We implemented a collection of machine-learning based models to generate out-of-sample predictions for the number of COVID-19 confirmed cases, as reported by the Centers for Disease Control and Prevention (CDC), for the time period between October 18th, 2020 and April 15th, 2022. These models included the model we propose: FIGI-Net, as well as Autoregressive statistical models, a collection of recurrent neural network (RNN) based models (GRU, LSTM), a stacked bidirectional LSTM (biLSTM), a set of LSTM-based models incorporating temporal clustering (TC-LSTM and TC-biLSTM), and a “naive” (Persistence) model43 to be used as a baseline. Additionally, for comparison purposes, we implemented five additional models commonly found in the forecasting literature: ARIMA28, Prophet44, and Transformer-based models (Transformer45, Informer46, and Autoformer47) both with and without clustering.
We used our models to retrospectively produce daily forecasts for multiple time horizons, h, ranging from h = 1, 2. . . , 15 days ahead. Visual representations of our predictions, alongside the actual observed COVID-19 confirmed cases, are presented in Figs. 3, 4, and 5 for county, state and national levels. We evaluated our model’s performance by comparing our out-of-sample predictions with subsequently observed reported data in each time horizon h using multiple error metrics that include: the root mean square error (RMSE), relative RMSE (RRMSE), and mean absolute percentage error (MAPE) metrics, as detailed in the section “Model evaluation”. In addition, we compared the prediction performance of our model with a diverse set of statistical and machine learning models, including state-of-the-art forecasting methodologies reported by the CDC (a comprehensive list of these models is provided in the section “Comparative analysis of COVID-19 forecasting models during critical time periods”). Finally, we assessed our model’s ability to anticipate the onset of multiple outbreaks during our studied time periods.
Determining the length of moving time windows for training
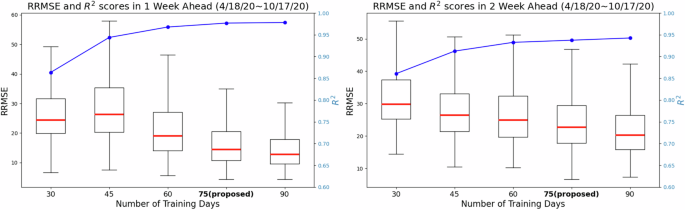
Given that neural network based models typically need a large amount of data to be trained48,49, we first investigated the minimum amount of data that would be necessary for our model to produce robust and reliable forecasts. We investigated the length of the moving window for training necessary to yield forecasts responsive to changes in disease dynamics due to changes in human behavior over time, vaccine availability, different transmission intensities for different variants (e.g. omicron), among other factors. Using observations from April 18th, 2020 to October 17th, 2020 we identified a time window size of 75 days to be a good compromise between reliable forecasting performance (shown in Fig. 2) and a short enough time period that would allow the model to continuously learn new transmission patterns. For more details, please refer to the Discussion.
The figure shows the RRMSE values and R2 scores (represented by the blue line) achieved by the proposed model using different training day lengths from first 6-month dataset (from April to Oct, 2020). For the 1-week horizon, 75-day and 90-day training lengths result in lower RRMSE prediction errors, with a median value below 20 and an approximate R2 score of 0.95. Moreover, increasing the training day length beyond 75 days leads to a 26.3% reduction in RRMSE for 2-week ahead prediction but does not further improve performance with longer training periods. These findings highlight the significant impact of training day length on short-term infection forecasting.
FIGI-Net forecasting performance at the county level
For each forecasting horizon h, we computed FIGI-Net’s prediction error metrics (RMSE and RRMSE) across all counties, over the time period: 10/18/2020 – 4/15/2022. Table 1 shows the prediction error values and percentage of error reduction with respect to the Persistence model of all the models for 1-day, 7-day and 14-day horizons. Based on the experiments regarding training length influence, we evaluated all comparative models using an optimal training length of 75 days in the following eighteen months of data. Our results demonstrate that FIGI-Net model has the greatest error reduction across all horizons (90%, 83%, and 35% RMSE reduction, correspondingly), followed by the biLSTM model (85% in horizon 1-day, 75% in horizon 7-day, and 34% in horizon 14-day RMSE reduction). To assess the significance of these error reductions, we performed the two-sided Wilcoxon signed rank test50 over the entire outcomes of tasks.
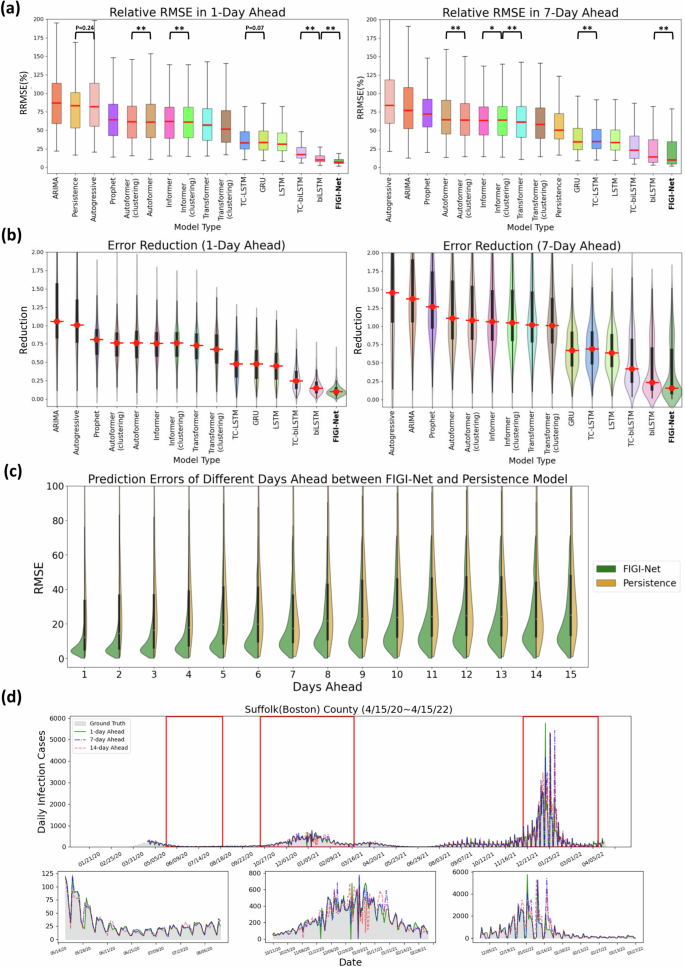
In Fig. 3(a), we visualize the forecasting ability of each model for both the 1-day and 7-day ahead tasks, including classic models such as Persistence (a naive rule stating yt+1 = yt)43, Autoregressive models(AR)51, ARIMA28, and Prophet, as well as deep learning-based models like GRU or LSTM architectures, and Transformer-based models such as Transformer, Autoformer, and Informer. We used the median RMSE score for each model as a means to order them in decreasing order (leftmost model with the worst performance, and rightmost with the best). Our first observation is that FIGI-Net scored the lowest median RMSE and relative RMSE (RRMSE) scores across all models for both the 1-day ahead and the 7-day ahead prediction task (approximately 6.98% of RRMSE at 1-day ahead, and 9.92% at 7-day ahead), followed by deep learning models with bidirectional components (TC-biLSTM with 17.08% and 23.03% of RRMSE and biLSTM with 9.83% and 14.28% of RRMSE, respectively). All recurrent neural network-based models showed improvement over the Persistence model (83.13% and 50.48% of RRMSE), the Autoreggressive model (81.57% and 83.87% of RRMSE), ARIMA (86.68% and 77.13% of RRMSE), and Prophet (64.3% and 71.91% of RRMSE). In Fig. 3(b), FIGI-Net reduced the errors by approximately 90% in the 1-day ahead prediction and by around 83% in the 7-day ahead prediction compared to the Persistence model. Moreover, Transformer-based models exhibit higher RRMSE values in the 1-day, 7-day, and 14-day ahead horizons compared to the recurrent neural network-based models (also shown in Fig. 3(a) and (b)). Transformer models typically require large amounts of data to perform optimally. Consequently, they underperformed in COVID-19 forecasting due to limitations in data granularity and temporal length. Also, COVID-19 data often lacks the seasonality and consistent patterns that Transformer models excel with, making it difficult for them to adapt to sudden changes. Additionally, the shorter data sequences typical in COVID-19 forecasting limited the model’s ability to capture complex patterns, while sensitivity to noise in the data further impacted performance. As a result, recurrent neural network-based models proved more effective for COVID-19 forecasting. A detailed description of the performance of each model is provided in Table 1.
This summary presents a comparative analysis of FIGI-Net’s performance at the county level for both the 1-day and 7-day ahead horizon tasks from Oct. 18th, 2020 to Apr. 15th, 2022. a Performance of each model over all 3143 counties presented as a box plot. The median is highlighted in red along with the 5th and 95th percentile whiskers. The models are ordered in decreasing order, with the most accurate model (lowest RRMSE) appearing on the rightmost side. Notably, FIGI-Net exhibits the lowest RRMSE compared to other models for both tasks, as confirmed by the two-sided Wilcoxon signed rank test. b Error Reduction between each model with Persistence model. Compared to other models, FIGI-Net provides the fewest erroneous forecasting results. c Performance comparison of FIGI-Net against Persistence (our baseline model). Comparison is shown as a set of violin plots across the different time horizons. Our model consistently displays a narrower distribution of prediction errors (RMSE) compared to the Persistence model. d shows daily prediction results of our model in 1-day, 7-day and 14-day horizons in Suffolk county in Massachusetts. The example demonstrates our model’s ability to provide highly accurate predictions for diverse locations and various time periods. *p-value < 0.05, **p-value < 0.01.
Figure 3 (c) focuses on the performance of FIGI-Net against Persistence model. For each time horizon, we generated a violin plot to visualize the RMSE scores of FIGI-Net (in green) and Persistence (in orange) across all counties. Our results show that FIGI-Net has a higher concentration of scores between the 0-20 range across all time-horizons, in comparison to Persistence model, where the scores of the orange distribution are widely spread. The main error difference between FIGI-Net and Persistence occur at the horizon 1, with a mean RMSE score of 9.97, in comparison to 79 from Persistence (a 89% reduction).
Despite the high variability in the data, the Wilcoxon signed-rank test comparing paired observations from the same counties under the two different models enabled us to detect significant differences. To further support our statistical comparisons between FIGI-Net and biLSTM (Supplementary Fig. 2 and Supplementary Fig. 5(a)), we conducted supplementary analyses using Cliff’s Delta52 and bootstrap53 approaches to address variability concerns in RMSE evaluation. Cliff’s Delta is a non-parametric effect size measure that quantifies the overlap between two distributions without assuming normality. In parallel, the bootstrap method generates resampled distributions of our error metrics, providing robust estimates of uncertainty and confidence intervals, even with skewed or limited datasets.
We performed an effect size analysis using Cliff’s Delta with 5,000 bootstrapping iterations, as shown in Supplementary Fig. 5(b). The results indicated Cliff’s Delta values of 0.224 (95% CI [0.196, 0.251]) for the 1-day forecast and 0.109 (95% CI [0.081, 0.139]) for the 7-day forecast, suggesting small but consistent effect sizes (ranging from 0.1 to 0.3). Notably, the RMSE of biLSTM was larger than that of FIGI-Net. Additionally, to reinforce our two-sided Wilcoxon signed-rank test findings, we performed 5000 bootstrap iterations by subtracting the RMSE of biLSTM from that of FIGI-Net, as shown in Supplementary Fig. 5(c). The results confirmed that the differences between FIGI-Net and biLSTM are statistically significant (p < 0.01), demonstrating the reliability of our analysis despite large standard deviations. Collectively, these analyses confirmed the robustness and reliability of FIGI-Net’s short-term forecasting performance, showing statistically significant results and small yet meaningful effect sizes.
For a more detailed comparison in short-term forecasting, we also evaluated the proposed model alongside other established models for 1-day and 7-day horizons during the Omicron outbreak. The results, presented in Supplementary Fig. 10 and Supplementary Table 6, demonstrate that FIGI-Net model efficiently handles short-term dynamic changes and achieves approximately a 59% reduction in prediction error. These results underscore the robustness and adaptability of our model in handling rapid and dynamic changes in infection trends, as was characteristic of the Omicron outbreak.
Figure 3 (d) showcases a visualization of the forecasts of our model for the county of Suffolk, Massachusetts, for the 1-day, 7-day and 14-day ahead tasks. Additionally to the full time period of the experiment, three periods from May to August 2020, October 2020 to February 2021, and December 2021 to March 2022, are also displayed. FIGI-Net accurately forecasted the daily infection trends in 1-day and 7-day ahead horizons, even when this county exhibited contrasting infection trends during the initial outbreak stages. However, we observed that the 14-day ahead forecasting trend yielded larger errors, particularly in cases where the infection numbers fluctuated significantly.
Forecasting performance at the state level
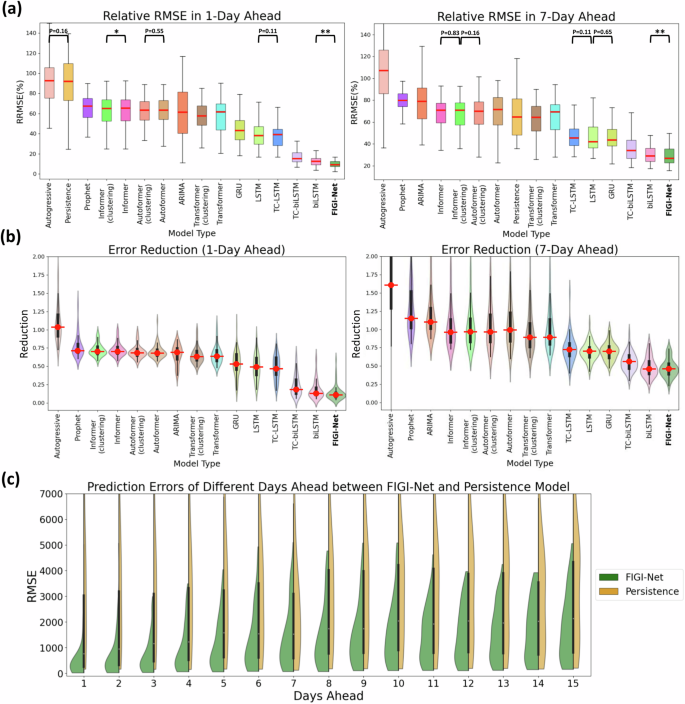
Similar to our county level experiments, we compared the performance of FIGI-Net against state-of-the-art models for the state-level geographical resolution. For FIGI-Net, which is a model that leverages high volumes county-level activity as part of its design, we decided to aggregate the county level forecasts into state level (details regarding the performance of FIGI-Net, trained on state-level data, can be found in the Supplementary Table 1). For other deep-learning based models, we adopted the same strategy as FIGI-Net to generate state-level forecasts, as these models require a sufficient amount of data. Additionally, for the rest of the models, we trained them using state-level data to obtain state-level outcomes. Then, we computed the RMSE and RRMSE for each state across 1-day, 7-day, and 14-day ahead horizons, as shown in Fig. 5(a). Our results, also shown in Fig. 4 and summarized in Table 2, show that FIGI-Net was able to score 1149.66 ± 1850.66 and 1935.36 ± 3458.21 in terms of RMSE for the 7-day and 14-day horizon, correspondingly. On the other hand, the score of the Persistence model was 2571.9 ± 5409.64 and 3330.82 ± 6072.91 (resulting in an error reduction of 54%, and 39% in each case). Next to FIGI-Net, we can see the biLSTM (with a score of 1198.43 ± 1806.38 and 1940.18 ± 3216.91) and TC-biLSTM models (1338.63 ± 2324.23 and 2225.14 ± 4116.09 in RMSE).
a At state level, FIGI-Net still exhibits the lowest RRMSE compared to other models for the 1-day and 7-day ahead horizon tasks. b Error Reduction at state level displays that FIGI-Net provides lower forecasting errors than other models and reduced errors by at least 53% compared to the Persistence model. c Performance comparison of FIGI-Net against Persistence at state level. This comparison also represents that our model consistently displays a much narrower distribution of prediction errors (RMSE) and provides much lower forecasting errors during the first four days. **:p-value < 0.01.
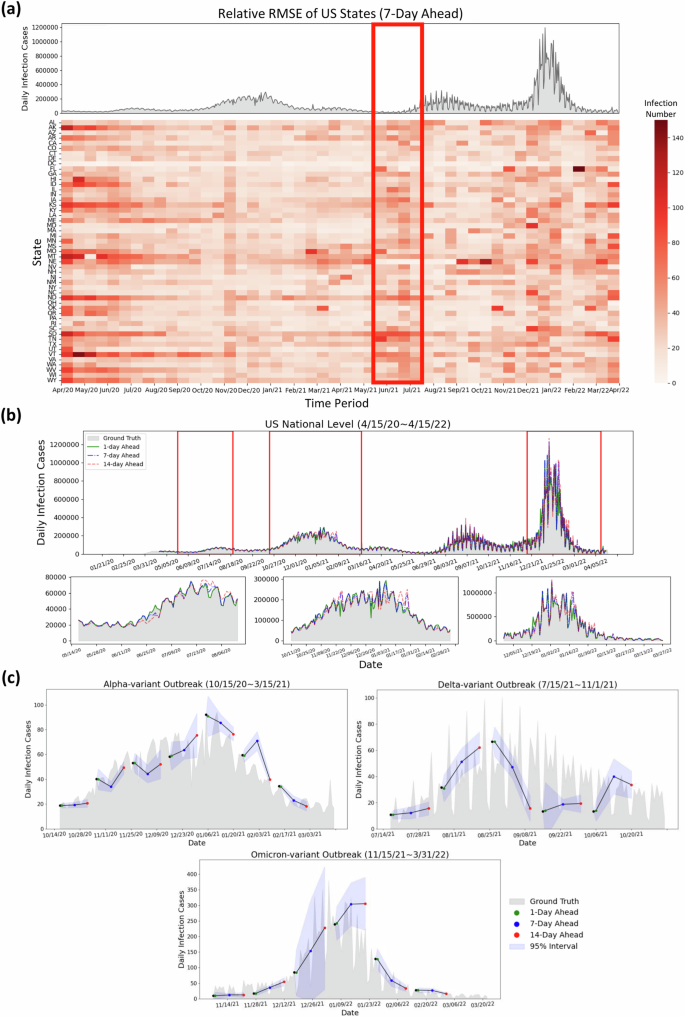
During the early stages of the pandemic, we observe the RRMSE values of FIGI-Net were notably higher. For instance, based on Fig. 5(a), Missouri, Montana, and Nevada displayed larger prediction errors between March to May 2021, as shown from the deep red color in the center of the RRMSE matrix. During this period, these states experienced a significant spike in COVID-19 activity, deviating from both previous months and the overall national trend. Alternatively, the red rectangle in Fig. 5(a) illustrates that the RRMSE errors often increase before the outbreaks or when the infection trend rapidly increases. This observation suggests that the proposed FIGI-Net model may provide early prediction outcomes for outbreaks. Similar trends are also evident in other day-ahead forecasting instances (shown in Supplementary Fig. 6).
a Relative RMSE performance among US states in the 7-day horizon at the state level is determined by the average relative RMSE of the last 7 days of each time period, compared to the national reported infection trend. The relative RMSE increased during the early period (April 2020 to May 2020) of the first upward trend, the time period (June 2021 to July 2021) before the Delta outbreak, and the increasing period (December 2021 to January 2022) of the Omicron COVID-19 outbreak. Missouri, Montana, and Nebraska have large relative RMSE values during March 2021 to May 2021. We can observe that the RRMSE errors often increase before the early stage of the next outbreaks or when the infection trend rapidly increases (red rectangle) (b) Daily prediction infection trends during different days ahead at the national level. It can be observed that the daily predicted infection trends at 1-day and 7-day ahead horizons show similarity to the reported data, while the 14-day ahead trend exhibits some fluctuations. c Daily predicted infection trends of the proposed model during the Alpha, Delta, and Omicron variant outbreaks with multiple day-ahead predictions. The proposed FIGI-Net model can provide a curve of predicted trends matching the observed report. The range of margin of error became larger when the trend of Omicron-variant outbreak increased.
National forecasting performance
Figure 5 (b) represents national level COVID-19 official reports contrasted with our model predictions across three different horizons. Similar to the observations in Fig. 3(c), our predictions were highly accurate at the 1-day and 7-day ahead horizons, with the RRMSE of 7.02% and 15.47%, respectively, compared to the national official reports. However, at the 14-day horizon, the discrepancies between predicted values and ground truth grew during high infection periods, resulting in an RRMSE of 27.94%. Table 3 shows the performance between FIGI-Net and other models at national level. Compared to our benchmark models, FIGI-Net improved forecast capacity can lead to a 86.5% reduction at 1-day horizon, a 60.98% reduction at 7-day horizon, and a 53.8% reduction at 14-day horizon in RRMSE score.
Additionally, we analyzed the daily performance of FIGI-Net at the national level across three main outbreak waves: the Alpha-variant wave (October 15th, 2020 to March 15th, 2021), Delta-variant wave (July 15th, 2021 to November 1st, 2021), and Omicron-variant wave (November 15th, 2021 to March 31st, 2022). By aggregating our county-level forecasts, we depicted the national prediction trends during these waves (Fig. 5(c)). Our model demonstrated an accurate prediction direction, maintaining an average error range of 12.17 infection cases at a confidence level of 95% during the Alpha and Delta-variant waves. During the Omicron-variant wave, the forecasting trend exhibited a wide range of confidence intervals. However, our model still successfully predicted trends up to the 14-day horizon. These above outcomes underscore the robustness of our FIGI-Net model in addressing substantial variations in infection numbers.
Geographical distribution of COVID-19 infection predictions of US counties
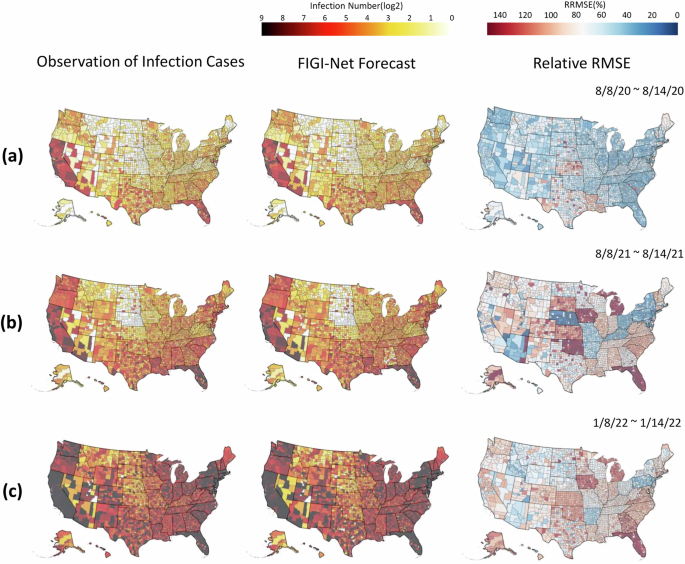
We conducted a geo-spatial analysis with the objective to identify possible geographical patterns in the performance of FIGI-Net. Figure 6 illustrates various geographical maps of US counties. The first column of Sections (a), (b), and (c) presents the average number of cases occurring for 3 distinct 1-week periods: the first halves of August 2020, August 2021, and January 2022 (periods just before the Alpha, Delta, and Omicron-variant waves began, respectively). With the objective to demonstrate the forecasting capacity of FIGI-Net, the second column presents the forecast average over the same time periods. Finally, the last column shows the relative RMSE incurred by FIGI-Net. FIGI-Net successfully predicted the COVID-19 activity shown in the 7-day ahead prediction maps compared to the observation maps. However, some counties had larger errors, as observed from the relative RMSE maps (the last column of Fig. 6). During these time periods, counties in Kansas and Louisiana displayed higher errors despite low or mild infection levels. Moreover, the counties in Oklahoma, Iowa, Michigan, and Florida exhibited larger prediction errors preceding the Delta-variant wave. Particularly in Florida, while the epidemic situation was severe, errors increased. Our model demonstrated higher accuracy and lower RRMSE values in the west coast and northeast regions during these time periods (see, for example, the third column of Fig. 6, while mid-west and south regions of the U.S. tended to display higher errors as the pandemic progressed (refer to Supplementary Movie 1 for the details of geographical distribution prediction and error maps in 1-day, 7-day, and 14-day ahead across all time periods from April 2020 to April 2022).
The figure shows the infection prediction distribution of the proposed model during different time periods at the county level. The early half of August 2020, August 2021, and January 2022 are represented in (a), (b), and (c), respectively. The first column of (a), (b), and (c) show the observed daily confirmed cases. The second column represent the 7-day (1-week) ahead prediction results, and the corresponding RRMSE score maps are shown in the last column. The proposed model provided the predictions that are similar to the observed daily reports of infection cases and had low RRMSE values in most counties. However, it had higher RRMSE values in the counties where the number of infection cases rapidly increased. Additionally, our model indicated higher RRMSE values in the counties of Kansas and Louisiana at these time periods. The counties in Michigan and Florida represented much higher RRMSE values during time periods (b) and (c) when the status of infections in these two states were severe.
Comparison between FIGI-Net and the CDC Ensemble Model in COVID-19 Forecasts
To further evaluate our approach, we compared the performance of our proposed FIGI-Net model with the COVIDhub_ensemble model54 (also known as the CDC model). The CDC model employs an ensemble methodology that combines the output of several disease surveillance teams across the United States, generating forecasts for the number of COVID-19 infections at the county, state and national levels. We collected the 1-week and 2-week ahead forecasts of the CDC model at county and state level, and compared them against FIGI-Net’s forecasts. Given the CDC model is an aggregated forecast (i.e. the total number of reported activity over the next 7 and 14 days, rather than a daily forecast over the same periods), we aggregated our daily predictions for the 1 to 7-day and 1 to 14-day horizons to facilitate a fair comparison between both models. The Persistence model of this task was also included as baseline.
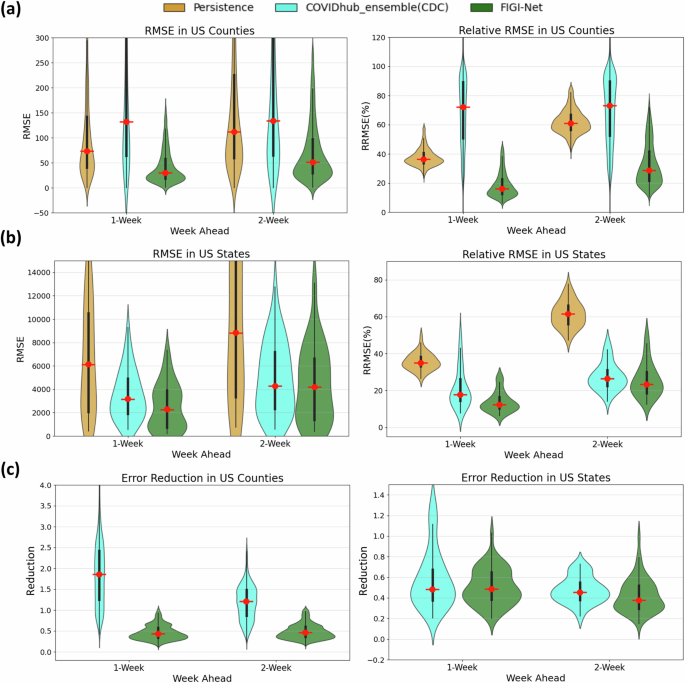
Shown in Fig. 7(a), the CDC model exhibited significantly higher average RMSE and RRMSE values at the county level compared to our FIGI-Net model. Our model achieved an approximate reduction of 58.5% in averaged RMSE and 53.28% in averaged RRMSE over the 1 and 2-week ahead horizons, respectively (see Table 4). At the state level (Fig. 7(b)), our proposed model consistently maintained the averaged reduction of 64.55% RMSE value and 64.48% RRMSE value (compared to the CDC ensemble and Persistence models, as shown in Table 5). Notably, the CDC model demonstrated better performance than the Persistence model in terms of lower error predictions for both 1 and 2-week horizons.
a The RMSE and relative RMSE values of the three models at the county level. b The comparison of prediction errors among these three models at the state level. Our proposed FIGI-Net model outperforms the Persistence model and CDC model in terms of lower prediction RRMSE errors. This indicates its enhanced capability to capture the complex dynamics of COVID-19 infection spread, with approximate 4.76% averaged reduction in errors observed across various prediction horizons. c The error reduction comparison between the CDC model and FIGI-Net. At the county level, FIGI-Net outperforms the CDC model, with errors approximately 58.5% lower than those of the Persistence model. At the state level, FIGI-Net continues to provide a 13% lower error reduction compared to the CDC model.
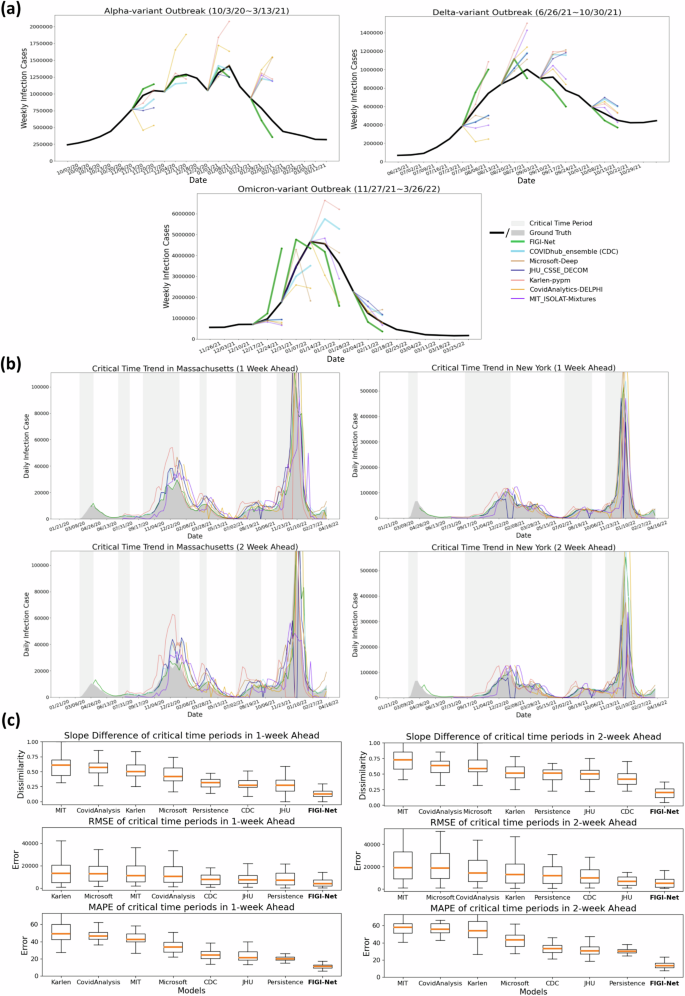
We also conducted comparative analyses between the COVID-19 forecasts of FIGI-Net with forecasting models officially reported on the CDC website54 (we selected models, including Microsoft-Deep, JHU_CSSE_DECOM, Karlen-pypm, CovidAnalytics-DELPHI, and MIT_ISOLAT-Mixtures, that provided sufficient infection prediction outcomes for 1-week and 2-week ahead horizons). Following the CDC weekly reporting criteria55, we aggregated the daily prediction cases into weekly prediction values. Specifically, we focused on three reported infection wave time periods and presented 1-week and 2-week prediction results of our model at the national level alongside those of other models (Fig. 8(a)). Our findings revealed that most models can accurately predict infection numbers during decreasing trends, but struggle to forecast the correct trend direction and COVID-19 official infection numbers during increasing trends. Interestingly, our FIGI-Net model correctly predicted the increasing direction of the infection trend before the outbreak wave began, from November 2021 to March 2023. Fig. 8b illustrates the examples of the predicted infection trends in Massachusetts and New York states by our FIGI-Net model and other forecasting models in 1-week and 2-week horizons. The results showed that the our model’s infection prediction trends are much closer to the reported data across different weeks ahead at the state level, and most of other models predicted accurate infection numbers when the infection trend increases. It is important to note that some models did not provide outcomes at some weekly time points, leading to 0 values in those models, which were subsequently removed in further comparison and analysis.
a Comparison of the forecasting results among different forecasting models during three different outbreak time periods at the national level. b Examples of COVID-19 infection prediction trends of different state-of-the-art forecasting models at the state level. The critical time period, which indicates a significant increase in COVID-19 infections, is highlighted in light grey color. c Performance evaluation of the forecasting methods during the critical time periods of COVID-19 infection in 1-week and 2-week horizons across the states. Slope difference, RRMSE, and MAPE were measured to assess the prediction number and trend accuracy of each model. Our proposed FIGI-Net model provided lower prediction errors in both 1-week and 2-week horizons during the critical time and may efficiently forecast the infection number and trend direction before the severe transmission of COVID-19. Here we also ranked them from high to low evaluation or error values according to the median values.
Comparative analysis of COVID-19 forecasting models during critical time periods
Given that identifying the beginning of a major outbreak is a crucial task in disease forecasting, we assessed the performance of our FIGI-Net model in early COVID-19 prevention and forecasting by measuring its ability to anticipate critical time periods marked by exponential growth of COVID-19 infection cases. We compared our model’s performance with other state-of-the-art COVID-19 forecasting models during this critical time period, using weekly data from July 20th, 2020 to April 11th, 2022. First, we identified these “critical” time periods as periods where the trend λ of COVID-19 activity (estimated as the coefficient of a linear model yt = λyt−1) remained above 1, indicating a sustained multiplicative growth for an extended period (Supplementary Fig. 7 to 9 for more details). Examples of such periods for Massachusetts and New York states are shown in Fig. 8(b). We compared the ground truth and predicted results of the models using slope similarity, RMSE, and MAPE values (see the equation (5) in the section “Model evaluation” for a definition to calculate slope similarity). Our FIGI-Net model efficiently and accurately predict infection case numbers and trend direction for 1-week and 2-week horizons during critical time periods (Fig. 8(c)). Table 6 and 7 presented the forecasting performance details among the prediction models and the statistical significance between our model and the others. Comparing with Persistence model, FIGI-Net model improved the RMSE value by at least 43% reduction in 1-week ahead and at least 57% reduction in RMSE in 2-week ahead forecasts. Additionally, our model achieved at least a 45% MAPE reduction in both week horizons and exhibited around 83% similarity in the slope of infection trend. These results indicate that FIGI-Net model can effectively adapt and refine forecasting trends when the pandemic intensities suddenly.