The mNGS dataset in this study comprised 402 adult patients admitted to the First Affiliated Hospital, Zhejiang University School of Medicine (FAHZU) between 8 March 2020 and 27 May 2023. These patients were suspected of having lung cancer or pulmonary infections. Inclusion criteria required patients to be aged ≥18 years and to have undergone BALF sampling within 72 hours of intubation to identify causative pathogens. Exclusion criteria included underlying leukaemia, absence of a definitive diagnosis after extensive follow-up, or lack of matching DNA and RNA mNGS data from BALF samples. The diagnosis of lung cancer was based on clinical suspicion, supported by laboratory results from cytology, flow cytometry, and/or tissue biopsy. Pathological information for all samples was assessed according to the 2015 WHO Histological Classification of Lung Cancer and determined from surgically resected tissue sections. The diagnosis of pulmonary infections was based on clinical suspicion and confirmed through standard microbiological diagnostics, including cultures, antigen/antibody tests, PCR, and sequencing. This study retrospectively analyzed archival materials at FAHZU under a no-patient-contact research protocol, which was approved by the FAHZU Institutional Review Board (IIT20220714A). Prior to sample collection, written informed consent had been obtained from patients, covering the use of residual samples for research purposes. According to Guidance of the Ministry of Science and Technology (MOST) for the Review and Approval of Human Genetic Resources, we could not share sequencing data with homo sapiens.
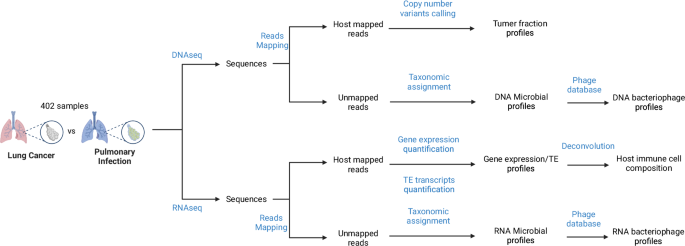
Here, we present a more condensed version of the methods fully described in Chen, Y. et al.12. The workflow is shown in Fig. 1. We make the raw sequencing data (unhost) freely available in NCBI Sequence Read Archive under the BioProject ID: PRJNA105676511, and scripts together with more downstream analysis results are accessible as the GitHub13.
The procedure of collecting BALF samples
BALF was performed using flexible bronchoscopy under local anaesthesia. The bronchoscope was advanced to a radiologically involved lung segment, and 100–150 mL of sterile 0.9% saline was instilled in 20–50 mL aliquots. After each instillation, fluid was gently aspirated and pooled. BALF specimens were immediately stored on ice and processed within 2 hours. Samples with visible blood contamination or recovery <30% were excluded.
BALF DNA/RNA sequencing methods
Wet lab BALF sequencing methods were described in previous study12,14. In brief, we recruited 123 lung cancer cases, 279 cases of pulmonary infections including tuberculosis, fungal, and bacterial infections, and 32 negative control cases that include conditions like immune pneumonitis, organizing pneumonia, and drug-related pneumonia. For BALF DNA sequencing, we treated 1 mL of BALF samples with 1 U benzonase and 0.5% Tween 20, incubating at 37 °C for 5 minutes to deplete host nucleic acids. Subsequently, 600 µL of this mixture was subjected to bead beating with ceramic beads in a Minilys Personal TGrinder H24 Homogenizer, followed by nucleic acid extraction from 400 µL of the sample using a QIAamp UCP Pathogen Mini Kit, with the final DNA elution in 60 µL. DNA quantity was assessed using a Qubit dsDNA HS Assay Kit. For BALF RNA sequencing, 1 mL BALF samples were centrifuged, and the precipitate was processed with TRIzol LS for RNA extraction using a Direct-zol RNA Miniprep kit. Library preparations for sequencing involved using 30 µL of DNA with the Nextera DNA Flex kit and 10 µL of purified RNA with the Ovation Trio RNA-Seq Library Preparation Kit. Library concentrations were quantified using a Qubit dsDNA HS Assay Kit, quality assessed via an Agilent 2100 Bioanalyzer with a High Sensitivity DNA kit, and sequencing performed on an Illumina NextSeq. 550 sequencer employing a 50-cycle single-end strategy2,3,4,15.
Generating microbial and host expression matrix
Detailed pipeline information for microbial and gene expression profiling is available at GitHub13. All parameters and database we used were showed with shell scripts. In short, we utilized a validated mNGS protocol aimed at comprehensive microbial composition analysis15,16,17. The process initiated with the use of fastp18 for the elimination of low-quality reads, duplicates, sequences shorter than 50 base pairs, and adapter contamination. To remove human genetic sequences, reads were aligned against the hg38 human reference genome using BWA (0.7.17)19. The generation of taxonomic profiles was facilitated by Kraken2 v2.0.7 and Bracken v2.5, which operated under default settings and employed a widely recognized database. To account for differences in sequencing depth, the sequencing reads identifying microbes were normalized to reads per million (RPM)20. Host gene expression was analyzed by aligning high-quality data to the human genome via HISAT221, using default settings, with gene-level quantification conducted through featureCounts. The aggregate gene counts were compiled using the featureCounts utility from the Subread package release 2.0.022.
In our GitHub repository and figshare (https://doi.org/10.6084/m9.figshare.29388539.v1)13, the kraken2_pipeline folder contains the following scripts for generating the microbial abundance matrix:
-
01_data_preprocessing.sh: uses fastp to remove low-quality reads and adapter sequences.
-
02_rmhost.sh: uses BWA and samtools to remove host reads by aligning to the human reference genome (hg38).
-
03_kraken2_bracken.sh: uses Kraken2 and Bracken to perform microbial taxonomic profiling.
-
04_relative_abudances_matrix.sh: calculates the relative abundance of each microbial taxon across all samples and summarizes the results at both the species and genus levels.
The RNAseq_pipeline folder includes scripts for read mapping, gene expression quantification, and profile generation:
-
01_data_preprocessing.sh: performs the same preprocessing as in the kraken2_pipeline using fastp.
-
02_map_to_reference_hg38.sh: uses HISAT2 to align reads to the hg38 reference genome.
-
03_featureCount.sh: uses featureCounts to generate gene-level read count matrices for individual samples. A specific version of the gene annotation file (GFF format) is provided to ensure result reproducibility.
-
04_readcount_to_expression_profiles.sh: merges the read count tables from all samples into a single gene expression matrix.
Microbial de-contamination
As previously reported in our methodology15,23, all wet-lab experiments were conducted under strict sterile conditions. Negative controls (PBS or sterile water) were included during nucleic acid extraction and library preparation to monitor potential contamination. These controls were processed in parallel with clinical samples throughout the experimental workflow. Our bioinformatic pipeline incorporated multiple layers of contamination control: ①Host DNA removal: Sequences were aligned to the human reference genome (GRCh38) and filtered. ②Decontam package application: We utilized the prevalence-based mode of the Decontam R package (v1.12.0), which statistically identifies contaminant taxa by comparing sequence frequencies between samples and negative controls. After generating microbial matrix, we used negative control to filter microorganisms which might be highly potentials as contamination. However, this step was quite independent for different laboratories. Thus, we provided our method as an optional procedure. First, negative controls were derived from BALF mNGS datasets of several (in our studies, there were 32, microbiological results and diagnosis of 32 negative control patients were listed in Table S3) individuals without infection or cancer, against which microbial abundance was compared. We then determined the mean and standard deviation of species’ relative abundances within these controls, establishing a threshold for positive detection at the mean plus three standard deviations. Second, microbes exceeding this threshold in the mNGS datasets of patients with lung cancer or infections were identified as ‘positive’ and included in our following microbial count analysis15.
Other omics information
We conducted secondary bioinformatic analysis employing various software tools. Firstly, we estimated the abundances of Transposable Elements (TE) using TEtranscripts24 and performed differential expression analysis with default parameters. Secondly, we identified immune-related genes (IRGs) using data from the ImmPort database (https://www.immport.org/home), and interferon-stimulated genes (ISGs) sourced from a referenced study25. Thirdly, to estimate the relative proportions of immune cells, we quantified transcript levels in TPM and employed digital cytometry via CIBERSORTx with the original gene signature file LM22 and 1000 permutations26. Fourthly, we identified tumor fractions or copy number variants through ichorCNA27, CNVkit28, and estimate29, adhering to the software instructions. Lastly, for bacteriophage annotation, we aligned cleaned reads against a curated phage database (CPD) using blastn30. Detailed parameters for each software or pipeline are outlined in our preprint manuscript12.
Differentially expressed genes (DEGs) and TE were identified in each group using the DESeq. 2 package, applying criteria of FDR ≤0.05 and Fold-change ≥1.531. Gene set enrichment analysis (GSEA) for DEGs was carried out using the REACTOME, KEGG, and GO databases by the fgsea package32,33,34. Significantly enriched pathways or biological processes were determined based on Fisher’s exact test (p-value < 0.05), following Benjamini-Hochberg adjustment. Latent variables were calculated by PLIER R package35. Wilcoxon rank-sum test assessed the difference between each group’s probability value.