Study site
Brazil is a continental-sized country with 203.1 million inhabitants, according to the 2022 census26. The country has 27 federative units (26 states and one federal district) with diverse population distribution, varying income levels, differing human development indices, and contrasting prevalence rates of arbovirus and respiratory diseases. Historically, arboviral diseases have been more common in tropical and equatorial regions, while respiratory diseases27 are more prevalent in the subtropical region28. Brazil’s climate is generally categorized into equatorial, tropical, semi-arid, highland tropical, and subtropical zones29, each influencing disease dynamics. The northern equatorial region, marked by high temperatures and humidity, creates ideal conditions for mosquito breeding, which contributes to the ongoing transmission of arboviruses like dengue and chikungunya30. Semi-arid areas in the northeast, characterized by extended droughts, may limit mosquito proliferation but still face arboviral outbreaks during the wetter seasons, especially in populated areas31. Recent temperature increases caused by climate change have been favoring the emergence of arboviral outbreaks in southern regions of Brazil32. Additionally, in regions with seasonal temperature variations, characterized by colder and drier months, there is a periodic increase in respiratory infections, including influenza or SARI caused by other viruses27,33,34,35.
Data sources
Our dataset integrates diseases and associated symptoms data from official public sources with Google search results, organized by the corresponding epidemiological week. Both data sources and data extraction methods are detailed below.
Official public sources (diseases and symptoms)
Dengue and chikungunya cases were obtained directly from the Brazilian Notifiable Diseases Information System (Sistema de Informação de Agravos de Notificação, SINAN, ftp://ftp.datasus.gov.br/dissemin/publicos/SINAN/), Brazil’s main platform for recording and monitoring cases of notifiable diseases. It employs standardized forms for each disease that collect data related to patient identification and the primary characteristics of the illness. Established in the 1990s, SINAN was created as part of a government initiative to improve public health surveillance. In 1998, it became mandatory for municipalities and states to report notifiable diseases across the country36. Over the years, SINAN has broadened its scope to encompass a broader range of diseases, reflecting changing public health priorities and the rise of new epidemics. Dengue cases have been reported to SINAN since 200137, while chikungunya was included in 201738.
SINAN data is publicly accessible through the file transfer protocol (FTP) service maintained by Brazil’s Unified Health System data platform (DATASUS). Dengue notifications are stored in the SINAN-Dengue database, while chikungunya notifications are available in the SINAN-CHIK database. Data extraction was performed using the R package microdatasus39, which facilitates access to SINAN records.
In the SINAN system, all suspected cases are notified according to the case definition for suspected cases established by the Pan American Health Organization (PAHO)40, which is based on clinical presentation. In our database, we included cases classified as probable, plausible, or confirmed, which encompass both laboratory-confirmed cases and those confirmed through clinical and epidemiological criteria. Discarded cases were excluded from the dataset. Case definitions for dengue and chikungunya follow PAHO guidelines. For dengue, symptoms should include a fever lasting 2 to 7 days, accompanied by at least two of the following: nausea, vomiting, rash, myalgia, headache, retro-orbital pain, petechiae, a positive tourniquet test, or leukopenia. For chikungunya, a suspected case is defined by the sudden onset of fever and severe arthritis or arthralgia that cannot be attributed to other conditions. For both diseases, it is required that individuals have either lived in or traveled to endemic or epidemic regions within 14 days prior to symptom onset, or who have an epidemiological link to a confirmed imported case41. Since the SINAN notification includes data on symptoms as defined in the notification form, these reported symptoms were also extracted. The variables selected from the SINAN database are detailed in Table 1.
Data on Severe Acute Respiratory Infections (SARI)—caused by influenza, COVID-19, or other viruses— were obtained from the Influenza Epidemiological Surveillance Information System (SIVEP-Gripe) and downloaded directly from OpenDataSUS42, the Brazilian Ministry of Health’s open data portal. The respective datasets can be located by entering ‘SRAG’ in the portal’s search field. Unlike the reporting of arboviruses to SINAN, only hospitalized SARI cases are reported to this system.
SIVEP-Gripe is Brazil’s primary surveillance system for SARI cases. Established in 2000, it began reporting SARI cases in 2009 as part of the response to the H1N1 influenza pandemic. In 2020, in response to the COVID-19 pandemic, the system was further adapted to monitor the concurrent circulation of SARS-CoV-2, influenza, and other respiratory viruses43. Since then, the mandatory notification of all suspected or confirmed COVID-19 cases that meet the criteria for SARI has been incorporated into SIVEP-Gripe, ensuring comprehensive tracking of severe respiratory infections during public health emergencies.
Severe acute respiratory infection (SARI) is defined as an individual with ILI who presents with dyspnea, respiratory distress, persistent chest pressure, oxygen saturation ≤94% in room air, or bluish discoloration of the lips or face44. ILI is characterized by fever accompanied by cough or sore throat, with symptom onset within the last seven days45. In contrast, under the framework of universal COVID-19 surveillance, ILI involves an acute respiratory condition defined by at least two of the following signs and symptoms: fever, chills, sore throat, headache, cough, runny nose, or disturbances in smell and taste senses. In children, nasal congestion is also considered in the absence of another specific diagnosis. In elderly individuals, additional criteria for severity should be considered, such as syncope, mental confusion, excessive drowsiness, irritability, and loss of appetite43. For suspected COVID-19 cases, fever may be absent, and gastrointestinal symptoms (such as diarrhea) may be present.
The SIVEP-Gripe dataset has been available through Opendatasus since 2009, which defines the period covered in our database. Similar to the SINAN system, a predefined list of reported symptoms is entered during the notification process and made available; therefore, we included them in our data extraction. The selected variables, along with the year they were added to the database, are detailed in Table 2. In 2019, variables for “vomiting” and “positive PCR for influenza” were added. Beginning in 2020, fields related to COVID-19 symptoms, PCR and antigen tests for COVID-19, and antigen tests for influenza were incorporated.
Since SIVEP-Gripe records only severe cases of respiratory diseases that require hospitalization, milder cases are reported in another system, e-SUS Notifica. However, the absence of mandatory reporting and challenges related to data cleaning and standardization on this database makes it highly susceptible to notification bias and underreporting. As a result, e-SUS Notifica is not suitable for accurately tracking the total number of influenza cases in the country. In contrast, the mandatory reporting of severe cases makes SIVEP-Gripe the most reliable system for monitoring respiratory disease cases in Brazil.
Web Search Data (diseases and symptoms)
Data on public interest in diseases and symptoms was obtained from Google Trends through its API. We extracted the Google Trends search index, Related Topics and Related Queries. Data extraction was performed using the gtrendsAPI package in R46, which facilitates direct access to Google Trends data.
The Google Trends search index measures the relative popularity of a search term over time and a geographical region. It ranges from 0 to 100, with 100 representing peak interest during the specified period and 0 indicating minimal or no search activity. This index does not reflect absolute search volumes; rather, it compares the search interest for a given term to its peak within the selected timeframe. The search index is normalized to account for variations in total search volumes across different regions and time periods. It is calculated by dividing the number of searches for a specific term by the total number of Google searches in the same location and timeframe. This normalization ensures that the index reflects relative interest, preventing larger regions or those with higher search volumes from skewing the data. As a result, it allows for meaningful comparisons across geographic areas and time periods, emphasizing fluctuations in public interest instead of absolute search metrics numbers.
Characteristic symptoms for chikungunya, dengue, COVID-19, influenza, and Severe Acute Respiratory Infections (SARI) were identified based on a consultation of official documents and scientific publications, including guidelines from the Brazilian Ministry of Health45,47, the World Health Organization (WHO)48,49, the Pan American Health Organization (PAHO)50, and selected peer-reviewed articles51. Symptom selection was guided by clinical distinctiveness and reporting frequency, prioritizing those most indicative of each disease. The most characteristic symptoms and their associated diseases are presented in Table 3.
To broaden the scope of search behavior analysis and account for local linguistic diversity, we also compiled a list of alternative expressions used to describe each symptom, including popular and regional terms commonly used throughout Brazil. Additionally, Google Trends distinguishes between searches for specific “terms” and broader “topics”. A search by “term” retrieves data based on exact keywords, while a “topic” search encompasses related terms and variations, including misspellings and regional differences. After identifying relevant search topics, we used their corresponding Freebase ID codes from Wikibase whenever possible, to include variations in spelling and language. These unique identifiers help disambiguate search terms, ensuring that queries remain focused on the intended diseases and symptoms while excluding unrelated meanings. For instance, the topic “Fever” (symptom) has the Freebase ID “/m/01s08g”, distinguishing it from non-clinical uses, such as in music or entertainment. A complete list of topics, search terms, Freebase ID codes, and their variations used in this study is presented at the Table 4.
The Google Trends search index data were extracted at the federative unit level, the highest geographic resolution available through the platform. The dataset covers the period from 2019 to 2024, as a five-year range is the maximum timespan for which the API provides weekly search index values. Although longer periods can be requested, the API returns data at a monthly resolution, which is not aligned with the epidemiological week format used in our official disease records. Additionally, because Google Trends normalizes the index within each extraction, combining data from different time windows can result in scale inconsistencies. To address this issue, values must be rescaled to ensure comparability. We present a normalization method for aligning time series across extractions, as detailed in Usage Notes section.
Alongside the search index for each query topic, we extracted additional data from the Related Topics and Related Queries sections available through the Google Trends platform. These features provide insights into user behavior by identifying topics and queries commonly searched in association with the original term. Related Topics refer to broader subjects linked to the query, while Related Queries capture specific search phrases entered by users. Both are ranked using two metrics: “Top” and “Rising”. Top indicates the most frequently searched items on a relative scale (where 100 represents the highest popularity). Rising highlights items with the greatest increase in search frequency over the preceding period. For each disease (dengue, chikungunya, influenza, and COVID-19) we collected Related Topics and Related Queries sorted by the Top metric, on a monthly basis, for each federative unit and for Brazil as a whole, covering the period from 2020 to 2024.
Data processing
Official public sources
For all diseases, we collected the report date, notification date, symptom onset date, federative unit of notification, and clinical symptoms. Although the data extracted from public databases are individual-level records, by the end of the processing workflow, we present the data as aggregated counts of cases and symptoms, grouped by epidemiological week and federative unit. After extracting the datasets, we selected variables of interest based on their availability within each dataset and across the years covered. For the SINAN dataset, we calculated the difference in days between the notification date and the symptom onset date. Records in which this difference exceeded 180 days were excluded to reduce distortions caused by errors, such as data entry errors or incorrect patient date of birth entries in the symptom onset field. Both the symptom onset and notification dates were converted to their corresponding epidemiological week start dates. An epidemiological week is a standardized time unit used in public health surveillance, typically defined as a seven-day period that starts on Sunday and ends on Saturday, facilitating consistent reporting and analysis of disease trends across different regions’ timeframes.
Symptom data were included starting in 2013 for dengue and in 2017 for chikungunya, reflecting the addition of these variables in notification forms only from those years onward. For symptom-related variables, we applied a standardization rule in which all values except for “1” (indicating “Yes”) were converted to missing values (NA). Finally, the data were aggregated by epidemiological week (based on symptom onset, data entry, and notification dates), federative unit, and final case classification. For each combination, we calculated the total number of dengue and chikungunya cases, as well as the number of cases in which each symptom was reported. Note that each case may or may not include symptom information; thus, symptom counts reflect the number of cases reporting each symptom within each group.
For dengue, we included all cases reported in the SINAN-DENGUE system, excluding those classified as “discarded,” i.e., cases with a final classification different from code “5”. For chikungunya, we included all cases reported in the SINAN-CHIK database that were confirmed as chikungunya, identified by a final classification code equal to “13”. For the SIVEP data, cases with a final classification of “2” (SARI due to another respiratory virus) or “3” (SARI due to another etiological agent) were grouped into a single category, while values of “9” were treated as “Not Available” (NA). For COVID-19 case counts, we considered SIVEP records with a final classification of “5” (SARI due to COVID-19), or those that tested positive via PCR or antigen tests for SARS-CoV-2. For influenza case counts, we considered records with a final classification of “2,” or those that tested positive via PCR or antigen tests for influenza. In instances where SARI cases tested positive for both COVID-19 and influenza, a new final classification value of “0” was assigned to indicate dual infection. Similar to the SINAN dataset, for symptom variables, any values other than “1” (indicating “Yes”) were converted to NA. This process resulted in the creation of three intermediate tables — SINAN_dengue_cases, SINAN_chik_cases, and SIVEP_cases — all of which are available in our data repository. We then combined the case counts from both data sources (SINAN and SIVEP) into a single final dataset that provides disease case counts aggregated by: (i) epidemiological week of symptom onset, and (ii) federative unit. This final aggregated table is referred to as the Arbo_SARI_disease_table in the data repository.
For each epidemiological week and federative unit, we counted the number of cases for the following categories: dengue, chikungunya, COVID-19, influenza, SARI caused by other respiratory viruses, SARI cases with no etiological agent information available (i.e., final classification or PCR and antigen test results unavailable), and the total number of SARI cases. The criteria for classifying cases in each disease category are summarized in Table 5. For dengue and chikungunya cases, we included both confirmed and suspected cases, excluding records with a final classification of “5” (discarded) or NA (Not Available). In the case of SARI, when a single record tested positive for both influenza and COVID-19, it was included in the time series counts for each disease but counted only once in the “total SARI cases” series. For diseases with data outside the available time frame in the respective dataset, the value is recorded as NA.
Web search data
After extracting interest indexes for all search terms from Google Trends, we organized the data aggregated by week and federative unit into a table, which is available in our data repository under the name GoogleTrends_search.
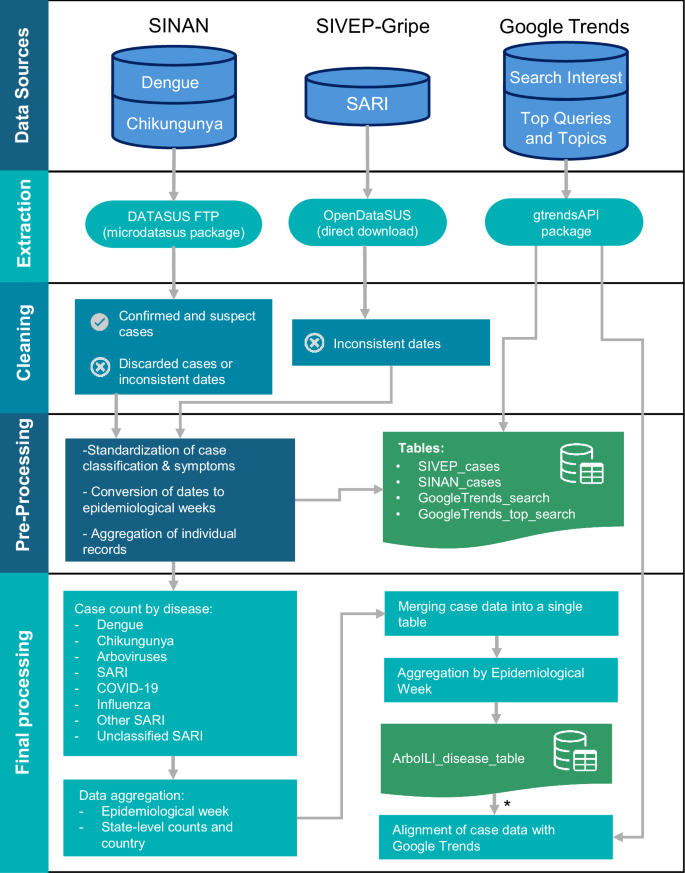
To identify patterns related to public interest in symptoms, we cross-referenced the Relate Topics associated from each disease (dengue, chikungunya, influenza, and COVID-19) with a predefined list of key symptoms for each disease, as outlined in Table 3. We also included topics that explicitly contained the word “symptom” (in Portuguese, “Sintoma”). Based on this approach, we created a binary variable, key_symptom, which indicates whether a given topic in the Related Topic database was related to the characteristic symptom of the corresponding disease. This information is stored in the GoogleTrends_related_topic table in the data repository. The corresponding data on Related Queries are available in the GoogleTrends_related_query table. A schematic overview of the data collection and integration process is presented in Fig. 2.