressed genes (Fig. 5f). Rv0968 is part of the cso operon, where ctpV lies. This operon is controlled by CsoR, which inhibits its own expression in normal conditions. However, when Cu is present, it attaches to CsoR and inhibits its function, enabling Rv0968 and ctpV expression49,50. The higher expression of both Rv0968 and ctpV (nonsignificant; adjusted p value = 0.053) was indicative of Cu accumulation inside these isolates (Supplementary Fig. 21). Moreover, the enrichment of ctpG, another P-type ATPase metal transport protein, has been previously associated with Cu accumulation in the cell, as well as the depletion of ctpB, a P-type ATPase Cu transport protein associated with Cu uptake in Cu-replete medium (nonsignificant; adjusted p value = 0.059)50,51. Nevertheless, genes associated with oxidative stress (i.e., sigE,furA,nuoB) or protein stress (i.e., rpsR,rplI,rpsQ) were not differentially expressed, indicating that Cu was not present at toxic levels (Supplementary Data 3)50. Altogether, these results are indicative of higher Cu accumulation in an L1.2.1 isolate compared to other L1 sub-lineages.
Discovery of DR-associated SVs in 41,134 M. tuberculosis isolates
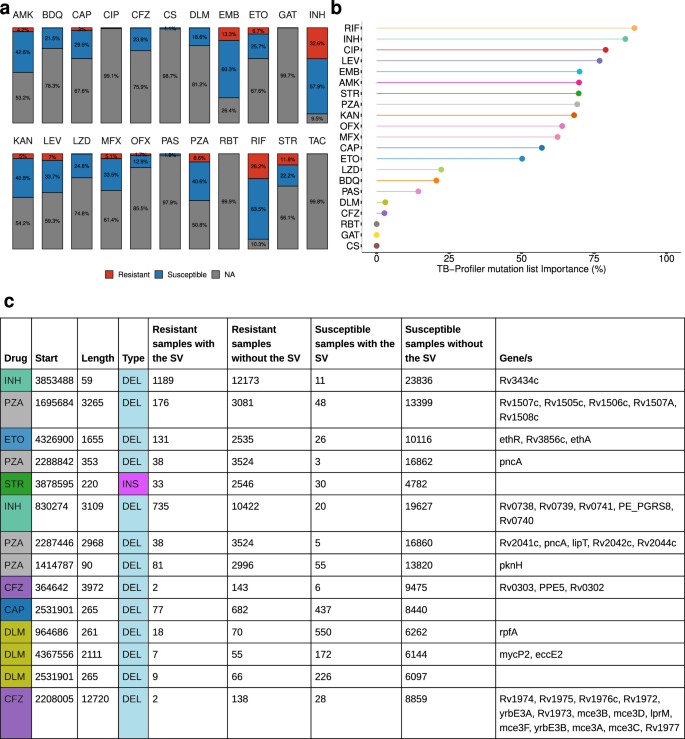
Using the Mtb-PRG and miniwalk, we aimed to discover DR-associated SVs across various first and second-line drugs for treating TB. The previous list of 41,134 isolates also contained phenotypic information for at least one drug, with first-line drugs being vastly more represented than second-line drugs (Fig. 6a). To search for associations, we used two parallel methods: Firth logistic regression and a supervised machine learning model (xgboost), testing each drug independently and accounting for population structure and TB-Profiler predictions52. Analyses were conducted at both the SV and gene levels (SV-gene burden test).
a Stacked barplot showing the percentage of isolates that are susceptible, resistant or have no information (NA) across the 41,134 isolates, for each drug. Percentage labels are only shown when the value is ≥1%. b Machine learning importance scores (%) of the TB-Profiler mutations predictions. TAC is not illustrated as only resistant isolates were available. c Thorough description of the 14 associated SVs, ranked by their adjusted p value (top – lower p value). AMK amikacin, BDQ bedaquiline, CAP capreomycin, CIP ciprofloxacin, CFZ clofazimine, CS cycloserine, DLM delamanid, EMB ethambutol, ETO ethionamide, GAT gatifloxacin, INH isoniazid, KAN kanamycin, LEV levofloxacin, LZD linezolid, MFX moxifloxacin, OFX ofloxacin, PAS para-aminosalicylic acid, PZA pyrazinamide, RBT rifabutin, RIF rifampicin, STR streptomycin, TAC thioacetazone. Source data are provided as a Source Data file.
Logistic regression revealed 14 non-canonical SVs (post-filtering) significantly associated (adj. p value <0.05) with seven DR phenotypes (Supplementary Data 4). Associations in PE/PPE genes were excluded due to their SV hotspot nature (list of discovered associations: see Supplementary Data 5). Xgboost outputs the estimated importance of each SV and TB-Profiler prediction to the analysis (ranging between 0–100%), indicating the relevance of these factors in classifying DR phenotypes. This analysis highlighted the contribution of the current list of WHO mutations, as identified by TB-Profiler, in predicting DR, with first-line drugs as well as fluoroquinolones and aminoglycosides having high importance scores (importance >50%). However, newer drugs such as bedaquiline, delamanid or clofazimine showed lower importance scores (Fig. 6b). The SV-gene burden test revealed 27 non-canonical genes associated with drug resistance across ten drugs (Supplementary Data 6). To increase confidence in our findings, we performed, for each associated gene and SV, a permutation analysis to estimate the likelihood of our observed p values under a null distribution. This showed that under the null hypothesis, there’s <5% chance of observing these associations (absolute z-score range 1.65–2.57), suggesting statistical significance for all variants.
For isoniazid (INH), we found two SVs and three genes associated with DR (Fig. 6c and Supplementary Figs. 22, 23). The most significant SV was a deletion overlapping Rv3434c (Fig. 6c), a gene previously associated with kanamycin (KAN) and amikacin (AMK) DR53. An overwhelming 99% of isolates with a mutation in this gene presented an isoniazid resistance phenotype, the vast majority from L2. Nevertheless, only five isolates did not present TB-Profiler predictions for INH DR. Another deletion spanning Rv0738-Rv0741 was predominantly found in resistant isolates (97%), with an Rv0740 knock-out having previously been associated with INH resistance54. Five isolates with SVs in Rv0740 did not have TB-Profiler predictions for INH DR. These findings underscore the potential of SVs in identifying additional resistance mechanisms, particularly in a widely distributed and commonly used drug like INH, not captured by TB-Profiler. For rifampicin, no non-canonical SVs or genes with SVs were detected in our analysis. For PZA, we found four SVs and six genes. The highest-scoring gene was pncA, with 2.3% of DR isolates having SVs overlapping that gene (Supplementary Fig. 23). An operon with SVs and also highly associated with PZA-DR was Rv1505c-Rv1507c, which plays a role in diacyltrehalose biosynthesis, a lipid that forms part of the cell wall55. The only gene with SVs associated with ethambutol (EMB) DR was accE5, whose encoded protein is found in the cell wall56 (Supplementary Fig. 23).
For second-line drugs, we found non-canonical SVs and genes with SVs for AMK, bedaquiline (BDQ), capreomycin (CAP), ciprofloxacin (CIP), clofazimine (CFZ), delamanid (DLM), ETO and streptomycin (STR) (Fig. 6c, Supplementary Fig. 23, and Supplementary Note 2). Our results highlight the importance of different types of SVs in DR, such as an insertion associated with STR DR, as well as the role of non-coding regions, such as a deletion upstream Rv2258c associated with CAP DR. The low importance scores of second-line, recently introduced drugs (BDQ, CFZ, and DLM) highlight the need to find more DR variants.