Chemicals and reagents
Vancomycin hydrochloride, bovine serum albumin (BSA), 1-Ethyl-3-[3-dimethylaminopropyl] carbodiimide hydrochloride (EDC), gold nanoparticles (40 nm diameter) in 0.1 mM phosphate-buffered saline (PBS) buffer, PBS, polyethylene glycol (PEG) 6,000 Da, avidin, Anhydrous Na2CO3, anhydrous NaHCO3, H2SO4, phosphate buffered saline (PBS) tablets, Tween®-20, membrane cut-off value 12 KDa, Pur-A-Lyzer™ Mega Dialysis Kit– were purchased from Sigma-Aldrich. EZ-Link™ Sulfo-NHS-LC-Biotinylation Kit was purchased from ThermoFisher Scientific (Waltham, MA). Sheep anti vancomycin IgG, donkey anti sheep/goat IgG: HRP, HRP-linked anti-his antibody, 3,3′,5,5′-tetramethylbenzidine (TMB) Core + were purchased from Bio-Rad (Hercules, CA). Water was prepared with Milli-Q water system (Millipore, Burlington, MA). HiFlow Plus TM membrane, glass fibre conjugate sheet, and cellulose fibre roll were from Merck Millipore. Other common reagents were bought from Sigma-Aldrich unless specified. Printing was conducted user Automated Lateral Flow Reagent Dispenser (ALFRD) (Claremont Bio, CA, USA) incorporated with Fusion 200 Pump (Chemyx, TX).
BSA-vancomycin conjugate preparation
Vancomycin was coupled to BSA using EDC. BSA (70 mg) was dissolved in 15 mL PBS (pH 7.4) and 1 mL of vancomycin solution (78 mg/mL in DH2O) was added dropwise into the beaker containing the BSA solution. Next, 1 mL of EDC (200 mg/mL in purified H2O) was added dropwise into the same beaker. The reaction mixture was left stirring at 500 rpm at room temperature (18–20 oC) for 1 h and then moved to a cold room, where it was kept overnight at 4 °C. The next day, the conjugate was purified by dialysis. The dialysis tube was kept spinning in a beaker filled with PBS stirring at 500 rpm, at 4 °C overnight. The PBS buffer was changed three times15,16. Quantification of the BSA concentration post the conjugation reaction was completed using a BCA protein assay kit (ThermoFisher). After dialysis, the vancomycin BSA conjugate (BSA-Van) was stored in aliquots at − 80 °C.
Development of indirect competitive enzyme-linked immunosorbent assay (ELISA)
An indirect competitive ELISA was developed to characterise the BSA–Van conjugate and to evaluate the quantitative binding ability of anti-vancomycin IgG. The secondary antibody used was donkey anti-sheep/goat IgG: HRP.
Briefly, 1.5 µg of BSA–Van was coated onto 96-well microtiter plates using 50 mM carbonate/bicarbonate buffer (pH 9.6) and incubated at 4 °C for 24 h. After three washes with washing buffer (0.05% Tween-20 in PBS), wells were blocked with PBS containing 1% BSA (w/v) for 1 h at 37 °C. Following another wash, 100 µL of anti-vancomycin IgG and 100 µL of vancomycin standard solutions were added to each well and incubated for 1 h at 37 °C. Plates were washed three times before adding 100 µL of the HRP-linked secondary antibody, followed by incubation for another hour at 37 °C.
After a final wash, 100 µL of TMB substrate was added and incubated at 37 °C for 15 min in the dark. The reaction was stopped with 50 µL of 2 M sulfuric acid per well, and absorbance was measured at 450 nm using a microplate reader (FLUOstar Omega, BMG Labtech, Germany). Calibration standards were prepared by serial five-fold dilution of vancomycin in serum, yielding concentrations of 1, 5, 25, 125, 625, 3,125, 15,625, and 78,125 ng/mL.
Quality control (QC) samples at three concentration levels (low, medium, high: Qc1, Qc2, Qc3) were used to evaluate assay accuracy. Intra-day precision was assessed by testing each QC level in triplicate on the same day. Inter-day precision was determined by testing the QC levels in triplicate on four consecutive days. Accuracy was evaluated as the percentage agreement between the measured and spiked QC concentrations15,16,17.
$${\text{Accuracy }}\left( {{\text{percentage}}} \right) = \left( {{\text{measured}}/{\text{known spiked}}} \right) \times {\text{100}}.$$
The calibration curve showed a linear relationship between absorbance and vancomycin concentration. Therefore, the limit of detection (LOD) and limit of quantification (LOQ) were calculated using the standard deviation of the response (σ) and the slope of the calibration curve (s), as follows:
$${\text{LOD}} = {\text{3}}.{\text{3}} \times \sigma /{\text{s}}$$
$${\text{LOQ}} = {\text{1}}0 \times \sigma /{\text{s}}$$
Biotinylation of the BSA-Van
A standard biotinylation procedure was followed to modify the BSA-Van conjugate with NHS-LC-biotin following the ThermoFisher protocol18. The product was purified by dialysis and the modification by biotin was confirmed through a HABA-avidin assay18. To further validate the conjugation, chip-based analysis was performed with the Agilent 2100 Bioanalyzer system following Protein 80 kit protocol19,20. The conjugates were compared with non-biotinylated conjugates and a BSA standard under non-denaturing conditions. Finally, the biotinylated BSA-Van was stored at -80 °C for further experiments.
Labelling anti-vancomycin IgG and conjugates with gold nanoparticles (AuNP)
AuNP labelling was performed by adding 100 µL of vancomycin IgG and 100 µL of biotinylated BSA-Van to separate 1 mL aliquots of gold nanoparticles in individual Eppendorf tubes. The pH of each AuNP solution was adjusted to 8.4 using 0.5 M NaOH. Tubes were vortexed for 5 s and placed on a shaker for 1 h at room temperature (RT). Subsequently, 25 µL of 10% BSA solution was added to each tube and incubated at RT for 3 min. The conjugates were then centrifuged at 13,500 rpm for 10 min.
After discarding the supernatant, the pellet was resuspended in 125 µL of running buffer. The running buffer used for vancomycin assays consisted of PBS containing 0.5% (v/v) Tween 20, 0.5% (v/v) Triton X-100, 0.5% (v/v) glycerol, 1% (w/v) sucrose, and 0.25% (w/v) BSA. The labelled conjugates were stored at 4 °C until use21.
LFA strip assembly
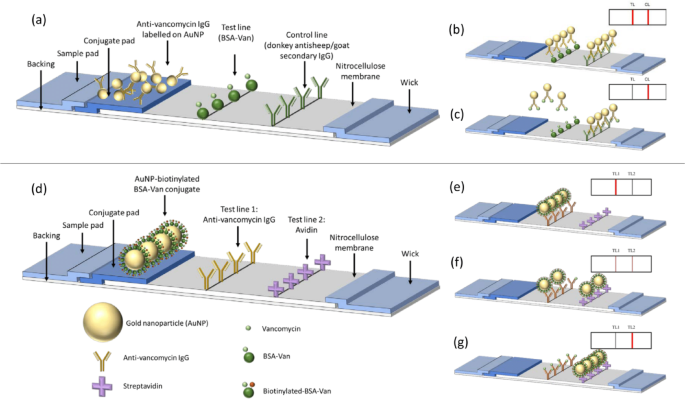
A traditional competitive LFA of vancomycin was developed and compared with the novel competitive LFA for vancomycin. Figure 1 represents the structure of both traditional competitive and novel competitive LFA.
For the traditional competitive LFA, vancomycin detection was based on competition between free vancomycin in the serum sample and the immobilised BSA–Van conjugate printed on the test line, both competing to bind to the anti-vancomycin IgG conjugated to gold nanoparticles. Upon application, vancomycin in the sample binds to the AuNP–IgG, forming immune complexes. Unbound AuNP–IgG is then captured by the BSA–Van on the test line. The resulting test line intensity is inversely proportional to the vancomycin concentration in the sample. Calibration standards for the traditional competitive LFA were prepared in vancomycin-free serum at concentrations of 0.12, 0.37, 1.11, 3.33, and 10 ng/mL.
For the novel competitive LFA, instead of using a traditional single test line and control line, this design incorporates two distinct test lines: one with anti-vancomycin IgG and another with avidin. Gold nanoparticles were conjugated to biotinylated BSA–Van. In the presence of vancomycin in the sample, the analyte competes with the conjugated vancomycin to bind the anti-vancomycin IgG on the first test line. Excess conjugate is then captured on the second test line via biotin–avidin binding. As the vancomycin concentration increases, the intensity of the first test line decreases, while the intensity of the second test line increases.
Conjugate pads were pre-treated with the optimised buffer alone for 24 h, followed by another 4-hour pre-treatment with the AuNP conjugates. The optimised buffer for vancomycin was the same as the running buffer. The final volume of AuNP conjugate dispensed was 30 µl per cm2 of the conjugate pad. The membrane backing card (7 × 30 cm) and the NC membrane (3.2 × 25 cm) were cut with a guillotine cutter and assembled with double-sided tape. The reagents [anti-Van antibody (1 mg/mL) or BSA-Van (1 mg/mL) or avidin (1 mg/mL) depending on the LFA method; Fig. 1] were taken into 1 mL syringes and dispensed on the nitrocellulose membrane using the ALFRD dispenser. The infusion rate of the syringes was 0.4 ml/min and the printing speed was adjusted to 32.5 mm/sec, allowing the volume of 0.205 µl/mm printed. The membrane with the printed test lines was left to dry at room temperature for 20 min before proceeding with strip assembly. The pre-treated conjugate pad and absorbent pad were assembled with double-sided tape onto the top and bottom of the strip, respectively. The 30 cm long NC membrane was cut into 5 mm strips, the standard width of commercial LFAs21.
Schematic of the structure of competitive LFA (a–c) and the novel competitive LFA (d–f). (a) The detection antibody can bind to both antibodies on test line (TL) and control line (CL). (b) Negative result. (c) Positive result. (d) The detection antigen conjugated on AuNP can bind to antibodies on test line 1 (TL1) whereas the biotin can bind to test line 2 (TL2). (e–g) As analyte concentration increases, signal at TL1 decreases while signal at TL2 increases.
Rationale for dual test-line configuration
Importantly, the novel LFA format intentionally replaces the conventional test/control configuration with two functional test lines: T1 and T2. The first line (T1) is coated with anti-vancomycin antibodies to capture analyte-bound conjugate, while the second line (T2) is coated with avidin to bind excess unbound biotinylated conjugate. Although T2 primarily serves a quantitative role, its consistent presence in blank and low-concentration samples offers an indirect verification of sample migration. The role of the traditional control line is to confirm test validity when no analyte is present; however, in our design, either or both test lines must be visible for the assay to be valid, rendering an independent control line unnecessary. Future versions may incorporate a dedicated control line with a distinct nanoparticle label to further enhance robustness.
LFA testing and readout
The assay was run with samples in optimised running buffer. A sample volume of 100 µL was used for vancomycin. Typically, the time between sample application and image capture was 5 min. The images were taken with a smartphone (Samsung A21s). After which, the images were analysed using R software to measure the intensity of the lines. The R code written for LFA reading is available at: https://github.com/vvasikasin/LFA_CAMO. Specifically, the analyses and relevant packages were implemented through a script within RStudio (PBC, Boston, MA). The core packages employed in these analyses comprised dplyr, hyperSpec, stats, baseline, and the Bioconductor package EBImage22. Briefly, the intensities of the test line 1 and test line 2 were normalised to the intensity of the background. The linear model was based on the concentration and the ratio of the relative intensity of both test lines to the background. The detail of this algorithm is described in Supplementary Appendix 1. The volume of sample applied to the LFA was optimised to achieve adequate membrane flow and signal development. Vancomycin required a larger volume (100 µL), likely due to its molecular size and lower permeability. This volume was selected empirically to ensure optimal signal intensity and reproducibility.
Calibration sample preparation for the novel LFA
To construct the calibration curve for the novel LFA, vancomycin-free serum samples were spiked with vancomycin at the following concentrations: 2.88, 14.40, 72, 360, 1,800, 9,000, and 45,000 ng/mL. Blood was collected from healthy adult volunteers and processed to obtain serum via centrifugation. Ethical approval for the study was obtained from the London Harrow Research Ethics Committee (19/LO/0219), and the study was registered on ClinicalTrials.gov (NCT04053140). Written informed consent was obtained from all participants in accordance with the Declaration of Helsinki. The methodology for serum preparation and spiking was adapted from validated clinical protocols for antibiotic quantification23.
Calculation of LOD and LOQ for the novel vancomycin LFA
LOD and LOQ were calculated using the standard deviation (SD) of blank samples and the calibration curve obtained from the assay. Three blank (0 ng/mL) serum samples were measured to determine the mean blank signal and its standard deviation. The LOD was defined as the signal equal to the mean blank plus three times the SD, and the LOQ as the mean blank plus ten times the SD.
The calibration curve followed a power-law equation of the form: Y = 0.022 × x0.6926. Where y is the measured signal and x is the vancomycin concentration. To determine LOD and LOQ in concentration units (ng/mL), the corresponding signal values were substituted into the inverse of the calibration equation.
Reproducibility and strip stability
The reproducibility of the novel vancomycin LFA was evaluated by assessing both intra-assay (within the same day) and inter-assay (between different days) precision. Three quality control concentrations—high (QC1: 45,000 ng/mL), medium (QC2: 360 ng/mL), and low (QC3: 2.88 ng/mL)—were tested. Each concentration was measured in five replicates (n = 5). Intra-day precision was assessed by running all replicates on the same day, while inter-day precision was evaluated by repeating the assay on five different days24.
To assess the stability of the novel LFA for vancomycin across different storage temperatures, a single batch of assembled LFA strips was divided into two sub-batches. One sub-batch was stored at room temperature, and the other at 4 °C for one week. After storage, all strips were tested using serum samples spiked with 2 µg/mL of vancomycin, and signal intensities were compared to freshly prepared strips.
Specificity testing
To evaluate the specificity of the vancomycin LFA, we tested structurally related and unrelated antibiotics: teicoplanin (a glycopeptide) and ceftriaxone (a β-lactam antibiotic). Stock solutions of teicoplanin and ceftriaxone were prepared in serum and diluted to final concentrations of 10,000 ng/mL, 45,000 ng/mL, and 100,000 ng/mL. These concentrations were selected to cover typical therapeutic levels (10,000 ng/mL), the upper limit of the vancomycin LFA detection range (45,000 ng/mL), and a high excess level (100,000 ng/mL) to test worst-case cross-reactivity. Each condition was tested in triplicate. After the assay developed, test strips were measured using the same image analysis software used for vancomycin calibration.
The signal intensity produced by the interferents (teicoplanin and ceftriaxone) was converted into vancomycin-equivalent concentrations using the vancomycin calibration curve. The cross-reactivity was calculated using the formula: cross-reactivity (%) = (measured concentration of VAN) / (expected concentration of interferent) X 1006.
Serum sample analysis and recovery
To evaluate the performance of the novel competitive LFA in a biologically relevant matrix, human serum samples were spiked with known concentrations of vancomycin and analysed using the developed assay. Three target concentrations were tested: 1,000 ng/mL, 500 ng/mL, and 70 ng/mL. Each concentration was tested in triplicate (n = 3). Recovery (%) was determined using the following formula:
Recovery (%) = (measured concentration) / (spiked concentration) X 10023.