Redundant genes are prevalent in prokaryotic genomes, with pathogens exhibiting higher redundancy ratios compared to non-pathogens
To effectively identify redundant genes in prokaryotic genomes on a large scale, we developed SoDpipe, a pipeline tailored for the automated analysis of redundant genes and their translation initiation signals in prokaryotes (see Methods, Supplementary Information, and Supplementary Fig. 1). We employed SoDpipe on 22,310 complete prokaryotic genomes from RefSeq, belonging to 6,450 species (Supplementary Data 1). We applied two criteria, widely used and consistent with previous studies23,24, to identify redundant genes, resulting in two datasets: Set-80 (80% identity and 80% coverage) and Set-100 (100% identity and 95% coverage). Set-100 comprised identical redundant genes most likely derived from recent duplications, while Set-80 may also include xenogeneic genes derived from HGT23. In total, we identified 2,432,287 redundant genes in 21,842 genomes, forming 800,486 redundant clusters in Set-80, meanwhile 1,042,374 redundant genes in 20,138 genomes, forming 307,360 clusters in Set-100. Additionally, we found a strong positive correlation between the number of redundant genes and prokaryotic genome size (Supplementary Fig. 2; Set-80: p-value < 0.001, r = 0.534; Set-100: p-value < 0.001, r = 0.246), supported by previous research7.
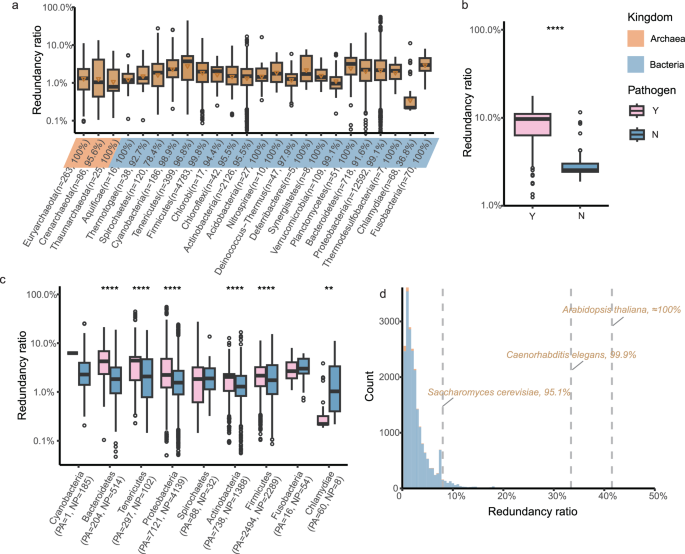
Given the compact and efficient nature of prokaryotic genomes, the prevalence of redundant genes within a bacterial genome may initially appear counterintuitive8. Yet, our analysis indicated that redundant genes were prevalent across most phyla of prokaryotic genomes, with their presence observed in 97.9% of genomes in Set-80 and 90.3% in Set-100 (Fig. 1a and Supplementary Fig. 3a). Considering the inherent variability in prokaryotic genome size and our analysis showing that prokaryotic genome size increases with the number of redundant genes, we have defined the redundancy ratio as the number of redundant genes divided by the number of protein-coding genes. As a result, the redundancy ratio in each prokaryotic genome showed large variability, ranging from 0% to 55.5%, with an average of 2.9% for Set-80 (the average redundancy ratio in Set-100 was 1.4%; Supplementary Data 1). Notably, after excluding phyla represented by only one genome, Tenericutes exhibited the highest average redundancy ratio, reaching 5.0% in Set-80 and 3.1% in Set-100, followed by Planctomycetes, Cyanobacteria, and Fusobacteria.
a The redundancy ratios for each phylum (n ≥ 5). The lower triangle indicates the average redundancy ratio. b The redundancy ratios for pathogens (n = 233) and non-pathogens (n = 64) in E. coli strains (p-value < 2.2e-16, Wilcoxon rank-sum test, two-sided). c The redundancy ratios for pathogens and non-pathogens in each phylum. The exact number and percentage of genomes with redundant genes are detailed in parentheses. Two-sided Wilcoxon rank-sum tests were performed, and p-values were adjusted for multiple testing using the false discovery rate method. d The distribution of redundancy ratios in prokaryotes and the ratio of three model eukaryotic organisms are represented as dashed lines. The percentage of prokaryotic genomes with a smaller redundancy ratio than each eukaryotic organism is indicated, separated by commas. Box plots indicate median (middle line), 25th, 75th percentile (box) and 5th and 95th percentile (whiskers) as well as outliers (single points). Source data are provided as a Source Data file. * p-value < 0.05; ** p-value < 0.01; *** p-value < 0.001; **** p-value < 0.0001.
To assess the difference in gene redundancy between pathogens and non-pathogens, we annotated the pathogenicity of each microbe (see Methods) and discovered that pathogens had a significantly higher redundancy ratio than non-pathogens (p-value < 0.001, Wilcoxon rank-sum test). Specifically, in Set-80, the redundancy ratio for pathogens was 3.3 ± 3.4%, compared to 2.3 ± 2.3% in non-pathogens. Similarly, in Set-100, the ratio was 1.7 ± 2.3% for pathogens and 1.0 ± 1.5% for non-pathogens, respectively. This conclusion is supported by observations at the strain level showing a significantly higher redundancy ratio in pathogenic E. coli strains (n = 233; 8.7 ± 3.1% in Set-80, 3.4 ± 1.8% in Set-100) compared to their non-pathogenic strains (n = 63; 3.1 ± 1.6% in Set-80, 1.2 ± 1.3% in Set-100; p-value < 0.001, Wilcoxon rank-sum test; Fig. 1b and Supplementary Fig. 3b). This suggested that some redundant genes were pathotype-specific and strain-specific, consistent with previous studies on 38 E. coli strains18. We found that, with the exception of the Chlamydiae phylum, pathogens consistently exhibited a higher redundancy ratio compared to non-pathogens across all phyla (Fig. 1c and Supplementary Fig. 3c). We further annotated functions of the redundant genes with the databases of COG (Clusters of Orthologous Groups)25 and found that pathogenic bacteria contained more redundant genes involved in ‘nucleotide transport and metabolism’ (class F, pathogens vs. non-pathogens: 0.8 ± 1.8% vs. 0.7 ± 2.7%, p-valueadj < 0.05, Wilcoxon rank-sum test, Supplementary Data 2), ‘secondary metabolite transport and metabolism’ (class Q, 1.4 ± 3.4% vs. 1.3 ± 2.8%, p-valueadj < 0.05), ‘energy production and conversion’ (class C, 4.5 ± 7.3% vs. 3.9 ± 6.5%, p-valueadj < 0.05), and ‘extracellular structures’ (class W, 0.45 ± 2.5% vs. 0.38 ± 1.8%, p-valueadj < 0.05) in Set-80. This observation is similar to findings from the previous study26, which demonstrate that fungal plant pathogens possess additional genes involved in secondary metabolism and in managing interactions with hosts compared to non-pathogens.
To better understand the degree of gene redundancy in prokaryotes, we calculated the redundancy ratio for several eukaryotic model organisms, including S. cerevisiae, Caenorhabditis elegans, and Arabidopsis thaliana. The results reveal that the redundancy ratio accounts for 8.0%, 33.4%, and 41.5% for these three eukaryotes in Set-80, respectively, and for 2.3%, 6.2%, and 14.5% in Set-100. Thus, most prokaryotes exhibit a lower redundancy ratio compared to eukaryotes. Specifically, the redundancy ratio of S. cerevisiae exceeds that of 95.1% of prokaryotic genomes with Set-80 parameters and 82.7% under Set-100 parameters, with the redundancy ratio of C. elegans and A. thaliana exceed 99.9% (95.4% in Set-100) and nearly 100.0% (nearly 100.0% in Set-100) of prokaryotes under Set-80 parameters, respectively (Fig. 1d and Supplementary Fig. 3d).
Redundant genes contribute to niche specialization
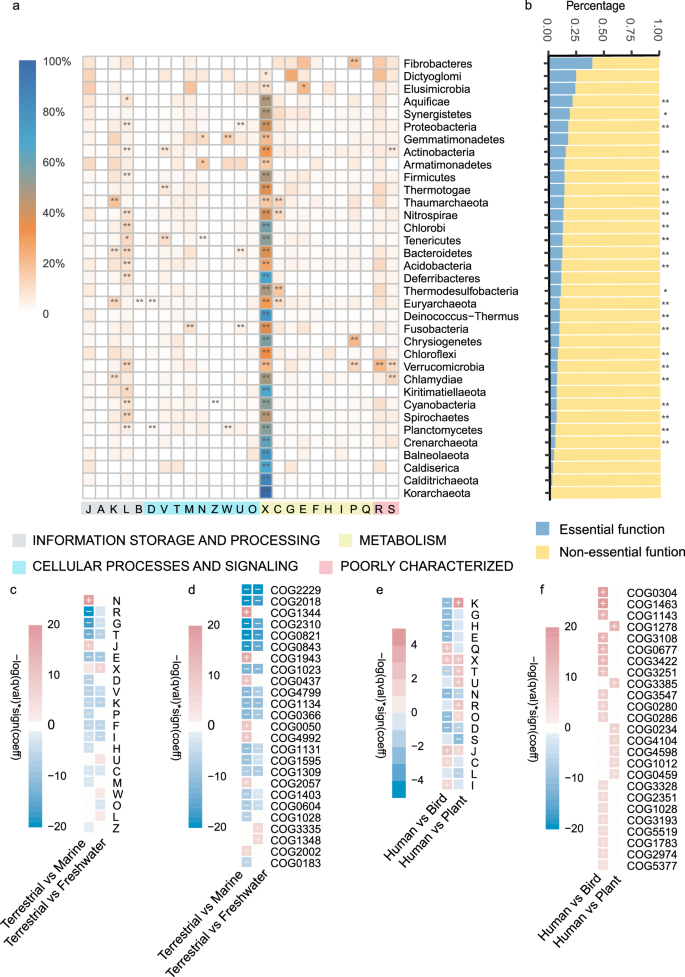
To survey the functional distribution of redundant genes in prokaryotes, we annotated all redundant genes and discovered their involvement across all COG functional categories, with a notable predominance in ‘mobilome: prophages, transposons’ (class X) (Fig. 2a and Supplementary Fig. 4a). Class X genes serve as vehicles for transmitting many vital genes associated with adaptive traits, thereby playing a crucial role in bacterial adaptation27. By leveraging comprehensive plasmid annotations in RefSeq, we further observed via mixed-effects linear regression that the redundant gene ratio increased with the number of plasmids (Set-80, p-value = 3.11 × 10−9, the slope coefficient = 1.633 × 10−3; Set-100, p-value = 1.43 × 10−8, the slope coefficient = 9.057 × 10−4). This suggests that mobile genetic elements may actively contribute to the formation of redundant genes. To further assess whether redundant genes tend to be indispensable for survival, we annotated them using the DEG (Database of Essential Genes) database28. The results revealed that, across most phyla, essential genes represented only a minor proportion of redundant genes (Fig. 2b, Supplementary Fig. 4b, Supplementary Data 3). A previous study on E. coli supported the above observations and highlighted that genes involved in fundamental cellular processes are highly essential and less frequently duplicated than those facilitating interactions and adaptation to diverse environments29.
a The heatmap presents the proportion of functions of redundant genes in each COG category. The black asterisk indicates a significantly higher proportion of the corresponding category (p-value < 0.05, Fisher’s exact test, two-sided). b The bar plot to the right presents the proportion of essential functions and non-essential functions, annotated by the DEG database. The black asterisk indicates a significantly higher proportion of non-essential than essential functions (p-valueadj < 0.05, Wilcoxon rank-sum test, two-sided, Supplementary Data 3). The hyphen represents all the other redundant genes that do not assign a COG function. c, d Changes in the functional abundance of redundant genes between the Terrestrial (n = 265), Marine (n = 383), and Freshwater (other aquatic environment excludes marine, including freshwater, lake, river, sediment, and sludge; n = 295) groups, presented as COG categories and COG IDs, respectively. The heat map shows the differentially abundant redundant functions, with the color scale calculated using q-values and beta coefficients generated from the general linear model in MaAslin2. The Terrestrial group was set as a reference. The scale indicates the degree of enrichment (red) or depletion (blue) compared to the Terrestrial group, with deeper colors signifying greater significance. e, f Changes in the functional abundance of redundant genes between Human (n = 297), Plant (n = 123), and Bird (n = 113) groups, presented as COG categories and COG IDs, respectively. The Human group was set as a reference. * p-value < 0.05; ** p-value < 0.01.
To advance the understanding of the functional distribution of redundant genes within prokaryotic genomes across different environments, we classified prokaryotes into six groups according to their isolation source (see Methods). By comparing microbes isolated from the Marine, Freshwater, and Terrestrial groups, representing three natural environments, we found that redundant genes in class N ‘Cell Motility’ were significantly more abundant in the Marine group, with genes such as COG1344 being associated with flagellum formation and swimming30 (p-valueadj < 0.05, Fig. 2c, d, Supplementary Data 4). In the Terrestrial group, we observed a significant enrichment of class G ‘Carbohydrate Transport and Metabolism’ (including COG1023, COG1134, and COG0366), which may be related to the nutrient scarcity typically found in aquatic environments31. Additionally, the abundance of redundant genes associated with signal transduction and stress response was significantly higher in the Terrestrial group (including COG2229 and COG2018). The greater environmental variability in terrestrial ecosystems likely necessitates more flexible signal transduction mechanisms that enable rapid adaptation to environmental changes32. The comparison of bacteria among Human, Plant, and Bird groups showed that Class K ‘Transcription’ (including COG1278, a cold shock protein, p-valueadj < 0.05, Fig. 2e, f, Supplementary Data 4) was significantly enriched in isolates from plants. Due to the large diurnal temperature fluctuations and pronounced seasonal climate changes that plants experience, in contrast to the relatively stable body temperatures of humans and birds, these redundant genes may enhance the ability of plant-associated microbes to tolerate cold stress33. Moreover, the typically higher body temperature of birds may impose demands on fatty acid synthesis and lipid metabolism, leading to the redundancy of COG0304 (3-oxoacyl-[acyl-carrier-protein] synthase, Class Q) in isolates from birds34. Our findings suggested that redundant genes might contribute to promoting niche specialization.
Redundant genes exhibit evolutionary expansion, potential functionality, and co-duplication with nearby translation initiation signals
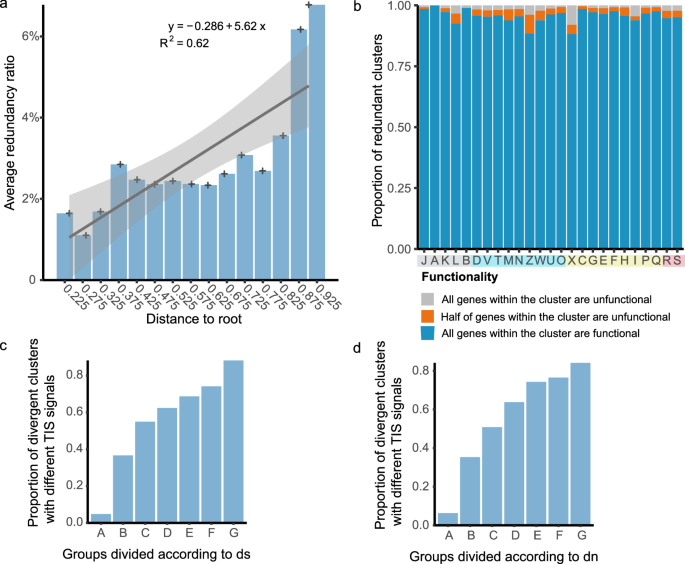
To uncover the evolutionary fate and consequences of redundant genes in prokaryotes, we calculated the phylogenetic distance of each species, based on the relatively complete phylogenetic tree provided by the All-Species Living Tree project35. We then grouped all genomes into 15 bins according to their phylogenetic distances with an interval of 0.05 and calculated the average redundancy ratio for each group. We found a positive linear relationship between the average phylogenetic distance and the average redundancy ratio (Fig. 3a, linear regression: R2 = 0.62, y = -0.286 + 5.62x, p-value < 0.05 in Set-80; Supplementary Fig. 5a, R2 = 0.74, y = 0.201 + 1.77x, p-value < 0.05 in Set-100). Across the majority of phyla, pathogens demonstrate a greater phylogenetic distance from the common ancestor compared to non-pathogens (Supplementary Fig. 5b). The results indicated that bacteria with greater phylogenetic distances were more likely to exhibit a higher redundancy ratio in their genomes, suggesting a trend of expanding redundant genes over the course of prokaryotic evolution.
a Relationship of redundancy ratio and phylogenetic distance in Set-80. The column shows the average redundancy ratio per bin, with the horizontal axis indicating phylogenetic distance. Plus signs mark mean distances, and linear regression with 95% confidence intervals depicts the relationship between redundancy ratio and phylogenetic distance. b Assessment of function in redundant clusters in all COG categories in Set-80. Based on the cognition that most clusters have a cluster size of two, clusters with a ratio > 0.67 were classified as unfunctional, <0.33 as functional, and those in between as semi-functional. c Relationship of translational divergence with synonymous substitution rate. Redundant clusters with ds ≤ 1 (78.5%) were binned at 0.2 intervals (groups A-F), while those with ds > 1 formed group G. The average synonymous substitution rate of each bin from A to G in the horizontal axis is 0, 0.07, 0.29, 0.49, 0.70, 0.90, 8.17, the vertical axis shows the proportion of clusters with changes in translation initiation in each bin. d Relationship of translational divergence with nonsynonymous substitution rate. All redundant clusters with dn ≤ 1 were binned at 0.02 intervals. The average nonsynonymous substitution rate of each bin from A to G in the horizontal axis is 0, 0.01, 0.03, 0.05, 0.07, 0.09, 0.13, and the vertical axis shows the proportion of clusters with changes in translation initiation in each bin. Source data are provided as a Source Data file.
Pseudogenization, recognized as a primary mechanism leading to gene loss36, is attributed to the accumulation of disabling mutations that cause the insertion of a premature stop codon or disrupt the reading frame37,38. After conducting a thorough search for premature stop codons in the recently formed redundant genes (see Methods), it was intriguing to find that only 5.9% of all redundant genes in Set-80 and 6.0% in Set-100 possessed these codons (Supplementary Fig. 5c, d), suggesting that the majority of these recently formed redundant genes did not evolve into pseudogenes due to premature stop codons. Additionally, an average of only 4.4% of the redundant genes in each genome possessed premature stop codons. An exceptionally high proportion of premature stop codons has been observed in the genomes of some Mycoplasma species (87.8% ± 14.4%), potentially due to these small parasitic bacteria evolving towards genome reduction39.
In order to infer whether redundant genes have potential functionality, we examined the 5’ upstream regions of all redundant genes and successfully detected translation initiation signal motifs in the majority of these genes, suggesting that most redundant genes had structurally complete upstream translation initiation signals for expression (see Methods). Due to the presence of premature stop codons and translation initiation signals serving as an indicator of gene functionality, we subsequently categorized the clusters of redundant genes into functional, semi-functional, and non-functional groups. The results revealed that 91.0% (89.7% in Set-100) of the clusters in Set-80 were functional, 3.4% (0.7% in Set-100) of the clusters were semi-functional, and 5.7% (9.7% in Set-100) of the clusters in Set-80 were unfunctional, indicating that most daughter genes potentially preserve functionality after duplication and short-term evolution. The higher proportion of semi-functional clusters in Set-80 compared to Set-100 hinted that over longer periods of evolution, one copy might sometimes degrade into a pseudogene while the other remained functional. Additionally, we found that high proportions of redundant gene clusters were functional across all COG categories, although variations were observed between different categories (Fig. 3b and Supplementary Fig. 5e). Specifically, the proportions of functional clusters were much higher in categories pertaining to vital functions, such as transcription, translation, and basic metabolism, while the categories related to cellular processes and signaling, as well as the mobilome, were more likely to lose functionality.
To better understand the duplication process in prokaryotes, we compared the translation initiation signal motifs in redundant gene pairs and discovered that 93.2% of these clusters in Set-100 possessed exactly identical signal motifs. Among the remaining 6.8%, further analysis using Tomtom40 showed that 40.3% of these clusters exhibited highly similar signal motifs, with at least a five-nucleotide overlap (Tomtom, p-value < 0.05). Given that the redundant genes in Set-100 can be regarded as having been recently duplicated, these findings suggested a co-duplication of translation initiation signals and coding sequences, with minor variations in translation initiation signals of some clusters possibly attributable to subsequent mutations.
However, identical translation initiation signal motifs were found in only 53.0% of the redundant gene clusters in Set-80, and 33.2% of the remaining clusters showed very similar signal motifs (Tomtom, p-value < 0.05). These relatively large variations compared to Set-100 prompted us to delve deeper into the evolution of translation initiation signals over time. We calculated the synonymous substitution rate (ds) and categorized them into groups A to F at intervals of 0.2, with all remaining ds values greater than one classified into group G. The results showed that the proportion of redundant gene clusters with divergent translation initiation signals increased with ds (Fig. 3c, r = 0.89, p-value < 0.05, groups A to F), peaking in group G. Likewise, redundant gene clusters with greater nonsynonymous substitution rates (dn) were more prone to experiencing mutations in translation initiation signals (Fig. 3d, r = 0.92, p-value < 0.01, groups A to F). These findings indicate that the translation initiation signals of redundant genes have predominantly evolved in conjunction with their coding sequences.
The roles of redundant genes in pathogens during clinical infections and evolution: case studies in A. baumannii and ECC
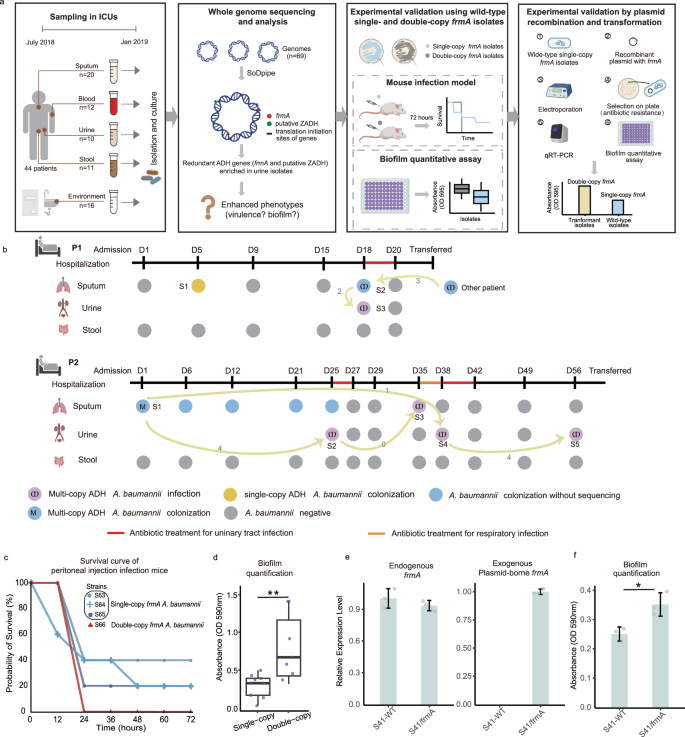
A. baumannii and ECC are notable nosocomial pathogens, with the capability to cause a wide range of infections, including septicemia, urinary tract infections, and pneumonia41,42. To elucidate the impacts of redundant genes and their translation initiation signals in clinical infections and evolution, we designed a time series observation in intensive care units (ICUs) of a tertiary hospital in Eastern China for six months. We collected a total of 69 A. baumannii strains, encompassing 53 strains from multi-sites of 44 patients in ICUs (sputum, n = 20; stool, n = 11; urine, n = 10; blood, n = 12) and 16 strains from environmental surfaces (Fig. 4a, Supplementary Data 5, the strains are designated S1 to S69). As a result, we identified 2,471 redundant clusters, containing 5,136 redundant genes, with each strain averaging 35 redundant clusters and 74 redundant genes. One ARG, eptA, was prevalently redundant (56/69, 81.2%) in the isolates. As for the virulence-related redundancy, one environmental isolate acquired redundant flmH, a gene specifically found in Aeromonas hydrophila for polar flagella formation, and another sputum isolate obtained redundant KPN_02274, an ompA gene from K. pneumoniae for the translocation of effector molecules of the type VI secretion system (Supplementary Fig. 6).
a Sampling design for the clinical implications of gene redundancy and experimental design to support the associations between ADH gene redundancy and enhanced pathogenicity. b Timeline of sample collection from two patients. Node colors indicate different isolation results and sequence types stratified by ADH gene copy number. The nodes linked by target curve arrows had the transmission relationships inferred by the cgSNV-based MST. The number of differential SNVs between the linked isolates was labeled on arrows. S1 to S5 represent different A. baumanni strains isolated from patients chronologically during the hospitalization. Periods of antibiotic administration are colored in the timeline. c Survival curves for mice infected by single- or double-copy frmA A. baumannii strains of the same clinically relevant lineage. The horizontal axis represents time after peritoneal bacterial injection, and the vertical axis represents the mouse survival rate. One strain with double copies of frmA (red) and three strains with a single-copy (blue) were tested, with five mice per strain (total n = 20). d The amounts of established biofilm on the plate lid of the 20 mice were measured. OD, optical density. p-value = 0.003, Wilcoxon rank-sum test, two-sided. Box plots indicate median (middle line), 25th, 75th percentile (box) and 5th and 95th percentile (whiskers) as well as outliers (single points). e Relative expression levels of endogenous frmA with S41-WT (A. baumannii S41) as the control strain and relative expression levels of exogenous plasmid-borne frmA with S41/frmA (A. baumannii S41/pYMAb2-frmA(ApmR)) as the control. f The amounts of established biofilm measured by a crystal violet staining assay at 595 nm. Error bars represent standard deviation, n = 3 independent replicates. Data are presented as mean values +/- SD. Source data are provided as a Source Data file. This figure was created with BioRender.com.
Remarkably, two genes within the ADH gene family, frmA (ADH class-III) and putative zinc-binding ADH (ZADH), were found as double copies (Set-80) in A. baumannii isolated from urinary tracts, in contrast to their single-copy presence in most strains from other sources (Fisher’s exact test, p-value < 0.05, Table 1, Supplementary Data 6, Supplementary Fig. 6). ADH serves as a pivotal element in modulating quorum sensing systems, enhancing bacterial motility, and facilitating biofilm formation and growth, with the latter responsible for nosocomial infections, especially in urinary tracts43.
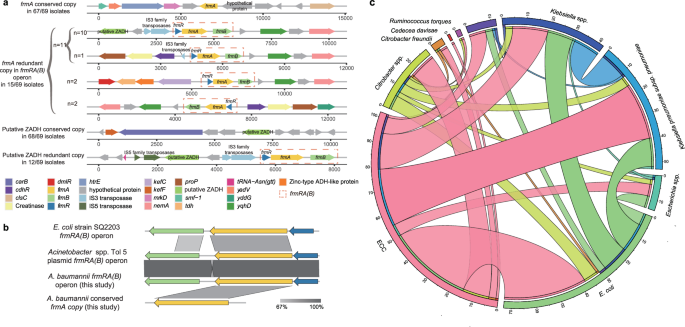
Upon examining the gene-environment surrounding frmA and ZADH copies, we observed that the singular frmA copy, flanked by two hypothetical proteins, remained conserved across the majority of strains (67/69, 97.1%). In contrast, the additional frmA copy, surrounded by frmR and frmB, was detected in only 21.7% (15/69) of the strains (Fig. 5a). These three genes constituted the frmRA(B) operon, which was situated within plasmid-related gene islands identified by IslandViewer and frequently positioned downstream of insertion sequences (ISs) (11/15, 73.3%). The frmRA(B) was early detected in E. coli, and a variant was recently detected in a plasmid, Acinetobacter spp. Tol 5 (pTol5, AP024709.1) (Fig. 5b). While both the frmA copies owned intact translation initiation signals, they displayed limited similarities regarding coding and translation initiation signals (Tomtom, p-value > 0.05). The additional frmA copy, along with its upstream translation initiation signal, nested within the frmRA(B) operon, was homogeneous between A. baumannii and pTol5 (Fig. 5b). Therefore, the frmRA(B) operon might have undergone HGT events from the close relatives of A. baumannii.
a The gene-environment of frmA copies and putative ZADH copies. b Comparison between frmRA(B) operons from A.baumannii (this study), E. coli strain SQ2203, and Acinetobacter sp. Tol 5 plasmid, as well as the conserved frmA copy in A.baumannii (this study). c HGT of redundant genes between ECC and human gut microorganisms.
Regarding the putative ZADH gene copies, while one copy surrounded by two hypothetical proteins was consistently retained across most strains (68/69, 98.6%), the additional copy, positioned downstream of a phage fragment, tRNA-Asn(gtt) and ISs within plasmid-related gene islands identified by IslandViewer, was observed in 17.4% (12/69) of the strains (Fig. 5a). The translation initiation signals of both copies were intact and significantly similar (Tomtom, p-value < 0.05) in most cases (11/12, 91.7%). It has been reported that tRNA-Asn(gtt) serves as a recognition site for recombination or counterbalancing the compositional differences between the phage and bacterial genomes to adjust the translation capacity during bacterial infection44. This suggests that the observed redundancy likely arose from recombination events and offers further evidence for the efficient expression of the putative ZADH gene.
Interestingly, the two ADH genes were found to be co-located and closely situated within the A. baumanni genome in all the redundancy events of putative ZADH. Specifically, the redundant ZADH was located upstream of the IS-frmRA(B) structure with a hypothetical protein in between (Fig. 5a). We defined isolates with this kind of genetic arrangement as the multi-copy ADH genotype. To elucidate the acquisition and transmission dynamics of multi-copy ADH genotype A. baumanni isolates, we therefore investigated longitudinal isolates from two patients (P1 and P2) who developed nosocomial urinary infections after ICU admission (Fig. 4b). By constructing a minimum spanning tree (MST) based on core genome single nucleotide variations (cgSNVs), we revealed both within-host persistence and between-host transmission of the pathogen. Notably, the multi-copy ADH genotype A. baumanni isolates invaded urinary tracts after the initial colonization in the respiratory tracts in both P1 and P2. In P2, this invasive urinary infection even persisted for a month, with the multi-copy ADH genes and their translation initiation signals remaining stable over time. Our findings suggest that the redundancy in the ADH gene family potentially offers the ability of colonized A. baumannii strains to invade urinary tracts and establish persistent infections.
To validate the association between ADH redundancy and virulence, we conducted mouse peritoneal infection assays using four clinical isolates with single-copy frmA or double-copy frmA, with five mice per strain. Results showed that all mice infected with double-copy frmA strains succumbed within 24 hours, whereas those infected with single-copy frmA strains exhibited a higher survival rate throughout the 72-hour observation (Fig. 4c). The results suggest that double-copy frmA isolates exhibit increased virulence. As ADH has been reported to play a role in biofilm formation43, we further investigated whether its redundancy enhances the biofilm mass in A. baumannii. Our quantitative biofilm assays revealed that isolates with double-copy frmA (n = 7) produced significantly more biofilm mass compared to single-copy frmA isolates (n = 9, Fig. 4d, p-value < 0.05, Wilcoxon rank-sum test).
Furthermore, we validated the effect of frmA redundancy by bacterial transformation experiments. We constructed a plasmid carrying an extra copy of frmA and introduced it into two single-copy frmA isolates, A. baumannii S41 and A. baumannii S65, randomly selected from the clinical isolates (Supplementary Fig. 7a and b, see Methods, Table 2). Quantitative reverse transcription PCR (qRT-PCR) confirmed the successful introduction and expression of the frmA copy, which significantly increased biofilm formation in both strains (A. baumannii S41: Fig. 4e, f; p-value = 0.02, Student’s t-test; A. baumannii S65: Supplementary Fig. 7c; p-value = 0.078, Student’s t-test). When we further enhanced the frmA expression in A. baumannii S65 by replacing its native promoter with that of the highly conserved gene ompA, biofilm formation increased accordingly (Supplementary Fig. 7c; p-value < 0.05, Student’s t-test). These results support a positive association between frmA expression levels and biofilm mass in A. baumannii (Supplementary Fig. 7c).
To gain further insights into the influence of redundant genes on resistance and virulence in pathogens, we downloaded 977 ECC genome assemblies from the NCBI and successfully reclassified 898 into six species17. Of these, 714 belonged to E. hormaechei, 65 to E. asburiae, 50 to E. cloacae, 26 to E. kobei, 25 to E. ludwigii and 18 to E. roggenkampii (see Methods). Using SoDpipe, we thus identified a total of 30,120 redundant clusters within 898 ECC genomes, containing 64,821 redundant genes. The redundancy ratio ranged from 0.2% to 7.9% in each ECC genome, averaging 1.5% (Supplementary Data 7).
The high proportion of E. hormaechei in our dataset, alongside its epidemiological predominance among ECC species in clinical and environmental settings42,45,46, motivated us to investigate whether redundant genes contribute to its dominance. By comparing E. hormaechei with all other ECC species, we discovered that 15 of 21 resistance and virulence-related redundant genes, including those conferring resistance to bactericidal compounds, anaerobic growth, mercury, and chemotaxis proteins, were significantly more prevalent in E. hormaechei (Fisher’s exact test, p-value < 0.05, Supplementary Data 8). Further analysis of these gene clusters suggested that most of them are potentially functional (Supplementary Data 8). Notably, 72.3% (516/714) of the E. hormaechei genomes possessed multiple copies of alkyl/aryl-sulfatase, a gene implicated in resistance to bile salts and sodium dodecyl sulfate47, whereas such redundancy was absent in other ECC species. Moreover, correlation analysis using a presence-absence matrix showed strong co-occurrence among certain redundant genes, particularly merR (P13111), merT (Q51769), and merP (P0A216), which are co-localized within the mercury resistance operon (mer; Supplementary Fig. 8a, b; Supplementary Data 8). Subsequent genome analysis of an isolate (GCF_000724505) demonstrated that two copies of the mer operon reside on separate plasmids, each carrying different antibiotic resistance and virulence genes. Among the 42 E. hormaechei isolates harboring redundant mer operons, we observed that the redundancy occurred in plasmid-associated regions in 61.9% isolates (26/42), suggesting that plasmid-mediated HGT commonly contributes to the spread and formation of mercury-resistance redundancy in E. hormaechei. As previously reported, the mer operon is often co-localized with other beneficial genes, such as those conferring antibiotic resistance, which may further enhance fitness48,49. In E. hormaechei, the retention of multiple plasmids carrying the mer operon not only strengthens mercury resistance but also enhances the overall fitness and competitiveness of the host strain across diverse environments. Overall, the high prevalence of these resistance- and virulence-related redundant genes in E. hormaechei likely contributes to its competitive advantage over other ECC species in different environments.
The redundant genes are primarily involved in membrane, transposition, metal resistance, and DNA recombination (Supplementary Fig. 8c), also indicating the pivotal role of HGT in acquiring multiple copies of resistance-related genes in the ECC genomes. To further detect HGT between ECC and their co-occurrent strains, we downloaded 456 human gut bacterial strains from the Human Microbiome Project (HMP) (Supplementary Data 9) and found that 1,218 of 48,565 genes in the ECC pangenome may transfer among gut bacteria via HGT. Among them, 59 were redundant genes (Fig. 5c and Supplementary Data 10). Notably, these transfers occurred almost exclusively among Gram-negative bacteria, particularly between ECC and species such as E. coli, K. pneumoniae, and Klebsiella species. The functions of the transferred redundant genes are largely associated with resistance and virulence, including resistance to silver, copper, and arsenic, as well as toxin-antitoxin systems. These findings highlighted the role of E. coli and Klebsiella as reservoirs for the horizontal transfer of multiple copies of resistance and virulence-related50,51, corroborating the aforementioned observations of A. baumannii.
We further discovered a gene island (Supplementary Data 11) that introduced 13 redundant genes along with 27 non-redundant genes, classified as a Copper and Silver Resistance Island (CSRI). This island included an array of metal resistance genes: five cation efflux system proteins, four copper resistance proteins, two silver-binding proteins (SilE), one putative copper-binding protein (PcoE), one sensor kinase (CusS), one transcriptional regulatory protein (CusR), one transcriptional activator protein (CopR), and one silver-exporting P-type ATPase. The identification of three insertion sequences (ISs) within the CSRI suggested that these metal resistance genes were likely gathered via upstream ISs. Given the extensive use of copper and silver ions as biocides in medical and healthcare fields, ECC’s enhanced ability to cope with copper and silver stress likely facilitates its survival52. Furthermore, since copper is essential for host defense, bacteria that develop copper resistance may better evade immune responses, representing a patho-adaptive strategy52. Similar to our observations on the mer operon, ample evidence indicates a positive correlation between heavy metal resistance and antibiotic resistance, with metal resistance facilitating the maintenance and spread of antibiotic resistance through co-selection53. Collectively, these findings shed light on the mechanisms underlying the transfer and persistence of redundant genes in hospital environments, highlighting the selective advantages they confer in colonization and pathogenicity.