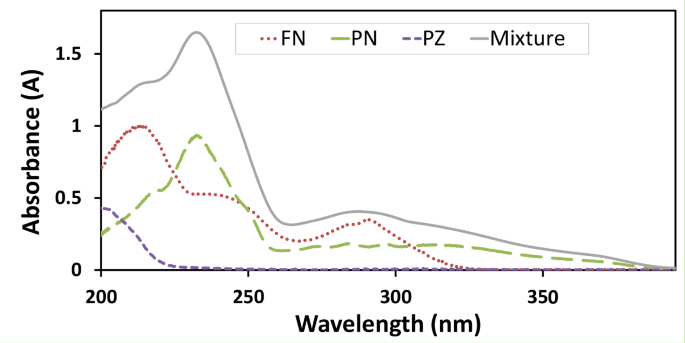
The simultaneous quantification of FN, PN, and PZ is of significant importance in veterinary medicine, as this combination offers broad-spectrum anthelmintic efficacy, simplifying treatment regimens and improving animal health. However, as illustrated in Fig. 1, the UV-Vis absorption spectra of FN, PN, and PZ in absolute ethanol exhibit substantial spectral overlap across a considerable wavelength range (200–300 nm). The broad and closely positioned absorption maxima of these analytes complicate their simultaneous determination using conventional UV-Vis spectrophotometric techniques. The spectrum of a ternary mixture (Fig. 1) further underscores this challenge, representing the additive spectral contributions of each component and highlighting the complexity of deconvoluting individual analyte concentrations. A 3D plot of the entire spectral dataset is provided in Supplementary Fig S2, which further illustrates the severe spectral overlap. To overcome these limitations and enable accurate simultaneous quantification, we developed a novel approach integrating AP-SPA for optimal wavelength selection with AT-SVR for model development.
Experimental design (Calibration and test Sets)
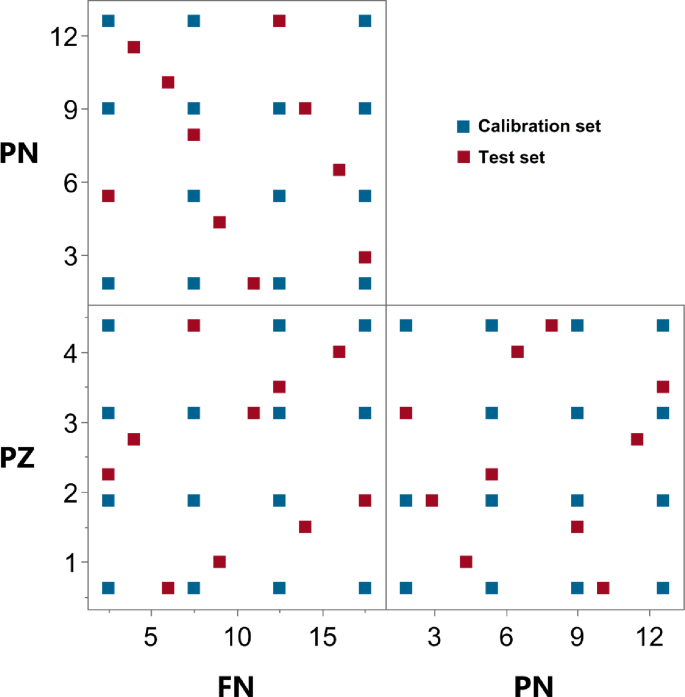
Developing robust and generalizable chemometric models for quantitative spectral analysis necessitates a carefully designed calibration set. An inadequately constructed calibration set can compromise model accuracy and limit its predictive capability for new, unknown samples31. To account for potential spectral interactions between FN, PN, and PZ, a multilevel experimental design was implemented to generate a calibration set comprising 16 mixtures (Supplementary Table S1). This design ensures that the concentrations of FN, PN, and PZ are varied independently, minimizing correlations between analyte concentrations and facilitating a more accurate assessment of their individual and combined spectral contributions. While the sample size is modest, this systematic design is highly efficient for capturing complex relationships in multivariate systems. For independent model validation, a test set of 10 samples was generated using a LHD, a space-filling technique chosen for its ability to provide representative coverage of the experimental domain with a minimal number of points24. The robust performance of the model on this external test set validates the sufficiency of our design. Figure 2 presents the scatter matrix plot visualizing the distribution of both calibration and test set samples within the three-dimensional concentration space of FN, PN, and PZ. This plot visually confirms the space-filling characteristics of both designs, demonstrating adequate representation of the experimental domain.
Optimization of chemometric model parameters
The predictive performance of the chemometric model is highly dependent on the selection of key parameters. To ensure the robustness and accuracy of our proposed method, a sensitivity analysis was conducted to optimize the variance threshold for data preprocessing and the plateau threshold for the AP-SPA stopping criterion. The effect of the variance threshold on the model predictive MSE is presented in Supplementary Fig S3. While the absolute minimum MSE for each analyte occurs at slightly different thresholds, a value of 0.1 was selected as it represents the best overall compromise, yielding consistently low prediction errors across all three analytes simultaneously (Supplementary Fig S3-A). This ensures that spectral noise is effectively filtered without removing crucial analytical information for any of the components.
The optimization of the AP-SPA plateau threshold is shown in Supplementary Fig S3-B. The results clearly indicate that a threshold of 0.0001 corresponds to the minimum achievable MSE for all three analytes. A less strict threshold results in higher error, while stricter thresholds show a trend of increasing error. Therefore, 0.0001 was confirmed as the optimal stopping criterion and used for all subsequent model development.
Adaptive Plateau-based successive projections algorithm (AP-SPA) for wavelength selection
To address the challenge posed by the significant spectral overlap, the AP-SPA was employed for optimal wavelength selection32. Unlike traditional SPA, which typically selects a pre-defined number of variables, AP-SPA incorporates an adaptive stopping criterion based on a plateau threshold (0.0001). This adaptive approach monitors the reduction in MSE during successive SPA iterations. The algorithm terminates when the improvement in MSE falls below this threshold, effectively preventing the inclusion of irrelevant or redundant wavelengths and mitigating the risk of model overfitting. Application of AP-SPA to the UV-Vis spectral data of FN, PN, and PZ mixtures resulted in a parsimonious selection of seven highly informative wavelengths. The specific wavelengths selected for the final model were [224, 262, and 343 nm] for FN, [208 and 328 nm] for PN, and [207 and 238 nm] for PZ. Supplementary Fig S4 visualizes these selected wavelengths, superimposed on the individual UV-Vis spectra of each analyte, highlighting their correspondence to regions of maximum absorbance and/or spectral differentiation. This targeted wavelength selection minimizes data redundancy and collinearity, leading to a more robust and computationally efficient calibration model. AP-SPA demonstrated enhanced computational efficiency compared to traditional SPA and exhaustive search methods, significantly accelerating the subsequent model building and validation steps by adaptively stopping the selection process and reducing data dimensionality.
Automated tuning support vector regression (AT-SVR) for quantitative modeling
To effectively model the potentially non-linear relationships between the UV-Vis spectra and the concentrations of FN, PN, and PZ, SVR with a Radial Basis Function (RBF) kernel was integrated with SPA. While traditional SPA often utilizes PLSR, PLSR linear nature can limit its performance when dealing with complex datasets exhibiting non-linearity32. To optimize the SVR model performance, a dual-stage hyperparameter tuning strategy was implemented exclusively on the calibration set using an internal 5-fold cross-validation (CV) procedure. This strategy combined a randomized search for broad exploration of the parameter space with a subsequent grid search for precise refinement. The performance of each hyperparameter combination was assessed based on its average cross-validated MSE across the five folds. Once the optimal hyperparameters were identified, a final model was trained on the entire calibration set and then evaluated a single time against the strictly external test set to ensure an unbiased assessment of its generalizability. Randomized search efficiently identified promising regions within the hyperparameter space, while grid search systematically evaluated all hyperparameter combinations within these refined regions to pinpoint the optimal parameter set. Supplementary Fig S5 illustrates this two-stage optimization, demonstrating the rapid identification of low-MSE regions by randomized search (upper panel) and the precise location of the global MSE minimum by grid search (lower panel). Table 1 compares the performance of the AP-SPA/AT-SVR method to PLSR for the simultaneous quantification of FN, PN, and PZ, based on MSE and R² for both calibration and test sets. Table 1 compares the performance of the AP-SPA/AT-SVR method to the traditional PLSR model. While the PLSR model showed a slightly lower calibration MSE for FN, the AP-SPA/AT-SVR model demonstrated superior predictive capability on the independent test set for all three analytes, as evidenced by lower MSE and higher R² values. This superior generalizability is the most crucial indicator of a model’s real-world utility. The advantage of our proposed method is particularly evident for PZ, where AP-SPA/AT-SVR significantly reduced the test set MSE from 0.09 (PLSR) to 0.02 and increased the R² from 0.941 to 0.972. These results demonstrate the superior predictive capability of the AP-SPA/AT-SVR approach, highlighting the synergistic benefits of adaptive feature selection and automated SVR modeling for complex spectral data analysis. Furthermore, to ensure the model was not overfitted, a residuals analysis was performed on the external test set (Supplementary Fig. 6). The plots show a random, non-systematic distribution of Studentized residuals around zero, confirming that the model is robust and generalizes well to new data.
Method validation using accuracy profile approach
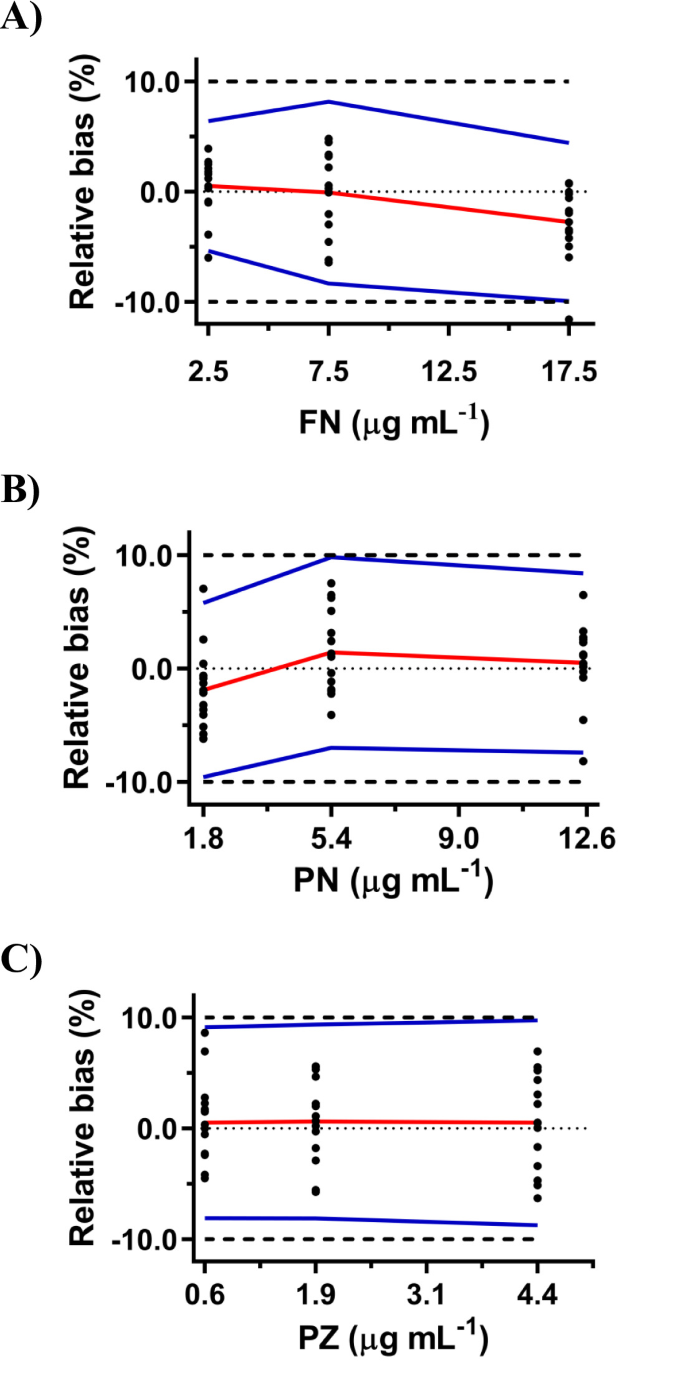
The proposed spectrophotometric method was validated for accuracy, precision, trueness, and linearity for the simultaneous quantification of FN, PN, and PZ, following the accuracy profile approach as outlined in the SFSTP guidelines26,27,28,29. The comprehensive validation results are summarized in Table 2. The relative bias values for all three analytes, consistently within ± 5% across the tested concentration ranges (Table 2), indicate excellent trueness of the method. Similarly, the repeatability and intermediate precision, expressed as Relative Standard Deviation (RSD), were consistently below the acceptance criterion of ≤ 5% across all concentrations (Table 2), demonstrating satisfactory precision both within-day and between-days. The accuracy profiles, visualized in Fig. 3, confirm that the β-expectation tolerance intervals fall well within the acceptance limits of ± 10% for all analytes across their validated concentration ranges. The relative expanded uncertainties were also within the acceptance criterion of ≤ 10% (Table 2). It is important to note that the measurement uncertainty was calculated using a “top-down” approach, which is directly integrated with the accuracy profile methodology33,34. This approach computes the total uncertainty by combining the variance from the intermediate precision—which empirically captures multiple random error sources from the validation design—with the uncertainty associated with the measurement bias. This provides a holistic and practical estimate of the uncertainty based on the method’s observed performance, as detailed in the cited literature35,36. The low risk values (all < 5%, Table 2) further substantiate the method reliability. Linearity, assessed over the concentration ranges detailed in Table 2, was excellent, with regression analysis yielding R² close to unity, slopes near 1, and intercepts approaching zero for all three analytes. For a direct graphical assessment of the model linearity and lack of bias, predicted versus actual concentration plots are provided in Supplementary Fig S7. In accordance with the total error approach, the validated quantification range of the method is empirically defined by the accuracy profiles (Fig. 3). The Lower Limit of Quantification (LLOQ) is established as the lowest concentration for which the method total error (encompassed by the β-expectation tolerance interval) is confirmed to be within the acceptance limits. Based on this criterion, the LLOQs for FN, PN, and PZ were determined to be 2.5 µg mL− 1, 1.8 µg mL− 1, and 0.6 µg mL− 1, respectively.
Accuracy profiles obtained with Adaptive Plateau-based Successive Projections Algorithm with Automated Tuning Support Vector Regression (AP-SPA / AT-SVR for A) fenbendazole (FN), (B) Pyrantel embonate (PN), and (C) Praziquantel (PZ). Relative bias (red line), ± 10% acceptance limits (—), 95% β-expectation tolerance limits (blue line), and relative back-calculated concentrations (•).
Application to veterinary dosage form analysis
To evaluate its practical applicability, the developed AP-SPA/AT-SVR UV-spectrophotometric method was applied to determine the content of FN, PN, and PZ in commercially available Cestal Plus veterinary chewable tablets. The results obtained with the proposed spectrophotometric method were statistically compared to those from a reported HPLC reference method4 using both t-tests and F-tests. Representative chromatograms confirming adequate separation and peak shape for the reference method are provided in Supplementary Fig S8. The statistical evaluation (Table 3) revealed no statistically significant difference (p >0.05) between the two methods, demonstrating excellent agreement. This strong concordance confirms that potential interference from excipients was negligible under the established sample preparation and dilution protocol. This confirms the accuracy and reliability of the proposed UV method for routine analysis of Cestal Plus tablets and highlights its potential to replace more complex, time-consuming, and costly chromatographic techniques for quality control purposes.
Greenness and whiteness assessment
This multi-tool approach to sustainability is consistent with the current state-of-the-art in green analytical chemistry, where a holistic view is essential for a true assessment of a method’s environmental impact. Recent studies have demonstrated the power of combining greenness, blueness, and whiteness metrics to provide a comprehensive sustainability profile for complex pharmaceutical analyses37,38. Our work builds on this paradigm by applying these principles to a novel chemometric model for veterinary pharmaceuticals, confirming that high analytical performance can be achieved in concert with significant environmental benefits. The GSST assigned a high greenness score to ethanol, the solvent used in this method, indicating its environmentally benign nature compared too many organic solvents commonly employed in HPLC methods (Supplementary Fig S9). The NEMI pictogram assessment yielded a fully green profile, signifying adherence to NEMI sustainability criteria. This favorable NEMI profile is primarily attributed to the use of ethanol, the absence of hazardous reagents, and the inherently low waste generation of UV spectrophotometry (Table 4). The modified Eco-Scale assigned a score of 84 (out of 100), reflecting the minimal use of hazardous substances and low waste production (Table 4 and Supplementary Table S3). The MoGAPI tool yielded a high score of 81, attributed to the simple and solvent-efficient sample preparation and the inherent greenness of UV spectrophotometry, which avoids extensive chromatographic separation and associated solvent consumption (Table 4 and Supplementary Table S4).
Furthermore, to provide a more holistic evaluation, a whiteness assessment was conducted based on the principles of White Analytical Chemistry (WAC). WAC complements greenness by balancing analytical performance (Red), eco-friendliness (Green), and practical/economic efficiency (Blue) into a single, unified metric39,40. We employed the Red-Green-Blue 12 (RGB12) algorithm for this purpose, as it provides a quantitative measure of a method’s overall sustainability and practicality41. The RGB12 algorithm yielded a high score of 91.9 for the proposed method, reflecting its excellent simplicity, directness, and use of a safe, readily available solvent (Table 4 and Supplementary Tables S5-S7). In contrast, typical HPLC methods for similar multi-analyte determinations often require significant volumes of hazardous organic solvents, generate substantial waste, and involve more complex sample preparation steps, resulting in lower whiteness scores. Therefore, the proposed spectrophotometric method presents a significantly greener and more sustainable analytical alternative, aligning with the growing emphasis on environmentally responsible analytical practices. Therefore, the proposed spectrophotometric method presents a significantly greener and more sustainable analytical alternative, aligning with the growing emphasis on environmentally responsible analytical practices. While the proposed AP-SPA/AT-SVR method offers significant advantages in terms of speed, cost-effectiveness, and environmental sustainability compared to HPLC, certain limitations must be acknowledged. First, UV-Vis spectrophotometry generally exhibits lower sensitivity than HPLC-MS/MS, making it less suitable for detecting trace-level impurities or metabolites in biological fluids. Second, the success of the chemometric model is heavily dependent on the design of the calibration set; any significant variation in the matrix excipients not accounted for in the calibration model could affect prediction accuracy. Finally, unlike HPLC, which physically separates components, this method relies on mathematical resolution, requiring specialized software and statistical expertise for model maintenance.