A reductionist model identifies CD4 T cell dysfunction during chronic Mtb infection
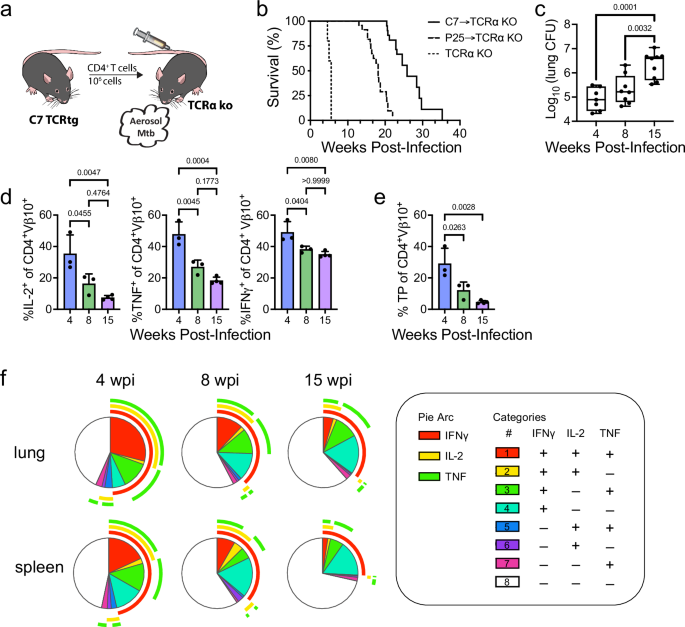
To study antigen-specific CD4 T cells in the setting of chronic Mtb infection, we developed an adoptive transfer model (Fig. 1a). We observed that intravenous transfer of P25 (i.e., Ag85b-specific CD4) TCR transgenic (TCRtg) T cells to TCRα KO mice prolonged the median survival time (MST) of recipient mice from five weeks to 18 weeks (Fig. 1b)18. Under the same infection conditions, B6 mice survive > 48 weeks19,20. P25 T cells gradually lose the capacity for cytokine production during chronic infection, but the diminished expression of Ag85b by Mtb in vivo raised the possibility that recrudescence was secondary to the inability of P25 T cells to recognize Mtb-infected cells21. Therefore, we transferred C7 (i.e., ESAT-6-specific CD4) TCRtg T cells into TCRα KO mice (C7→TCRα). C7 T cells conferred long-term protection, but ultimately the mice succumbed with an MST of 26 weeks, which is significantly reduced compared to B6 mice (Fig. 1b). The greater susceptibility of C7→TCRα mice was accompanied by a progressive increase in lung bacterial burden (Fig. 1c). We used this model to investigate why immunity failed.
a Schematic of C7→TCRα adoptive transfer model. Illustration from NIAID NIH BIOART Source (bioart.niaid.nih.gov/bioart/281 and 506)” (b) Survival curves of infected TCRα KO, P25→TCRα KO (n = 20), and C7→TCRα KO (n = 13). The difference between the groups was statistically significant (p < 0.0001) as determined by the log-rank test. c Lung CFU of infected C7→TCRα mice at 4, 8, and 15 wpi. Data is pooled from two independent experiments, with n = 7-9 mice per time point. Box plots indicate median (middle line), 25th, 75th percentile (box) and minimum and maximum (whiskers). d Frequencies of IL-2, TNF, or IFNγ or (e) Triple producing (TP; IL-2+TNF+IFNγ+) C7 T cells. Representative data of three independent experiments, n = 3-4 mice per time point. Bars, mean ± SD. f Cytokine production and T cell polyfunctionality visualized using SPICE plots. c–e Statistical significance was analyzed by one-way ANOVA with Bonferroni’s multiple comparison test. Source data are provided as a Source Data file.
An advantage of the C7→TCRα model is the large number of antigen-specific T cells that can be assessed independently of their function. Purified CD4 T cells from C7 mice contained variable numbers of non-transgenic naïve CD4 T cells (see methods), so ESAT6-specific CD4 T cells were identified using ESAT6/IAb (E6) tetramers or anti-Vβ10 antibodies (Supplementary Fig. 1a). Cytokine production in response to ESAT6 peptide stimulation was used as a measure of T cell function. The frequency of C7 T cells producing IL-2, TNF, or IFNγ decreased over time, accompanied by a loss of polyfunctional T cells and an increase of non-cytokine-producing T cells (Fig. 1d–f). Thus, diminished function of ESAT6-specific CD4 T cells correlates with increasing bacillary burden.
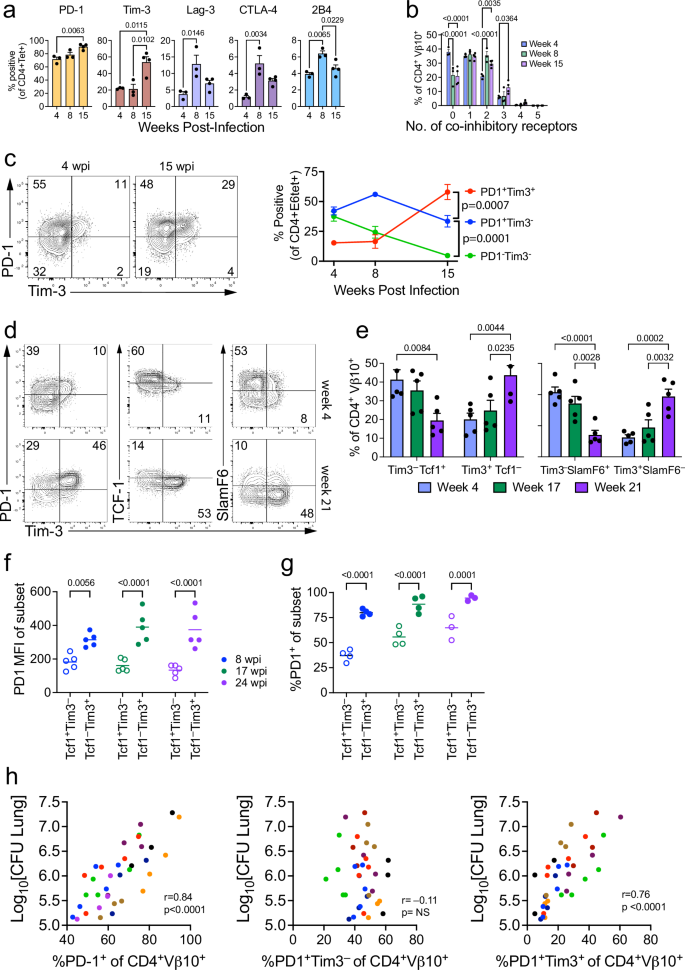
To determine whether loss of function was associated with the acquisition of co-inhibitory receptor expression, PD-1, TIM-3, LAG-3, CTLA-4, and 2B4 expression was measured between 4- and 15-weeks post-infection (wpi). PD-1 and TIM-3 were the most highly expressed, and the number of co-inhibitory receptors expressed by C7 T cells increased over time (Fig. 2a, b). The frequency of PD-1+TIM-3+ C7 T cells increased over time, a characteristic of terminally exhausted CD8 T cells (Fig. 2c)13,22. In parallel, C7 T cells resembling progenitor exhausted CD8 T cells (TPEX; i.e., TIM-3–TCF-1+ or TIM-3–SLAMF6+) diminished while those resembling terminally exhausted CD8 T cells (TTEX; i.e., TIM-3+TCF-1– or TIM-3+SLAMF6–) increased (Fig. 2d, e)13,22. The percentage of TIM-3+TCF-1– C7 T cells expressing PD-1 was higher compared to their TIM-3–TCF-1+ counterpart (Fig. 2f). Consistent with a terminally exhausted phenotype, PD-1 levels were significantly higher on TIM-3+TCF-1– C7 T cells (Fig. 2g). Although the kinetics of the expression of co-inhibitory receptors varied between experiments, we always observed CD4 T cells expressing high levels of PD-1 and TIM-3 late during infection
a Frequencies of CD4+ ESAT-6 Tetramer+ C7 T cells expressing PD-1, TIM-3, Lag-3, CTLA-4, or 2B4. b Percentage of C7 T cells (from ‘a’) expressing 0–5 co-inhibitory receptors in the lungs of infected mice. c Representative flow plots and quantification of C7 T cells expressing PD-1 and/or TIM-3. d Representative flow plots of C7 T cells expressing TCF-1, SLAMF6, or PD-1, and TIM-3, at 4 and 21 wpi. e Frequencies of C7 T cells expressing TCF-1, SLAMF6, or TIM-3. f, g PD-1 median fluorescence intensity (MFI) of TCF-1+TIM-3– and TCF-1–TIM-3+ C7 T cells (f) and the percentage of TCF-1+TIM-3– and TCF-1–TIM-3+ C7 T cells expressing PD-1 (g). h Correlation of lung CFU with percentage of PD-1+, PD-1+TIM-3–, or PD-1+TIM-3+ C7 T cells. Independent experiments, each with two or three timepoints are indicated by different colors. Representative data of two (a, b) or more than three (c–g) independent experiments. h Cumulative data from 2 independent experiments. a–c n = 3, 3, and 4/group at 4, 8, 15 wpi, respectively. d–f n = 5/group. g n = 4, 4, and 3/group at 4, 17, 24 wpi, respectively. Data represent mean ± SEM. Statistical significance was analyzed by two-way ANOVA (e–g) with Bonferroni’s (a), Tukey’s (c, e), Fisher’s LSD multiple comparison test, or one-way ANOVA with Tukey’s multiple comparison test (b). Pearson correlation (two-tailed) was calculated using data from three independent experiments consisting of 36 individual mice (h). Source data are provided as a Source Data file.
We next asked whether there was a correlation between the exhausted phenotype of C7 cells and progressive lung infection. Analysis of all individuals from two independent experiments encompassing multiple time points demonstrated a strong positive correlation between the frequency of PD-1+ C7 T cells and CFU (Fig. 2h, left). As PD-1 is also a marker of activated T cells, we next determined the correlation between PD-1+TIM-3– (i.e., activated or TPEX) and PD-1+TIM-3+ (TTEX) C7 cells. There was no significant correlation between the PD-1+TIM-3– phenotype and lung CFU; in contrast, there was a striking correlation with the PD-1+TIM-3+ C7 phenotype, which was largely independent of time point or experiment (Fig. 2h). While the high number of C7 cells could create competition for antigen and APC in this model, we suggest that this would have the opposite effect on T cell exhaustion: namely, that less TCR stimulation would preserve function. These data suggest that the emergence of CD4 T cell exhaustion late during infection could contribute to recrudescence and disease. However, the causal relationship remains to be established. Development of T cell exhaustion could result in loss of Mtb control, or alternatively, an increase in bacillary burden and antigen load could result in T cell exhaustion. Nevertheless, these data show that the development of terminally exhausted CD4 T cells correlates with ongoing bacterial replication and disease progression in this model.
Exhaustion contributes to CD4 T cell dysfunction in an adoptive transfer model
Based on the co-expression of PD-1 and TIM-3 by a large fraction of C7 CD4 T cells during the chronic phase of infection in this model, which resemble CD8 TTEX cells, we hypothesized that C7 CD4 T cells were becoming exhausted11,13. To test this possibility, lung parenchymal CD4 T cells from C7→TCRα mice were sorted (Supplementary Fig. 1b), transcriptionally profiled at 4 and 25 wpi by RNAseq and compared to splenic CD4 T cells from uninfected C7 TCRtg mice (Fig. 3a).
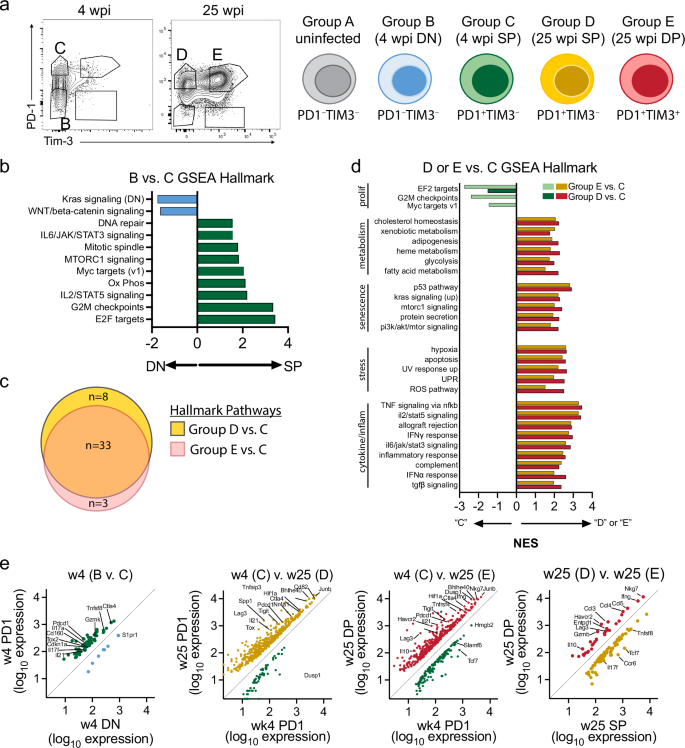
a Flow plots and schematic of sorted cell populations at 4 and 25 wpi used for RNAseq analysis. The color of the cells visually represents the different cell populations in (b–e). b GSEA results of mouse-ortholog Hallmark pathways significantly enriched among Group B or Group C C7 T cells. c Venn diagram representing the overlap of Hallmark pathways significantly enriched in Groups D or E compared to Group C. d GSEA Hallmark pathways significantly enriched among Group D or Group E compared to Group C. e Scatter plots of DEGs ( | log2FC | ≥ 1 and Padj < 0.05) between indicated groups. NES, Normalized Enrichment Score; False Discovery Rate (FDR), < 0.05 for all pathways shown. For each cell population analyzed, there were 4 (Groups B, C) or 5 (Groups D, E) biological replicates. Source data are provided as a Source Data file.
At 4 wpi, CD4 T cells from C7→TCRα mice were mostly PD-1–TIM-3– (Double Negative, DN; Group B) or PD-1+TIM-3– (Single Positive; SP; Group C). Compared to CD4 T cells from uninfected C7 mice, 3777 and 4268 genes, respectively, were upregulated (log2FC≧1, FDR < 0.05) (Supplementary Data 1). Jun and Fos were among the most highly induced genes, and Gene Set Enrichment Analysis (GSEA) identified gene signatures associated with T cell activation in both Group B and C C7 CD4 T cells. Only 62 differentially expressed genes (DEGs) were identified between Group B and C cells (Supplementary Data 2). Among these genes were Il17a, l17f, Il21, and Tox2, suggesting that the PD1+ C7 CD4 T cells contains Th17- and Tfh-like cells. The EF2 targets, G2M checkpoint, and IL-2/STAT5 signaling Hallmark Pathways were enriched among Group C cells, indicating that PD1+ C7 CD4 T cells undergo greater proliferation and activation than Group B cells (Fig. 3b).
By 25 wpi, nearly all C7 CD4 T cells were PD1hiTIM3– (Group D) or PD-1hiTIM-3+ (Double Positive; DP; Group E) (Fig. 3a). Most Hallmark pathways that were significantly enriched were identified both in group D and E when compared to group C (Figs. 3c, Supplementary Data 3, 4). The only Hallmark pathways upregulated in group C were “E2F targets,” “G2M checkpoints” and “Myc targets v1,” consistent with the greater proliferative activity of CD4 T cells during the early phase of Mtb infection (Fig. 3d). The sharing of upregulated pathways by Group D and E suggests that the difference in the duration of infection (i.e., 4 vs 25 wpi) was more important in determining the transcriptional program of the cells than their expression of co-inhibitory receptors (i.e., PD1hiTIM3– vs. PD-1hiTIM-3+). Highly enriched signaling pathways in groups D and E included those induced by T cell cytokines (IL-2, IFNγ), inflammatory cytokines (IL-6, TNF, IFNα), and anti-inflammatory cytokines (TGFβ). Lipid and other metabolic pathways were also detected (fatty acid metabolism, cholesterol homeostasis, glycolysis). Genes from pathways associated with cellular stress were enriched in C7 CD4 T cells from 25 wpi including “apoptosis,” “hypoxia,” “ROS pathway,” “UPR,” and “UV response up,” which often indicate cell stress and DNA damage. The upregulation of these pathways could be driven by the infection and might contribute to T cell dysfunction.
Compared to Group C (4 wpi), 463 genes in Group D and 489 genes in Group E significantly changed expression, and 80% of these were upregulated at 25 wpi (Supplementary Data 5, 6). Ccr4, Ccr6, Il17f, Il17re, and Il23r, were expressed more at 4 wpi, suggesting that Th17 responses diminished during chronic infection. The expression of negative regulators and co-inhibitory receptors by Group D and E C7 CD4 T cells (PD1hiTIM3– and PD1hiTIM3+, respectively) were increased (e.g., Ctla4, Entpd1, Foxp3, Lag3, Nfil3, Pdcd1, and Tigit) (Fig. 3e, Supplementary Data 5, 6). Tox and Tox2, and Nr4a1, Nr4a2, and Nr4a3, are important transcriptional regulators of CD8 T cell exhaustion and were increased among Group D and E C7 CD4 T cells. Among the Group E CD4 T cells, we noted increased expression of genes associated with CD4 T cell exhaustion (e.g., Prdm1, Il10, Il21)23 and terminal differentiation (Klrg1 and Tbx21). Thus, both 25 wpi CD4 T cell populations had features resembling exhausted CD8 T cells observed in tumors and chronic viral infection13,24,25.
Only 108 genes differed between 25wpi PD1hiTIM3– and 25wpi PD1hiTIM3+ CD4 T cells (Fig. 3e, Supplementary Data 7). Like the 4 vs. 25 wpi comparison, Group E CD4 T cells not only expressed higher levels of genes associated with exhaustion (Havcr2, Entpd1, Lag3, Il10) but also soluble mediators (Ifng, Ccl3, Ccl4, Ccl5) and genes associated with a cytolytic phenotype (Gzmb, Nkg7, Prf1). In contrast, Group D CD4 T cells expressed higher levels of Slamf6 and Tcf7, indicating a similarity to CD8 TPEX cells. We conclude that in the C7→TCRα model, C7 CD4 T cells acquire a transcriptional signature during the chronic phase of infection that resembles exhausted CD8 T cells.
Senescence signature among CD4 T cells during chronic infection
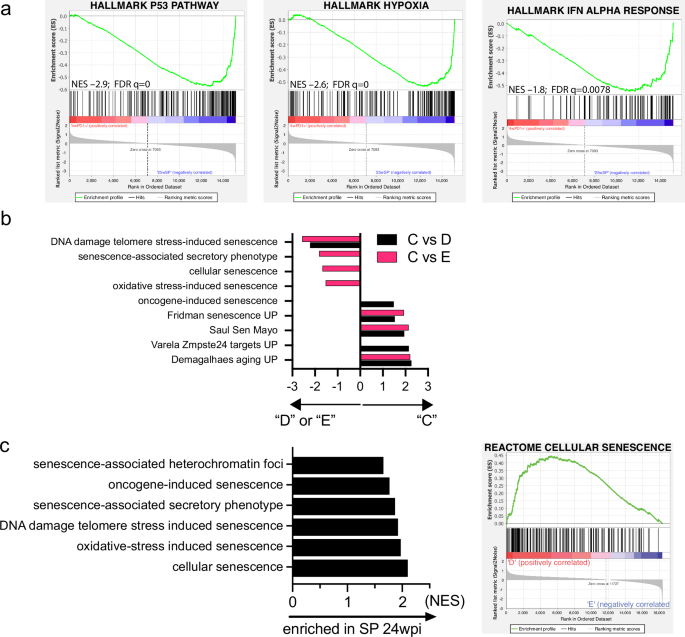
Three Hallmark pathways that were significantly enriched during chronic infection (i.e., expressed in Group D and E, compared to Group C C7 CD4 T cells) were the “p53 pathway,” “Hypoxia,” and “Interferon alpha response” gene sets (Figs. 3d, 4a). p53 is the best studied human tumor suppressor gene and is the most frequently mutated gene among human cancers. However, physiologically, the p53 pathway orchestrates key responses, often in response to DNA damage, oxidative damage, or metabolic stress, and generates a variety of cellular responses including DNA repair, cell cycle arrest, apoptosis, and senescence26. Activation of the p53 pathway has recently been linked to T cell senescence27,28. Both Type I IFN and hypoxia can also lead to cellular senescence (Fig. 4a)29,30,31.
a GSEA Enrichment plots between Group C and Group D T cells for the p53, Hypoxia, and IFN alpha Hallmark pathways. b GSEA results of senescence-related signatures enriched in Groups D or E compared to Group C. X axis, NES. c Senescence pathways enriched in Group D compared to Group E (left) and the Cellular Senescence GSEA enrichment plot (right). FDR, < 0.05 for all pathways shown. Source data are provided as a Source Data file.
We hypothesized that transcriptional pathways in response to cell stressors (UV damage, hypoxia, apoptosis, UPR, ROS pathway) could be mechanistically linked by activating the p53 pathway. Therefore, we considered whether the Hallmark pathways of “protein secretion,” “Mtorc1” and “PI3K/AKT3/mTOR” signaling, cytokine signaling (IFNγ, IFNα, TNF, TGFβ, IL-6) during chronic infection could indicate CD4 T cell senescence (Fig. 3d). Analysis using a compendium of “Senescence” signatures (Supplementary Data 8) revealed significant enrichment of the ‘Saul Sen Mayo’ and ‘Fridman Senescence Up’, and other gene sets32, which indicate that a program of cell senescence is active late during infection (Fig. 4b). Group D and E cells differed with respect to transcriptional programs associated with senescence. Several senescent gene sets were significantly enriched in Group D (PD1hiTIM3–) but not Group E (PD1hiTIM3+) C7 CD4 T cells (Fig. 4c). Also, PD-1 and TIGIT are associated with senescence and aging33,34. The genes encoding p53 and p16 were frequently shared in a leading-edge analysis. Thus, while chronic infection leads to changes consistent with senescence and exhaustion in C7 CD4 T cells, PD1hiTIM3– C7 cells (Group D) are more typical of senescent T cells while PD1hiTIM3+ C7 cells (Group D) represent exhausted T cells.
Few lung-resident CD4 T cells express multiple effector function during chronic infection
Given the surprising heterogeneity of CD4 T cell responses, especially regarding the detection of both effector and dysfunctional features, we next assessed whether the apparent heterogeneity could be resolved using scRNAseq to analyze polyclonal CD4 T cell responses. We investigated whether CD4 T cells become dysfunctional during chronic Mtb infection by comparing CD4 T cells from the lungs of B6 mice early (4 – 6 wpi) vs. late (39 – 50 wpi) after Mtb infection. CD4+CD44– T cells expressed little PD-1 or TIM-3, consistent with these cells being naïve (Fig. 5a). Early after infection, most CD44+E6tet– CD4 T cells (antigen-experienced excluding ESAT-6-specific) expressed PD-1 and a smaller population also expressed TIM-3. More ESAT-6 specific CD4 T cells (i.e., CD44+E6tet+) expressed both PD-1 and TIM-3. In contrast, nearly all CD4 T cells expressed PD-1 during chronic infection. Importantly, the ESAT6-specific CD4 T cells were predominantly PD-1+TIM-3+ late during chronic infection (Fig. 5a). Thus, CD4 T cells in chronically infected mice express both PD-1 and TIM-3, a phenotype associated with CD8 TTEX cells11,13.
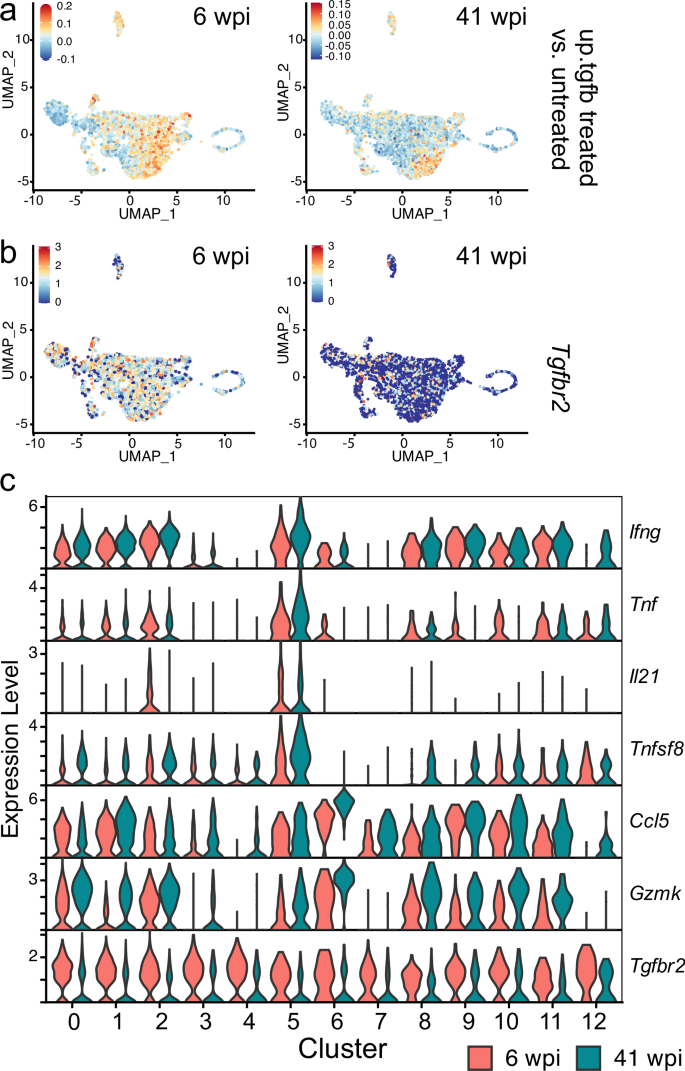
a–d Data is from 24 mice in 3 experiments infected for 4–6 (early) or 39–50 (late) weeks. Box plots indicate median (middle line), 25th, 75th percentile (box) and minimum and maximum (whiskers). E6tet+, Esat-6 tetramer+. a Percentage of total CD4 T cells (left) or E6tet+ CD4 T cells (right) that express both PD-1 and TIM-3 during chronic infection. b Percentage of EdU+ CD4+CD44+ T cells early (4–6 wpi) or late (39–50 wpi) after infection. c Percentage of CD4+CD44+ T cells producing IL-2, IFNγ, or TNF, after stimulation with ESAT61-15 (E6-stim). d Percentage of ESAT6-tetramer+ (E6tet+) CD4+CD44+ T cells among total CD4 T cells, early or late after infection. a, b, d two tailed t-test; (c) two-way ANOVA with Bonferroni’s multiple comparisons test. e–i Total lung parenchymal T cells were sorted and analyzed by scRNAseq. e UMAP projection of CD4 T cells clustered based on scRNAseq analysis and cluster designations. Outlined clusters all have a Th1 core signature. f Distribution of CD4 T cells among the different clusters, normalized by the total number of CD4 T cells from each subject (n = 2). Data are presented as mean values ± SD. Statistical testing was performed using a two-way ANOVA with Šídák’s multiple comparisons test. g Dot plot of differentially expressed genes. h Dot plot of genes associated with effector functions or co-inhibitory receptors. i UMAP projections overlapped with expression of Ifng and Il21. p-values: *, p < 0.05; ****, p < 0.0001; ns, no significant difference. Source data are provided as a Source Data file.
We next assessed the function of CD4 T cells during chronic infection. Significantly fewer cells proliferated compared to early timepoints (Fig. 5b). Late during infection, fewer CD4 T cells produced IFNγ and TNF after ESAT6 stimulation, and polyfunctionality was lost over time (Fig. 5c). Importantly, the frequency of ESAT-6 specific CD4 T cells was similar at both timepoints post-infection; thus, differences were not due to altered numbers of ESAT6-specific CD4 T cells (Fig. 5d). These data show that activated CD4 T cells develop a dysfunctional T cell phenotype during chronic infection.
To characterize CD4 T cell heterogeneity during chronic infection, CD3+TCRβ+IV−ve parenchymal lung T cells were flow sorted from Mtb-infected mice 6 (n = 2) and 41 (n = 2) wpi for analysis by scRNAseq and TCRseq (Supplementary Fig. 2a). After sequencing and quality control, 2838 (6 wpi) and 4723 (41 wpi) CD4 T cells were available for analysis. Cell clustering based on gene expression (excluding TCR genes) identified 13 clusters (Fig. 5e). Only the proportion of Cluster 0 and Cluster 4 differed between 6 and 41 wpi (Fig. 5f, Supplementary Fig. 2b). The CD4 clusters were annotated using transcriptional signatures and key canonical genes (Fig. 5g, h; Supplementary Fig. 2c, Supplementary Table 1, Supplementary Data 9). Clusters 7–12 accounted for 13% of the CD4 cells. Cluster 7 were Tregs (Foxp3, Ctla4, Il2ra, Ikzf2). Cluster 8 and 11 were proliferating cells (Mki67 and Top2a in G2/M or G1/S cell cycle phase, respectively). Cluster 9 expressed high levels of Klrd1, Klrg1, Tbx21 (T-bet) and Cx3cr1, indicating that these were terminally differentiated CD4 T cells. Cluster 10 expressed a type I IFN signature (Isg15, Bst2, Ifit1, Ifit2 and Ifit3). Cluster 12 were Th17 cells (Ccr4, Ccr6, 1l17re, Rorc, Aqp3).
Six clusters, including the four largest clusters (0, 1, 2, 3), all expressed Cxcr3, Cxcr6, Id2, Nkg7, Ccl5 and Ifng, suggesting that they were different forms of Th1 cells (Fig. 5e). We used FindMarkers to compare Clusters 0, 1, 2, 3, 5 and 6, to naïve-like CD4 T cells (Cluster 4; Ccr7, Sell, Lef1) (Supplementary Data 9). Cluster 6 cells expressed high levels of Ccl3, Ccl4, Gzmb and Eomes; some cells expressed Pfn1. As Eomes promotes a cytotoxic program in human CD4 T cells, Cluster 6 could be cytotoxic CD4 T cells35,36. Clusters 0, 1, and 3 were difficult to annotate. Cluster 0 resembled Th1 cells (Cxcr3, Cxcr6, Id2, Nkg7, Ccl5, Ifng) and Cluster 1 expressed genes indicative of tissue residency (Itga1, Jaml, CD226)37,38. Finally, upregulated genes in Cluster 3 included Tcf7, Il7r, Il2rb, Ly6a, Cxcr3, S1pr1, and Klf2 suggesting these are Tscm39. Ifng transcript was detected in Clusters 0, 1, 2, 3, 5 and 6, but Clusters 2 and 5 expressed considerably more Ifng than the other clusters. Cluster 5 cells expressed multiple effector genes required for protection including Tnf, Il21, Lta, and Tnfsf8 (Fig. 5h; Supplementary Data 9)19,40,41. CD4 T cells belonging to Cluster 5 are the key polyfunctional effector CD4 T cells in the lungs of infected mice. Cluster 2, which is adjacent to Cluster 5, also expressed Ifng, Tnf, Il21, and Tnfsf8, but lower amounts (Fig. 5h). In chronically infected B6 mice, many parenchymal CD4 T cells had a PD-1+TIM-3+TCF7– phenotype based on flow cytometry, which is typical of CD8 TTEX cells11,13. However, this cell surface phenotype poorly correlated with the transcriptional signature apart from Cluster 2, which we believe are exhausted. Notably, when the DEGs from the C7 RNAseq were used as a gene signature to probe the B6 scRNAseq dataset, the genes upregulated in Group D and E (vs. Group C, Fig. 3e) were most highly expressed by Cluster 2 and 5. The DEGs upregulated by Group E (vs. Group D) were specifically expressed by Cluster 2, indicating that these polyclonal CD4 T cells resemble exhausted T cells (Supplementary Fig. 2d).
Finally, polyclonal CD4 T cells in chronically infected B6 mice also expressed p53, senescence, and hypoxia signatures, particularly within Clusters 0, 1, 2, and 5 (Supplementary Figs. 3a–d). Thus, it appears that Cluster 5, which is only 3-4% of the CD4 T cells, are the true polyfunctional effectors that express the highest levels of IFNγ, IL-21, and TNF (Fig. 5I). In contrast, the majority of other Th1 cells appear to be hypofunctional, especially with respect to cytokine production.
Early and late CD4 T cell responses express distinct gene signatures
We identified statistically significant DEGs that were upregulated at 6 or 41 wpi using Seurat FindMarkers (Supplementary Data 10). PANTHER was used to determine molecular pathways that were statistically over-represented in the input list compared with randomly selected genes42. Annotated pathways including “T cell activation (P00053),” “JAK/STAT signaling (P00038)”, “Hypoxia response via HIF activation (P00030),” and “Inflammation mediated by chemokine and cytokine signaling (P00031),” were predicted to be upregulated at 6 wpi. No pathways of immunological relevance were significantly upregulated at 41 wpi. Next, genes that were upregulated at 6 or 41 wpi in each cluster were used to query the Reactome database (Supplementary Data 11). We focused on Clusters 0, 1, 2, 3, and 5, since they were of greatest interest. These five clusters were enriched for TCR signaling and CD28 costimulation pathways (except Cluster 3) at 6 wpi (Supplementary Table 2). Clusters 0, 2, and 5 appeared most activated based the identification of multiple signaling pathways including those for interferon, IL-2, and IL-21 signaling (Supplementary Table 2, Supplementary Data 11). At 41 wpi, only Clusters 2 and 3 were enriched for TCR signaling and none of the clusters were enriched for cytokine signaling pathways.
Regulatory pathways including CTLA4 signaling and signaling by the IL-27 and TGFβ inhibitory cytokines were enriched at 6 wpi but not 41 wpi. TGFβ signaling was confirmed by applying a TGFβ signaling signature to the UMAP projection (Fig. 6a). Genes induced by TGFβ were upregulated in several clusters at 6 wpi, but by 41 wpi, the signature was prominently expressed only by CD4 T cells in Cluster 1. Consistent with these data, Tgfbr2 was more highly expressed at 6 wpi than 41 wpi (Fig. 6b). Interestingly, genes in the “Downregulation of TGFB receptor signaling (R-MMU-2173788),” pathway were also significantly overrepresented at 6 wpi, suggesting that TGFβ receptor signaling was being actively regulated early but not late during the immune response to Mtb. Pathways pertaining to Notch and Runx signaling, which can synergize with other transcription factors to promote T cell activation and Th1 differentiation, were enriched in several clusters at 6 wpi but not 41 wpi (Supplementary Data 11). Genes upregulated in several clusters at 6 wpi, and to a more limited degree at 41 wpi, were enriched in pathways pertaining to cell stress and injury including senescence, apoptosis and necrosis, and hypoxia (Supplementary Table 1, Supplementary Data 11). Differences in individual genes were detected between 6 and 41 wpi. Various combinations of Ifng, Ccl5, Tnfsf8, Gzmk, Nkg7, and Il7r were significantly increased in Clusters 0, 1, 2, and 5 at 41 wpi (Fig. 6c, Supplementary Data 10). We speculate that the increased expression of these genes at 41 wpi could reflect an alteration of the regulatory environment of the Mtb lesion, or a consequence of greater antigen load. Increased expression of these effectors could drive more inflammation or even be a sign of CD4 T cell dysfunction.
Large clonal expansions of activated CD4 T cells are maintained during chronic infection
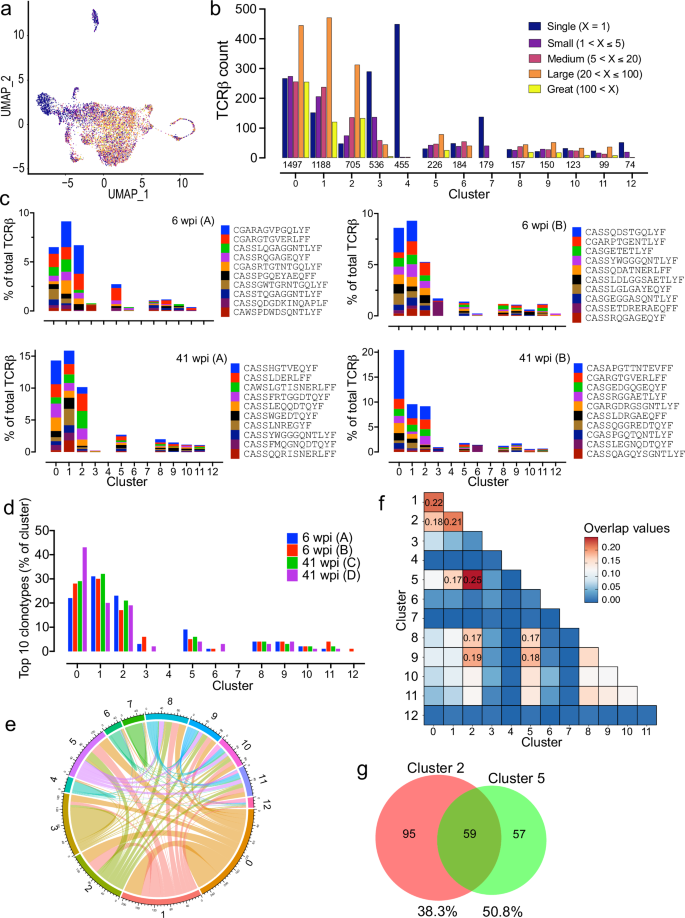
TCR analysis was used to identify CD4 T cells responding to Mtb since clonal expansion following activation is a cardinal feature of antigen-specific T cell responses. We identified 5573 paired TCRαβ; 2321 from 6 wpi and 3252 from 41 wpi. There were 1965 unique clonotypes based on amino acid sequences, which were categorized by the size of the expansion (Fig. 7a, b). The majority of TCRs in all clusters belonged to highly expanded clonotypes (i.e., > 5), except for Clusters 3, 4, 7, and 12. In Cluster 4, 99% of the TCR clonotypes were singletons, consistent with their annotation as naïve CD4 T cells. The 10 most abundant clonotypes accounted for 21% of CD4 T cells. We assessed the top 10 clonotypes for each subject (Fig. 7c). More than 75% of these highly expanded clonotypes had a Th1, Trm-like, Tfh-like, or Teff phenotype (i.e. clusters 0, 1, 2, 5, respectively) and were largely absent from the naïve, Treg and Th17 clusters. Overall, the Th1, Trm-like, and Tfh-like clusters were dominated by highly expanded clonotypes (i.e., containing > 20 cells), which accounted for 47-63% of the TCRs in these clusters. As a control for antigen-specificity, C7 CD4 T cells from uninfected mice were transferred into each B6 mouse one day before their infection. The C7 cells were identified based on their CDR3β sequence (Supplementary Data 12)43. Like endogenous CD4 T cells, most C7 cells were in the Th1, Trm-like, Tfh-like, or Teff clusters (Supplementary Data 12). These data show that while public TCRs for Mtb antigens exist and convergent recombination occurs, most clonal expansions were derived from a single CD4 T cell. Importantly, the detection of Mtb-specific CD4 T cells from the same clonotype among different transcriptional clusters implies that their differentiation and activation is influenced by extrinsic factors in microenvironments of the infected lung. These data show that Mtb infection is driving CD4 T cell responses in these clusters.
a A UMAP projection colored by the size of the TCR clonotype expansions. b Distribution of TCR count and clonotype size in each CD4 cluster. The number of total TCRs is listed above each cluster designation. c For each subject analyzed by TCRseq, the distribution of the 10 most abundant CDR3β among the 13 clusters is shown. d For each subject, the percentage of the total TCRs in each cluster accounted for by the top 10 TCR clonotypes. e A chord diagram of shared TCRs between clusters for Mouse 1 (46 wpi, A). f The overlap of TCR clonotypes between clusters based on the Jaccard Similarity Index. g Sharing of clonotypes between Clusters 2 and 5 determined by Venn analysis95.
Many of the highly expanded clonotypes were distributed among several clusters (Fig. 2e). To quantify the degree of TCR sharing, we calculated the Jaccard Similarity Index between each cluster pair. Significant sharing was detected between Clusters 0, 1, 2, and 5 (Fig. 7f). The greatest amount of TCR sharing was between Cluster 2 and 5, which were adjacent to each other on the UMAP projection and shared some transcriptional characteristics (Fig. 5). Sharing of unique clonotypes for Cluster 2 and Cluster 5 was 38.3% and 50.8%, respectively (Fig. 7g). Cluster 5 had the greatest expression of genes associated with CD4 effector function. Finding that Cluster 0, 1, and 2 CD4 T cells, while highly related by TCR clonotype and parenchymal location to Cluster 5 Teff cells, did not express any effector T cell transcriptional program, suggests these cells were hypofunctional or possibly paralyzed44.
TCR affinity affects the development of exhaustion
As TCR affinity and signaling strength affect T cell differentiation45,46,47,48, we hypothesized that TCR affinity would also influence the development of T cell dysfunction. C24 TCRtg mice express a TCR that also recognizes ESAT64-17 as do C7 cells, but with higher affinity49. C7 and C24 were transferred at a 1:1 ratio into TCRα KO mice and assessed at 4 and 34 wpi (Supplementary 1c). Early post-infection, no consistent phenotypic differences were observed between the two T cell clonotypes. At 34 wpi, the number of C7 and C24 cells was similar (Supplementary Fig. 1d). However, more C7 T cells resembled progenitor exhausted CD8+ T cells (i.e., TCF-1+TIM-3–), and more C24 T cells resembled terminally exhausted CD8+ T cells (i.e., PD-1+TIM-3+) (Fig. 8a). C24 cells expressed more T-bet (Fig. 8b), and more produced IFNγ (Fig. 8c), indicating a more terminally differentiated phenotype. There were no differences in IL-2 or TNF production (Fig. 8c).
a–c C7 (black) and C24 (green) TCRtg cells were co-transferred (1:1 ratio) into TCRα KO mice. Closed circles, females; open circles, males. Parenchymal C7 and C24 cells were analyzed at 4 (n = 8) and 34 (n = 6) wpi. a Percentage of C7 or C24 cells that expressed TCF-1+TIM-3– (left) or PD-1+TIM-3+ (right). b T-bet MFI. c Percentage of C7 or C24 cells that produced of IL-2, TNF, or IFNγ after ESAT6 peptide stimulation in vitro. Data are representative of two independent experiments. Statistical significance was analyzed by two-way ANOVA. d–g scRNAseq was performed on parenchymal CD4 T cells after co-transferred C7 and C24 cells, 24 wpi. d UMAP projection of C7 and C24 cells clustered based on scRNAseq analysis. e Location of C7 and C24 cells in the UMAP projection based on TCR sequence. f Proportions of C7 or C24 T cells within each cluster. g Feature plots of genes of referred to in the text. Source data are provided as a Source Data file.
We next performed scRNAseq of co-transferred of C7 and C24 cells at 24 wpi. We identified 1,181 C7 and C24 cells based on their CDR3β sequences, which were clustered into 9 populations (Fig. 8d, Supplementary Data 13). Clusters 2, 4, and 8 were predominantly C24 T cells; Clusters 0, 1, 3, 5, and 7 were predominantly C7 T cells; and Cluster 6 was mixed (Fig. 8e, f). Clusters 0 and 3 were Th17 cells (Rorc, Ccr6), although only Cluster 0 expressed Il17a (Fig. 8g). Cluster 1 had a memory phenotype (Il7r, Tcf7, Klf2); Cluster 5 had a cytotoxic signature (Gzmb, Prf1); Cluster 7 were Tregs (Foxp3); and Cluster 8 were proliferating cells (Mki67). Clusters 2, 4, and 6 contained Tbx21 expressing cells, but only Clusters 2 and 4 significantly expressed Ifng. Cluster 4 also expressed Il21. These results align with more IFNγ production by C24 cells. The reduced effector capacity of Cluster 2 in the context of low Il7r and high Hif1a suggests chronic stimulation50 or hypoxia51,52. These two clusters are reminiscent of Clusters 2 and 5 in B6 mice. In that analysis, Cluster 5 were highly functional effector CD4 T cells, and Cluster 2 consisted of CD4 T cells that had lost expression of some, but not all, effector genes (Fig. 5). The identity of Cluster 6, which mostly consisted of C7 T cells, was ambiguous. These cells expressed high levels of genes associated with CD8 T cell exhaustion (Nr4a1, Nr4a3, Egr2), suggesting Cluster 6 contained exhausted T cells53,54.
Our findings align with studies showing that high affinity TCR clones skew towards a Th1 phenotype45, establishing that TCR signaling strength contributes to the varied differentiation states of the T cells. The higher affinity C24 T cells express Ifng and Il21 and skew towards a Th1 phenotype. The lower affinity C7 T cells skew towards a Th17 phenotype and are more dysfunctional. However, another factor to consider is that higher affinity T cells can outcompete the lower affinity T cells for antigen, raising the possibility that antigen availability in this model can modulate T cell phenotype. Finding that the two clones clustered separately, independently of TCR gene expression, demonstrates that TCR affinity influences T cell differentiation and function during chronic Mtb infection.