Chronic STm infection triggers a robust CCL11 response in the MLN
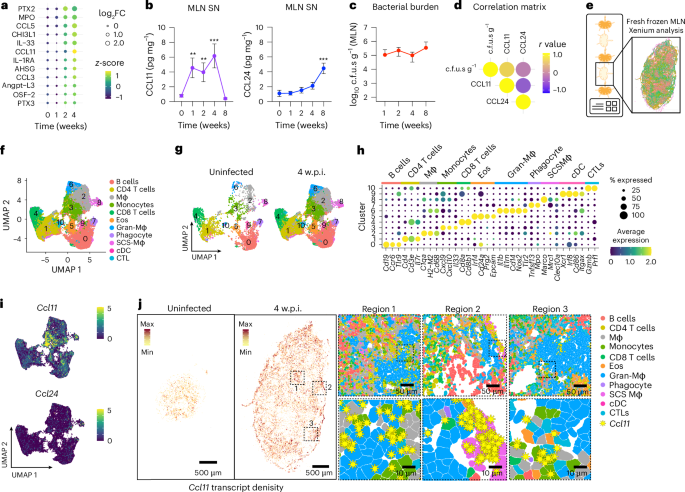
MLN serve as a major reservoir of persistent STm colonization5, although the underlying mechanisms remain elusive. To better understand how host responses both control and permit STm survival over prolonged periods, we orally infected 129×1/SvJ (129 WT) mice and monitored cytokines, chemokines and soluble mediators in MLN supernatants across 4 weeks (Fig. 1a and Extended Data Fig. 1a). Analysis using a broad cytokine array detected minimal changes in analytes during the first week post infection (w.p.i) compared with uninfected controls. By 2 weeks, however, significant elevations were noted in pro-inflammatory mediators linked to neutrophil responses, such as MPO, PTX2 and CHI3L1, with further increases in pro-inflammatory cytokines/proteins, chemokines and proteins involved in tissue repair and angiogenesis by 4 w.p.i. (Fig. 1a and Extended Data Fig. 1a). Interestingly, levels of eosinophil-related mediators of chemotaxis and activation, such as CCL11, CCL5 and IL-33, increased by 4 w.p.i.24,25 (Fig. 1a and Extended Data Fig. 1a). MLN levels of CCL11 peaked at 4 w.p.i relative to baseline and declined by 8 w.p.i., and this increase correlated with higher MLN bacterial loads. Levels of CCL24, another eosinophil chemoattractant, did not rise during the first 4 weeks but showed a modest increase at 8 w.p.i. (Fig. 1b–d). In the spleen, CCL11 remained at baseline despite substantial STm burdens, whereas CCL24 rose slightly by 4 w.p.i, indicating that CCL11 induction is MLN specific (Extended Data Fig. 1c–e). Together, these data reveal a localized cytokine programme that probably drives eosinophil recruitment during persistent MLN infection.
129×1/SvJ mice were orally infected with STm SL1344 (108 c.f.u.s) and monitored for 8 weeks. a, Heat map of significantly regulated soluble mediators in MLN supernatants at 1, 2 and 4 w.p.i. compared to uninfected controls (pooled samples from 5 mice per group across 2 experiments; n = 10 per condition). b, ELISA of CCL11 and CCL24 in MLN supernatants at 1, 2, 4 and 8 w.p.i. compared to uninfected controls (n = 8–10 mice per time point, 2 experiments). c, Bacterial burden in the MLN at 1, 2, 4 and 8 w.p.i. (n = 14 mice per time point). d, Correlation matrix between MLN bacterial levels and MLN tissue levels of CCL11 and CCL24. Data from mice at 1, 2, 4 and 8 w.p.i. (n = 10 mice per time point). e, Schematic representation of Xenium single-cell in situ analysis of MLN tissue sections from uninfected controls and mice at 4 w.p.i (n = 2–8 mice per group, 3 experiments). f, UMAP plot of MLN immune cells from Xenium single-cell in situ analysis in uninfected controls and mice at 4 w.p.i.: cluster 0 (salmon), cluster 1 (beige), cluster 2 (grey), cluster 3 (olive green), cluster 4 (high green), cluster 5 (orange), cluster 6 (light blue), cluster 7 (purple), cluster 8 (pink), cluster 9 (magenta) and cluster 10 (cyan). g, UMAP plot of clusters 0–10 represented in uninfected controls and mice at 4 w.p.i. h, Heat map of top 1–5 differentially expressed genes within each cluster in f (cut-off: log2FC = 1.41, P < 0.05). i, UMAP plots showing total Ccl11 and Ccl24 expression in uninfected controls and mice at 4 w.p.i. j, Ccl11 expression levels in the MLN of uninfected controls and mice at 4 w.p.i. Ccl11 enriched regions are outlined, segmented cells are pseudocoloured according to clusters in f, and yellow symbols indicate Ccl11 transcripts.Data are presented as mean ± s.e.m. All analyses are two-tailed and data were analysed using Student’s t-test (a), Kruskal–Wallis with Dunn’s post hoc test (b), Pearson correlation (d) and Wilcoxon rank-sum test (h). Exact P values in Supplementary Table 1.**P < 0.01, ***P < 0.001.
Spatial transcriptomics reveals discrete macrophage profiles and Ccl11 sources in chronically infected MLN
Xenium single-cell in situ profiling of MLN from uninfected and STm-infected mice at 4 w.p.i. (Fig. 1e,f) identified 11 distinct UMAP clusters, with distinct differences between uninfected controls and STm-infected mice at 4 w.p.i. (Fig. 1g). Major lymphocyte clusters defining B cells (Cluster of differentiation [Cd] 19, Ccr6 [C-C chemokine receptor type 6] and Tlr9 [Toll-like receptor 9]) and CD4 T cells (Cd3 epsilon, Cd4 and Il-7r [Interleukin-7 receptor]) were present in cluster 0 and cluster 1. In addition, cluster 4 expressed markers for CD8 T cells (Cd8a, Cd8b1) and cluster 10 expressed markers for cytotoxic lymphocytes (for example, Prf1 [Perforin-1] and Gzmb [Granzyme B]). Seven additional clusters exhibited myeloid signatures. Cluster 6 showed increased expression of M1-like macrophage markers (for example, Il-1b [Interleukin-1 beta], Nos2 [Nitric oxide synthase 2] and Tlr2), while cluster 8 had increased expression of M2-like macrophage markers (for example, Marco, Clec10a [C-type lectin domain family 10 member A] and Mrc1 [Mannose receptor, C type 1]). Cluster 2 had high expression of H2-M2 [histocompatibility 2, M region locus] and Cd68, a general marker for macrophages. Cluster 3 expressed high levels of the cytokines and chemokines Il-33, C-X-C motif chemokine ligand 9 [Cxcl9] and Cxcl10. Cluster 9 represented dendritic cells expressing Itgax, Irf8 [Interferon regulatory factor-8], Xcr1 and Cd86. In addition, two granulocyte-like clusters were identified: cluster 7 which was delineated by high expression of neutrophil-associated genes (for example, Mpo [Myeloperoxidase], Tlr5 and Tnfsf13 [encoding BAFF]), and cluster 5 which contained transcripts associated with eosinophils (for example, Prg2 [eosinophil major basic protein], Cd24a [a glycosylphosphatidylinositol-anchored surface protein], Epcam [encoding EpCAM] and Irf4), while not expressing MHC-encoding genes (Fig. 1f–h). Comparing the profiles of uninfected and infected mice revealed a notable increase in cells belonging to several different clusters, with major effects in the different myeloid populations (Fig. 1g).
Spatial analysis highlighted distinct organization patterns for most clusters within the MLN. In the infected MLN, clusters 6 and 8 macrophages were localized differently; cluster 8 macrophages were primarily found in the capsule and subcapsular sinus (SCS), and cluster 6 macrophages formed dense, granuloma-like structures throughout the MLN at 4 w.p.i. (Extended Data Fig. 1f,g). The eosinophil cluster (cluster 5) was primarily observed in proximity to the granuloma structures and to the SCS, while the remaining myeloid clusters localized across the infected MLN (Extended Data Fig. 1f). To test whether classic signalling pathways shape these granulomas, we mapped interferon-γ (Ifng) transcripts in the infected MLN. IFNγ expression was enriched immediately adjacent to granuloma structures (Extended Data Fig. 1h), consistent with IFNγ’s established role in controlling chronic STm infection5,9.
Ccl11, but not Ccl24, transcripts were increased in STm-infected MLN (Fig. 1i and Extended Data Fig. 2a) and broadly distributed at low levels across multiple cell types, with the highest expression in monocytes, macrophages and eosinophils (Fig. 1i,j and Extended Data Fig. 2b). These data were confirmed using flow cytometry and fluorescence microscopy (Extended Data Fig. 2c–e). Spatial mapping showed distinct Ccl11 transcript enrichment across three regions in infected MLN tissue sections (Fig. 1j). Region 1, encompassing the T-cell zone (cluster 1 cells), and region 3, near the periphery of granuloma macrophages (cluster 6 cells), showed Ccl11 transcripts primarily in the monocyte cluster (cluster 3) cells. However, in region 2, located in the MLN capsule/subcapsular sinus area, Ccl11 was mainly expressed in SCS macrophages (cluster 8 cells) (Fig. 1j).
Depleting monocytes/macrophages using anti-CSF1 antibodies at 4 w.p.i. reduced CCL11-producing cells, total MLN CCL11 protein and eosinophil recruitment to the MLN (Extended Data Fig. 2f–l). These data demonstrate that these myeloid subsets are the essential drivers of the CCL11–eosinophil axis during persistent STm colonization.
CCL11 neutralization selectively reduces MLN eosinophils and increases local STm burdens
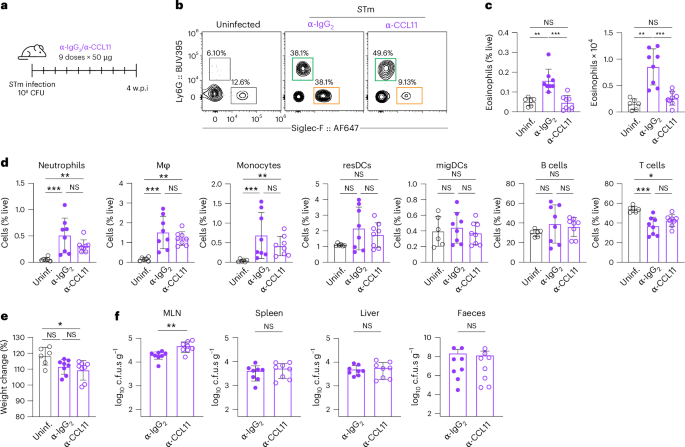
To test the contribution of CCL11 to persistent STm infection, mice received anti-CCL11 or an isotype control antibody starting at 3 days post infection and every 3 days thereafter until sacrifice at 4 w.p.i.26 (Fig. 2a). CCL11 blockade markedly reduced eosinophils in the MLN but not in the spleen, further indicating a tissue-restricted effect (Fig. 2b,c and Extended Data Fig. 3a–c). Numbers of neutrophils, monocytes, dendritic cells and macrophages were unchanged (Fig. 2d and Extended Data Fig. 3b). Although survival and weight were unaffected (Fig. 2e), anti-CCL11 treatment significantly increased STm colony-forming units (c.f.u.s) in the MLN (Fig. 2f) without altering bacterial levels in spleen, liver or faeces (Fig. 2f). Collectively, these data demonstrate that CCL11-mediated eosinophil recruitment helps to control pathogen burden in the MLN.
a, Schematic of the CCL11-neutralization approach. Mice were orally infected with STm (108 c.f.u.s) and treated with either anti-CCL11 antibodies or anti-IgG2 antibodies starting at day 3 post infection and continuing every 3 days until sacrifice at day 28 (n = 6–8 mice per group, 2 experiments). b, Representative FACS plots of neutrophils and eosinophils from MLN in mice treated with anti-IgG2 or anti-CCL11 at 4 w.p.i. compared to uninfected controls. c, Quantification of frequency (left) and absolute numbers (right) of MLN eosinophils in mice treated with anti-IgG2 (purple closed circles) or anti-CCL11 (purple open circles) at 4 w.p.i. compared to uninfected controls (black open circles). d, Quantification of frequency of live neutrophils, macrophages (Mϕ), monocytes, resident dendritic cells (DCs), migratory DCs, T cells and B cells from MLN in mice treated with anti-IgG2 or anti-CCL11 at 4 w.p.i. compared to uninfected controls. e, Percentage of body weight variation (day 28 vs day 0) in mice treated with anti-IgG2 or anti-CCL11 at 4 w.p.i. compared to uninfected controls. f, Quantification of bacterial levels in MLN, spleen, liver and faeces in mice treated with anti-IgG2 or anti-CCL11 at 4 w.p.i. Data are presented as mean ± s.e.m. All analyses are two-tailed and data were analysed using Kruskal–Wallis with Dunn’s post hoc test (c–e) or Mann–Whitney U-test (f). Exact P values in Supplementary Table 1. Uninf., uninfected. NS, non-significant; *P < 0.05, **P < 0.01, ***P < 0.001.
Distinct phenotype of MLN eosinophils during persistent STm infection
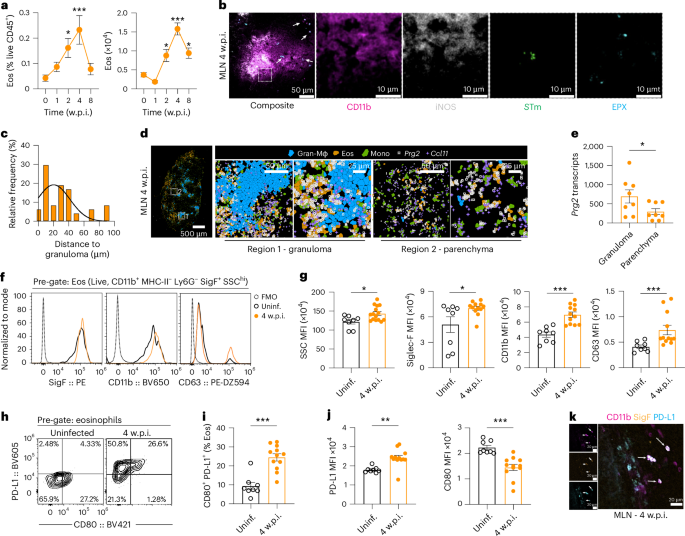
Prompted by the effect of CCL11 neutralization on bacterial control, we quantified the kinetics of eosinophil recruitment during chronic infection. Eosinophils expanded rapidly in the MLN, rising by 2 w.p.i., peaking at 4 w.p.i. and remaining elevated through 8 w.p.i. (Fig. 3a). Immunofluorescence and spatial transcriptomics showed EPX/Prg2+ eosinophils encircling CD11b+iNOS+ granulomas at a mean distance of ~30 μm; Prg2 transcripts were significantly enriched in cells within granulomatous regions versus the parenchyma (Fig. 3b–e). Concomitantly, eosinophil numbers declined in the blood and spleen of mice during the 4 weeks of infection (Extended Data Fig. 4a–c), indicating recruitment from systemic immune reservoirs.
129×1/SvJ mice were orally infected with STm (108 c.f.u.s) and monitored for up to 8 w.p.i. a, Quantification of frequency (left) and absolute numbers (right) of live eosinophils at 1, 2, 4 and 8 w.p.i. compared to uninfected controls (n = 7–12 mice per time point, 2–4 experiments). b, Representative immunofluorescence image of MLN sections from mice at 4 w.p.i showing eosinophils (EPX, cyan) localized at the periphery of CD11b+ (magenta) and iNOS+ (grey) granulomas containing STm (green). Arrows highlight eosinophils (n = 5 mice, 2 tissue sections per mouse). c, Quantification of distance between eosinophils and granuloma structures (n = 5 mice, 2 experiments). d, Representative image from Xenium analysis showing Prg2 transcripts (white symbols) in eosinophils at the periphery of granuloma macrophages (gran-Mϕ). Segmented cells are pseudocoloured according to UMAP cluster in Fig. 1f. e, Quantification of total Prg2 transcripts within 50 μm of granuloma structures and in the MLN parenchyma (n = 8 mice, 3 experiments). f, Representative FACS histogram for activation markers on MLN eosinophils from uninfected controls (black) and mice at 4 w.p.i (orange). Fluorescence minus one control (FMO) is shown as dashed black line. g, Quantification of MFI of activation markers: SSC-A, Siglec F, CD11b and CD63 on MLN eosinophils from uninfected controls and mice at 4 w.p.i. (n = 8–12 mice per group, 3 experiments). h, Representative FACS plots showing PD-L1 and CD80 on MLN eosinophils from uninfected controls and mice at 4 w.p.i. i, Quantification of the frequency of CD80+ PD-L1+ eosinophils from MLN in uninfected controls and mice at 4 w.p.i. (8–12 mice per group, 3 experiments). j, Quantification of PD-L1 and CD80 MFI on MLN eosinophils in uninfected controls and mice at 4 w.p.i. (8–12 mice per group, 3 experiments). k, Immunofluorescence image of infected MLN at 4 w.p.i showing CD11b+Siglec F+PD-L1+ positive eosinophils (n = 3 mice, 2 experiments). Data are presented as mean ± s.e.m. All analyses are two-tailed and data were analysed using Kruskal–Wallis with Dunn’s post hoc test (a), Wilcoxon signed-rank test (e) or Mann–Whitney U-test (g,i,j). Exact P values in Supplementary Table 1. *P < 0.05, **P < 0.01, ***P < 0.001.
Activated eosinophils are marked by increased granulation (side scatter (SSC)-area), maturation (Siglec-F), migration/adhesion (CD11b) and degranulation (CD63)27. Phenotyping across organs showed that MLN eosinophils from STm-infected mice have a distinct activation profile: relative to uninfected controls, eosinophils from the small intestine, colon, spleen or blood, they upregulate all four marker categories (Fig. 3f,g and Extended Data Fig. 4d,e). Strikingly, most MLN eosinophils were PD-L1hi, and a PD-L1+CD80+ subset emerged (Fig. 3h–k), a phenotype previously associated with antimicrobial and immunoregulatory activity in the gastrointestinal tract27. Together these data define a tissue-restricted, highly activated eosinophil population positioned around granulomas and poised to deliver antibacterial and immunomodulatory effects during persistent STm infection.
Eosinophils mediate local antimicrobial control of persistent STm
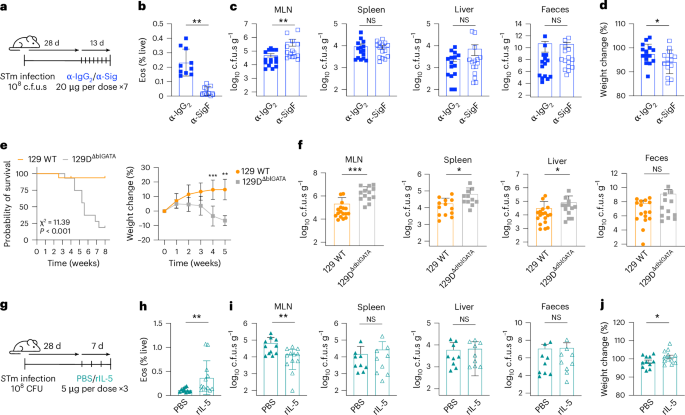
To test whether eosinophils actively restrain STm during persistence, we depleted them after infection was established. Mice infected for 28 days received anti-Siglec-F or isotype antibody (Fig. 4a). Anti-Siglec-F treatment efficiently depleted MLN eosinophils (Fig. 4b and Extended Data Fig. 5a) and caused a significant increase in MLN STm c.f.u.s, while bacterial loads in spleen, liver and faeces were unchanged (Fig. 4c), mirroring the phenotype of CCL11 neutralization (Fig. 2f). Depletion produced only a modest weight loss and no survival defect (Fig. 4d). Thus, eosinophils provide a compartmentalized antimicrobial function that limits STm expansion specifically within the MLN during chronic infection.
a, Schematic of transient eosinophil depletion using anti-Siglec F antibodies. Mice were orally infected with STm for 4 weeks (108 c.f.u.s) before receiving 7 doses of either anti-IgG2 or anti-Siglec F antibodies every 2 days before sacrifice on day 41 (n = 15 mice per group, 3 experiments). b, Quantification of the frequency of live MLN eosinophils in infected mice treated with anti-IgG2 (blue filled squares) or anti-Siglec F (blue open squares) at day 41 (n = 10 mice per group, 2 experiments; FACS plots in Extended Data Fig. 5a). c, Quantification of bacterial levels in MLN, spleen, liver and faeces in infected mice treated with anti-IgG2 or anti-Siglec-F at day 41. d, Percentage of body weight variation (day 41 vs day 28) in infected mice treated with anti-IgG2 or anti-Siglec F antibodies. e, Kaplan–Meier survival curve (left) comparing infected 129 WT mice (orange) and genetically eosinophil-deficient 129ΔdblGATA1 mice (grey) over 8 weeks along with body weight variation (right) in the same groups (n = 14–16 mice per group, 4 experiments). f, Quantification of bacterial levels in MLN, spleen, liver and faeces in 129 WT and eosinophil-deficient 129ΔdblGATA1 mice at 4 w.p.i. g, Schematic of transient eosinophilia by rIL-5 treatment. 129X1/SvJ mice were infected for 4 weeks before receiving 3 doses of rIL-5 or PBS every 2 days and sacrifice on day 35 (n = 11–12 mice per group, 3 experiments). h, Quantification of the frequency of live MLN eosinophils in infected mice treated with PBS (closed teal triangles) or rIL-5 (open teal triangles) at day 31. FACS plots in Extended Data Fig. 5g. i, Quantification of bacterial levels in MLN, spleen, liver and faeces in infected mice treated with PBS or rIL-5 at day 35. j, Percentage of body weight variation (day 35 vs day 28) in infected mice treated with PBS or rIL-5. Data are presented as mean ± s.e.m. All analyses are two-tailed and data were analysed using Mann–Whitney U-test (b–d,h–j), mixed effects analysis with Šidák’s multiple comparisons test (e) or Mantel–Cox analysis (e). Exact P values in Supplementary Table 1. *P < 0.05, **P < 0.01, ***P < 0.001.
Eosinophil-deficient mice are susceptible to persistent STm infection
To assess whether the absence of eosinophils influences acute STm infection dynamics, we examined the susceptibility of eosinophil-deficient ΔdblGATA1 mice using a C57BL/6 background, which is known for its heightened susceptibility to STm infection. Eosinophils did not influence acute typhoid-like disease as ΔdblGATA1 mice displayed WT-like weight loss, survival and systemic dissemination (Extended Data Fig. 5b,c). To assess persistence, ΔdblGATA1 mice were backcrossed to the 129×1/SvJ background. Baseline immune physiology was comparable between 129 WT and 129ΔdblGATA1 mice (Supplementary Table 2). After oral STm infection, however, eosinophil-deficient mice began losing weight by 4 w.p.i. and showed marked mortality by 6 w.p.i. (median survival 37 days; Fig. 4e). Histopathology at 4 w.p.i. revealed pronounced caecal and colonic inflammation and tissue damage in 129ΔdblGATA1 mice (Extended Data Fig. 5d), accompanied by elevated caecal STm burdens, implicating eosinophils in both bacterial control and maintenance of mucosal integrity. Although neither genotype exhibited STm bacteraemia, ~50% of eosinophil-deficient animals developed anaerobic bacteraemia, consistent with barrier leakage (Extended Data Fig. 5e).
MLN sections from both groups showed plasma cell expansion, but WT mice formed well-organized histocytic granulomas that were smaller and less defined in eosinophil-deficient 129ΔdblGATA1 mice (Extended Data Fig. 5f). Strikingly, 129ΔdblGATA1 mice harboured significantly higher STm loads in the MLN and increased levels in spleen and liver at 4 w.p.i. compared with WT mice (Fig. 4f). Together, these data demonstrate that while eosinophils are dispensable for acute infection, they are essential during chronic colonization to preserve mucosal and granuloma architecture and to limit systemic and MLN Salmonella persistence.
Transient eosinophilia enhances antimicrobial defence during persistent STm infection
To test whether increasing eosinophil numbers augments control of persistent STm, mice infected for 28 days received 3 intraperitoneal (IP) injections of recombinant IL-5, a cytokine that stimulates eosinophil maintenance, proliferation and maturation28, or PBS (Fig. 4g). rIL-5 treatment doubled MLN eosinophil numbers without altering body weight or disease severity (Fig. 4h and Extended Data Fig. 5g). Strikingly, this selective eosinophilia produced a significant reduction in MLN bacterial levels, while levels in spleen, liver and faeces were unchanged (Fig. 4i,j), Thus, boosting eosinophils improves local antimicrobial control specifically within the MLN during persistent Salmonella infection.
Eosinophils shape MLN granuloma immune cell composition
Since previous research has demonstrated that eosinophils can modulate TH1/TH2 balance in other settings23,29, we first compared MLN T cells in 129 WT and eosinophil-deficient 129ΔdblGATA1 mice. Flow cytometry of MLN CD4+ and CD8+ T cells showed no differences in TH1 (T-bet+), TH2 (GATA3+) or Treg (Foxp3+) frequencies (Extended Data Fig. 6a,b). Thus, altered T-cell polarization does not explain the heightened bacterial burden in 129ΔdblGATA1 mice.
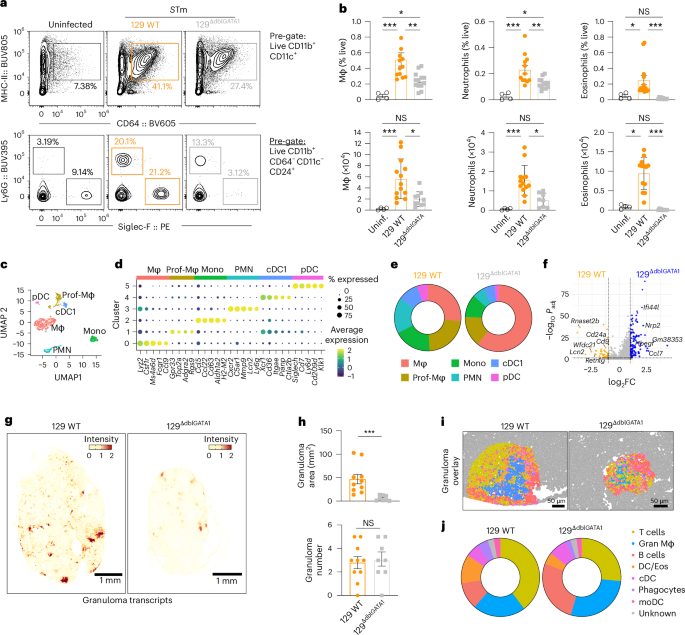
Eosinophil loss selectively reduced MLN neutrophils and macrophages, but monocytes and dendritic cells remained unchanged (Fig. 5a,b and Extended Data Fig. 6c,d); no effects were seen in the spleen (Extended Data Fig. 6e). Single-cell (sc)RNA-sequencing of CD11b+ and CD11c+ cells resolved six major populations: macrophages, proliferating macrophages, monocytes, neutrophils, cDC1s and pDCs (Fig. 5c,d), and confirmed a reshaped myeloid compartment in eosinophil-deficient mice, driven by shifts in proliferating macrophages, neutrophils and monocytes (Fig. 5e and Extended Data Fig. 7f). Transcriptionally, 129ΔdblGATA1 mice MLN showed reduced inflammatory/neutrophil-associated genes, such as Lcn2, Retnlg, Cd24a and Cd9, and increased levels of interferon-stimulated and chemotaxis genes including Ifi44, Ifi44l, Ccl2 and Ccl7 (Fig. 5f and Supplementary Table 3).
129 WT mice and eosinophil-deficient 129ΔdblGATA1 mice were orally infected with STm (108 c.f.u.s) for 4 weeks, after which immune populations in the MLN were analysed. See also Extended Data Figs. 6–8. a, Representative FACS plots of MLN macrophages (Mϕ), neutrophils and eosinophils in 129 WT and 129ΔdblGATA1 mice at 4 w.p.i. and in uninfected controls. b, Frequencies (top) and absolute numbers (bottom) of Mϕ, neutrophils and eosinophils in the MLN of 129 WT (orange) and 129ΔdblGATA1 mice (grey) at 4 w.p.i and in uninfected controls (black open circles) (n = 4–13 mice per group, 2–3 experiments). c, UMAP plot of myeloid subsets from scRNA-seq in 129 WT and 129ΔdblGATA1 mice at 4 w.p.i.: Mϕ (salmon), PMN (cyan), monocytes (green), proliferating Mϕ (prof-Mϕ, beige), cDC1 (light blue) and pDC (magenta); pooled samples from 3–5 mice, n = 2 mice per group, 1 experiment. d, Heat map of top 3–5 differentially expressed genes defining different myeloid clusters. e, Myeloid cluster proportions in 129 WT and 129ΔdblGATA1 mice. f, Volcano plot of differentially expressed genes in 129 WT and 129ΔdblGATA1 mice (cut-off: log2FC > 1, adjusted P = 0.05). Up in WT, orange; up in 129ΔdblGATA1, blue. g, Representative Xenium images of granuloma-associated transcript levels (Itgam, Nos2, Il-1b, Il-1rn and Tlr2) in 129 WT and 129ΔdblGATA1 mice at 4 w.p.i. (n = 6–8 mice per group, 3 experiments). h, Quantification of average granuloma size (top) and numbers (bottom) in 129 WT and 129ΔdblGATA1 mice at 4 w.p.i (defined by CD11b+iNOS+ foci, by immunofluorescence) (n = 8–10 mice per group, 3 experiments). i, Representative Xenium images of granuloma clusters in MLN tissue from 129 WT and 129ΔdblGATA1 mice at 4 w.p.i. Segmented cells are pseudocoloured according to UMAP clusters in Extended Data Fig. 8b (n = 6–8 mice per group, 3 experiments). j, Granuloma cluster proportions in 129 WT and 129ΔdblGATA1 mice at 4 w.p.i. Data are presented as mean ± s.e.m. or means (e,j). All analyses are two-tailed and data were analysed using Kruskal–Wallis with Dunn’s post hoc test (b), Wilcoxon rank-sum test with Bonferroni-adjusted P values (f) or Mann–Whitney U-test (h). Exact P values in Supplementary Table 1. *P < 0.05, ** P < 0.01, *** P < 0.001.
Macrophage states shift away from bactericidal programmes in the absence of eosinophils
Flow cytometry of macrophage subsets showed fewer iNOS+ (M1-like) macrophages, a subset in which STm persist10, while numbers of CD206+ (SCS-like) macrophages, reported to harbour replicating STm in vitro17, were unchanged (Extended Data Fig. 7a–c). iNOS+ macrophages displayed elevated CD14, MHC-II and CXCL9 (classical activation), while CD206+ cells upregulated the M2-marker CD301 (Extended Data Fig. 7d). scRNA-seq of the myeloid compartment further resolved four macrophage populations after subsetting (Extended Data Fig. 7e–h). Macrophages from infected 129ΔdblGATA1 mice were enriched for programmes linked to heparin/immunoglobulin binding and scavenger receptor activity, pathways that are typically associated with macrophage homeostasis and fibrosis. In contrast, macrophages from infected 129 WT mice expressed genes associated with growth factor, chemokine and NF-kB binding pathways linked to inflammatory pathogen-clearing states (Extended Data Fig. 7i,j).
Granulomas are smaller and less populated in the absence of eosinophils
Spatial Xenium mapping showed that mice contained markedly smaller CD11b+iNOS+ foci in 129ΔdblGATA1 versus 129 WT mice (Fig. 5g,h). Unbiased clustering of cells within and 50 μm around these foci identified eight granuloma-associated populations (Extended Data Fig. 8a,b). Eosinophil-deficient mice had a selective reduction of (1) T- and B-cell clusters, (2) DC/eos, cDCs clusters and (3) phagocyte clusters (Fig. 5i,j and Extended Data Fig. 8c). Pseudobulk analysis confirmed reduced expression of Prg2, Retnla, Rorc, Cdh1, Cd14 and Tlr2, markers of fully organized granulomas, in eosinophil-deficient mice (Extended Data Fig. 8d–f). Thus, eosinophils are required not only for granuloma size but also for recruitment or retention of multiple immune subsets within the lesion.
Transient depletion of eosinophils with anti-Siglec-F antibodies (days 28–42) recapitulated these defects: granuloma numbers were unchanged, but individual lesions were significantly smaller. In contrast, rIL-5-induced eosinophilia did not alter granuloma size (Extended Data Fig. 8g–j). Collectively, these data demonstrate that eosinophils maintain granuloma integrity, thereby supporting organized lesions that limit STm in the MLN.
Eosinophil deficiency increases intracellular STm burdens
To assess intracellular infection, we used a tdTomato-expressing STm that fluoresces inside host cells10. Macrophages were the main STm reservoir in both mouse genotypes, but eosinophil-deficient mice had fewer infected monocytes (Fig. 6a,b). Despite this, per-cell bacterial burden was higher in 129ΔdblGATA1 macrophages, as reflected by mean fluorescence intensity (MFI), versus 129 WT mice (Fig. 6c,d). Subtyping revealed fewer iNOS+ (M1-like) macrophages among infected cells in 129ΔdblGATA1 mice, whereas CD206+ SCS-macrophages were disproportionately infected and carried higher bacterial loads (Fig. 6e,f and Extended Data Fig. 8k,l). Thus, eosinophils sustain bactericidal myeloid subsets, especially M1-like macrophages, and thereby limit intracellular STm persistence in the MLN.
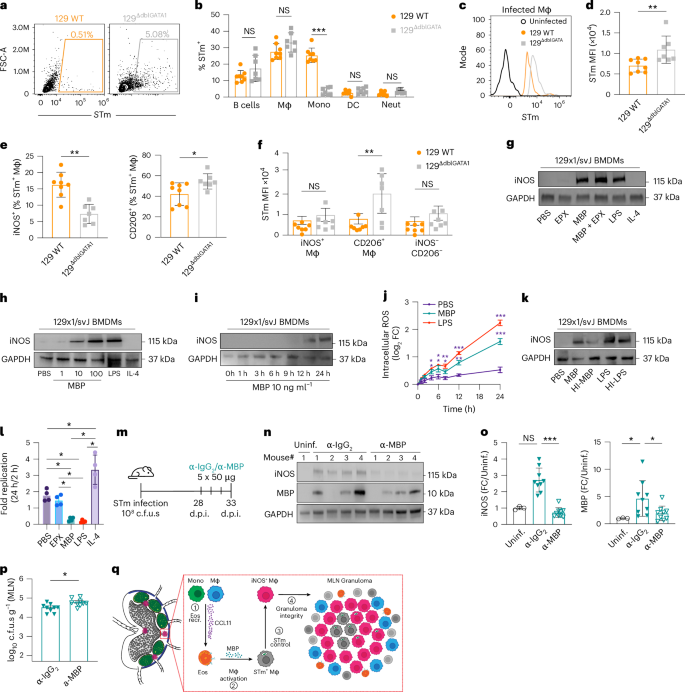
Eosinophil-deficient 129ΔdblGATA1 mice and 129 WT mice were orally infected with tdTomato-expressing STm (108 c.f.u.s) for 4 weeks to assess eosinophil-dependent bacterial control in the MLN. a, Representative FACS plots of tdTomato-STm in 129 WT and 129ΔdblGATA1 mice at 4 w.p.i. b, Frequency of tdTomato-STm+ immune cells in 129 WT and 129ΔdblGATA1 mice (n = 7–8 mice per group, 2 experiments). c, Representative FACS histogram of tdTomato-STm+ macrophages (Mϕ) in 129 WT (orange) and 129ΔdblGATA1 (grey). Uninfected controls (black) served as a negative control. d, MFI of tdTomato-STm within Mϕ in 129 WT and 129ΔdblGATA1 mice. e, Frequency of tdTomato-STm+ iNOS+ or tdTomato-STm+ CD206+ Mϕ in 129 WT and 129ΔdblGATA1 mice (n = 7–8 mice per group, 2 experiments). f, MFI of tdTomato-STm within iNOS+, CD206+ and iNOS−CD206− Mϕ in 129 WT and 129ΔdblGATA1 mice (n = 7–8 mice per group, 2 experiments) (see also Extended Data Fig. 7k). g–i, Western blots of iNOS and GAPDH in whole-cell lysates from 129 WT BMDMs stimulated with recombinant EPX (100 ng ml−1), MBP (100 ng ml−1), MBP and EPX (100 ng ml−1 each), LPS (100 ng ml−1) or IL-4 (20 ng ml−1) (g); dose-dependent MBP effects (1–100 ng ml−1) (h); kinetics of MBP effects (10 ng ml−1, 1–24 h) (i). n = 3 biological replicates (see also Extended Data Fig. 9). j, Intracellular ROS levels measured over time in 129 WT BMDMs stimulated with MBP (10 ng ml−1) and LPS (100 ng ml−1) (n = 3 biological replicates). k, Western blots of iNOS and GAPDH in 129 WT BMDMs stimulated with MBP, heat-inactivated (HI)-MBP, LPS or HI-LPS for 24 h (3 biological replicates). l, Fold-replication of STm in 129 WT BMDMs pre-stimulated with MBP (10 ng ml−1), EPX (10 ng ml−1), LPS (100 ng ml−1) or IL-4 (20 ng ml−1) for 24 h before gentamycin protection assay (bacteria enumerated at 2 h and 24 h, 4 biological replicates). m–p, Schematic of MBP neutralization using anti-MBP antibodies. Mice were infected with STm for 4 weeks and then given 5 daily doses of anti-IgG2 or anti-MBP antibodies before sacrifice on day 33 (m); western blots of iNOS, MBP and GAPDH in whole MLN lysates from uninfected and infected mice treated with anti-IgG2 or anti-MBP at 33 days post infection (n); quantification of iNOS (left) and MBP (right) protein levels in MBP-neutralized mice compared to IgG2-treated mice and uninfected controls (n = 3–9 mice per group, 2 experiments) (o); MLN bacterial burden in MBP-neutralized mice compared to IgG2-treated mice (n = 9 mice per group, 2 experiments) (p). q, Proposed model showing how macrophages and monocytes recruit eosinophils to the MLN through CCL11-dependent signalling. Recruited eosinophils become activated in STm-infected MLN and release MBP which leads to STm control within macrophages, increased macrophage iNOS levels and sustained granuloma integrity. All analyses are two-tailed and data were analysed using Mann–Whitney U-test (b,d,e,p); two-way ANOVA with uncorrected Fisher’s LSD test (f), two-way ANOVA with Tukey correction (j), one-way ANOVA with Tukey correction (l) or Kruskal–Wallis with Dunn’s post hoc test (o). Exact P values in Supplementary Table 1. *P < 0.05, **P < 0.01, ***P < 0.001.
Eosinophil-specific MBP primes macrophages for antimicrobial activity
Mature murine eosinophils release the secondary granule proteins MBP and EPX upon activation30. MBP is a cationic protein that is toxic to parasites and some mammalian cells31,32. EPX is a haem peroxidase that, with H2O2, oxidizes halides and nitric oxide and can damage microbes33. Both EPX+ and MBP+ cells localize to the periphery of MLN granulomas (Extended Data Fig. 9a). To test direct antibacterial activity, we incubated recombinant MBP or EPX with STm at concentrations previously reported to be antimicrobial31,34. Neither protein reduced bacterial viability (Extended Data Fig. 9b). Instead MBP, but not EPX, induced dose- and time-dependent increases in macrophage iNOS and intracellular reactive oxygen species (ROS), mirroring LPS responses, while EPX did not alter Arginase-1 levels (Fig. 6g–j and Extended Data Fig. 9c–e). To confirm specificity, heat-denatured MBP lost activity after treatment. In addition, human MBP reproduced the mouse MBP effects on macrophages. (Fig. 6k and Extended Data Fig. 9e,f), Strikingly, MBP-primed macrophages restricted intracellular STm replication to the same extent as LPS-polarized (M1) controls without increasing cell death (Fig. 6l and Extended Data Fig. 9g)35, reinforcing a direct network between eosinophils and macrophages in maintaining STm control.
To further assess the contribution of MBP to antimicrobial immunity during persistent STm infection in vivo, mice were infected for 28 days before receiving 5 consecutive daily doses of either polyclonal anti-MBP or isotype control antibodies (Fig. 6m). MBP blockade significantly reduced iNOS expression in MLN tissue (Fig. 6n,o) and concomitantly increased STm c.f.u.s in the MLN (Fig. 6p). Together, these findings establish MBP as a previously unrecognized eosinophil-derived licensing signal that converts macrophages into iNOS-high antibacterial effectors, linking eosinophil activity to granuloma integrity and durable control of persistent Salmonella.