Scarce PGx information in Africa
As a ground truth dataset, we curated PharmGKB7, the largest publicly available database of pharmacogenetic drug-gene interactions, and obtained a list of 1111 drugs for which there is, at least, one PGx association reported. A PGx association is characterised by a chemical-gene-variant triplet that has been reported in the scientific literature or is part of a recommendation report, for example, from the CPIC consortium. Importantly, one pair of chemical-gene can have multiple annotations if several variants from the same gene have been reported to interact with the given chemical. In addition, the association can influence the pharmacokinetics (PK) or pharmacodynamics (PD) of the drug, including efficacy, dosage and toxicity considerations. To provide an overview of the data available in PharmGKB, at this stage of the analysis we considered both PK/PD effects. Of the annotated drugs, we found that only 14.2% of them are indicated for communicable diseases (Fig. 1A). Moreover, infectious disease-related drugs have on average, less annotations per drug when compared to non-communicable diseases (Fig. 1B). These data highlight the need to increase PGx studies in infectious diseases.
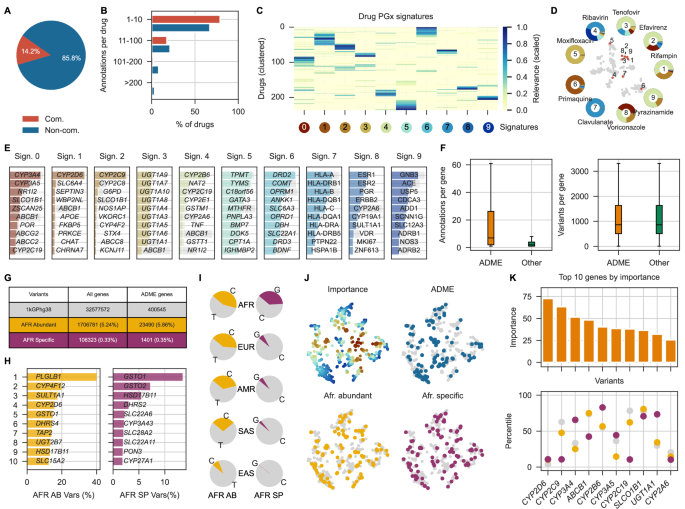
A Percentage of drugs for communicable vs non-communicable diseases with at least one significant pharmacogenetic association in PharmGKB. B Distribution of communicable vs non-communicable drugs according to their number of PharmGKB annotations. C Clustering of drugs according to their PGx signature. D T-SNE clustering of drugs according to their PGx signature. Examples of infectious disease drugs belonging to different signature groups are highlighted. E Number of PharmGKB annotations and number of genomic variants in ADME vs Non-ADME genes. The boxes include the interquartile range (with the inner line denoting the median), and the whiskers encompass the 5 and 95th percentile. F Top ten genes in each of the PGx gene signatures. G Percentage of African Abundant and African Specific variants in the 1kGPhg38 whole genome annotation and specifically in ADME genes. H Top ten ADME genes by number of African Abundant and African Specific variants. I Example of an African abundant variant distribution across populations corresponding to rs7255816 (gene: CYP4F12) and example of an African specific distribution across populations corresponding to rs680055 (gene: CYP3A43) (AFR: Africa, EUR: Europe, AMR: America, SAS: South-Asia, EAS: East-Asia). J T-SNE of PharmGKB-annotated genes represented by signature association. Overall importance in signature analysis is shown in the top left, ADME genes are highlighted in the top right, and top 10% of genes according to African abundant and African specific variants are shown in the bottom. K Top ten genes selected by their importance to PGx signature and representation of the African Abundant (yellow) and African Specific (purple) variants they harbour. Source data for B,H,K are provided as a Source Data file.
To gain a global understanding of the PharmGKB data, we identified sets of pharmacogenes that tend to co-occur across drugs. In Fig. 1C, at a high level, it can be seen how drugs can be partitioned into 10 pharmacogene “signatures” (Methods), with some drugs being very specific to one signature (e.g., clavulanate and moxifloxacin) and others showing a more varied profile (e.g. ribavirin and voriconazole) (Fig. 1D). Interestingly, seemingly unrelated drugs such as tenofovir, rifampicin and pyrazinamide, shared association with the same signature (Fig. 1D) defined by pharmacogenes such as CYP2B6 and NAT2 (Fig. 1E). Of note, some signatures were specific to one or a few genes (e.g., signature 0: CYP3A4 and CYP3A5, which co-occur as expected; signature 1: CYP2D6, signature 2: CYP2C9), while others encompassed gene families (signature 3: UGTs, signature 7: HLAs).
In addition, we were interested in understanding the distribution of PharmGKB annotations by gene, in particular comparing Absorption, distribution, metabolism and excretion (ADME) vs non-ADME genes. We used a list of 283 genes curated by the PharmADME working group, defined by their involvement in the ADME processes of drugs8. As such, ADME genes are expected to be more frequently associated with PGx effects. Indeed, we observed that in our curated data ADME genes harbour a much higher proportion of significant annotations in PharmGKB when compared to non-ADME genes (Fig. 1F).
In parallel to the curation of PharmGKB data, we also focused on the identification of genes that will be more relevant to downstream applications of our study because they carry variants that are prevalent in Africa. To that end, we used the 1000 Genomes Project data9, which contains whole genome sequencing (WGS) and deep whole exome sequencing (WES) of 2504 individuals from 26 different populations. The populations are grouped into the following categories: Africa (AFR), Europe (EUR), America (AMR), South Asia (SAS), East Asia (EAS). First, we analysed the distribution of variants between ADME and non-ADME genes, demonstrating that, despite ADME gene variants being annotated much more frequently in PharmGKB, both ADME and non-ADME genes harbour a similar number of genetic variants (Fig. 1E). Next, we set out to classify the variants as non-African (i.e. variants that are present in less than 20% of the African samples), Africa-abundant (i.e., variants that are present at a higher proportion in African populations, but are also common to others; AFR-abundant) or Africa-specific (i.e., variants that, on top of being abundant, are present at a much higher proportion with respect to any other ancestry; AFR-specific). Briefly, an AFR-abundant variant was defined as a variant having an AF over 0.20 in African populations regardless of its AF in the remaining populations, and an AFR-specific variant was defined as a variant within the set of AFR-abundant variants overrepresented at least 8x with regards to the remaining populations (Methods, Fig. S1). Overall, ADME genes maintained the proportion of AFR-abundant and AFR-specific variants in ADME vs non-ADME genes (Fig. 1G). Among the genes harbouring more AFR-abundant and AFR-specific variants, we find members of the CYP450 family and numerous transporters (Fig. 1H). An example of the AF distribution across populations of an AFR-abundant and AFR-specific variant is showcased in Fig. 1I. The distribution of AFR-abundant and AFR-specific variants annotated in PharmGKB is maintained between ADME and non-ADME genes (Fig. S2).
Finally, we plotted the PharmGKB-annotated genes according to their signature association (Fig. 1J). The top ten genes important to the PGx signatures are relevant ADME genes, including several CYP450 family members and transporters (Fig. 1K). Interestingly, genes encoding for the CYP3A4, CYP2B6 and UGT1A1 enzymes harbour a relatively high proportion of AFR-specific variants.
ML/AI-based pharmacogene prioritisation
Next, we set out to develop an ML/AI method to predict drug-pharmacogene pairs, using PharmGKB as a reference dataset. Given the scarce PGx information available on the African population, we devised a strategy that leverages orthogonal information available for drugs and genes to assist in the training procedure. Previously, in tasks, such as drug activity prediction, we showed that incorporating publicly available data from chemoinformatics and bioinformatics resources can improve the predictive power of ML models, especially when few training data points are available10. Here, we primarily used a biomedical knowledge graph that collects drug and gene information of multiple types, including drug-gene interactions, gene expression profiles in cells, drug side effects, protein function and cellular localisation, etc. From this knowledge graph, we selected types of biomedical relationships that we reasoned made sense in the context of PGx, for example, protein abundances in tissues, protein-protein interactions, participation of proteins in pathways, and drug targets and metabolic genes (Tables S1 and S2). This information was retrieved in ‘embedded’ form from the Bioteque11, i.e., in 128-dimensional vectors assigned to each drug and each gene in our dataset. In brief, for each type of relationship and data source (e.g., protein-protein interactions from the STRING database12) in the knowledge graph, we obtained an embedding vector for each entity (gene, in this case), such that similar vectors correspond to proximal nodes in the original network. In addition to knowledge graph embedding vectors, we also obtained sequence embeddings for genes13 (i.e., learned numerical representations of protein sequences such that similar embeddings correspond to similar sequences), as well as multiple descriptors for the drugs. As drug descriptors, we used a representative repertoire of tools, including 2D chemical fingerprints, arrays of physicochemical properties14, bioactivity profiles, and ADME calculations15 (Table S3). Collectively, we assembled, for each drug and gene in our dataset, a compendium of numerical arrays that captured the knowledge available for them in a vectorial form that is amenable to downstream ML tasks (Fig. 2A).
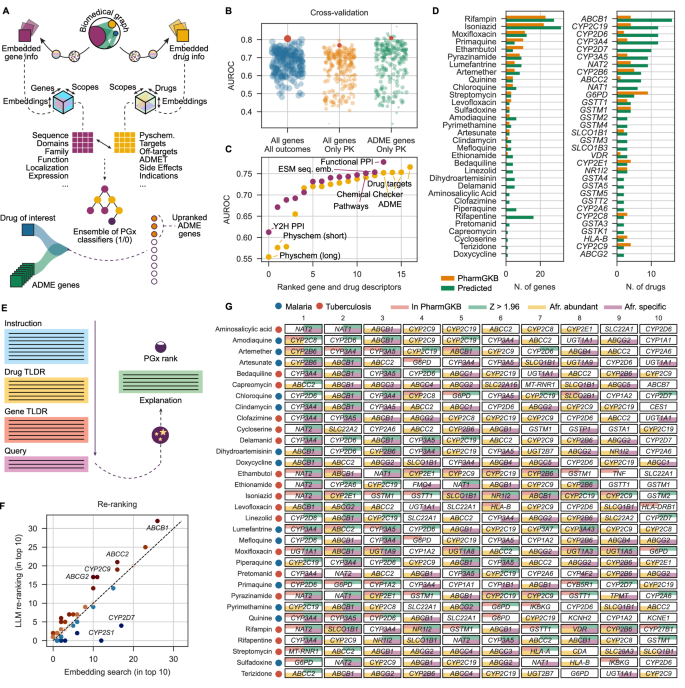
A Graphical representation of the pipeline to extract gene and drug embeddings for the training of the PGx classifier. B Cross-validation performance measured by the Area Under the Curve (AUROC) of the different PGx classifiers and the performance of each of the three ensembles (red dot). C Top quartile performance (AUROC) of classifiers using different individual drug and gene descriptors. D Number of genes associated with each of the selected drugs for the study according to PharmGKB vs predicted by the PGx classifier, and number of selected drugs associated with the top genes according to the PGx classifier vs the number of annotations in PharmGKB. Source data are provided as a Source Data file. E Graphical representation of the LLM interrogation for the selection of the top ten PGx associations per drug. F Genes in the top ten most important PGx associations to the drugs of study according to the LLM vs the initial PGx classifier. G Top ten genes with predicted PGx association per drug (z-score >1.96, green) and annotated by their association in PharmGKB (orange). In yellow, we show genes that have at least 50 AFR-abundant variants and, in purple, genes that have at least one AFR-specific variant.
While the collected orthogonal information above is not explicitly related to PGx, it constitutes a good starting point for a supervised ML approach whereby a drug-gene pair predictor is trained based on their known drug-gene associations in PharmGKB (outcome variable; y) and the drug/gene embeddings or vectors (features; X). More precisely, we formulated the problem as three binary classification tasks, corresponding to three training sets of decreasing size, namely all drug-gene pairs in PharmGKB with any type of PGx association, all drug-gene pairs with a PK association, and drug-ADME gene pairs with a PK association (Fig. 2B). For each binary classification task, we defined as positive (1) the known drug-pharmacogene associations, and we randomly sampled a set of 10x unlabelled drug-pharmacogene pairs to be used as negatives (0), preserving the relative frequency of drugs and genes observed in PharmGKB. We then trained an ensemble of models for each task in a fully automated manner, each member of the ensemble corresponding to a model trained with a certain type of drug and gene features (e.g., drug side effects and protein-protein interactions) (Methods). In a cross-validation procedure, we found that aggregating results across the ensemble consistently yielded high performance (Fig. 2B), with AUROC scores around 0.8 for all three binary classification tasks. Of note, we found that biologically-informative representation for drugs such as drug target profiles, Chemical Chequer bioactivity signatures16,17, and predicted ADME properties (including, among others, interactions with CYPs), were most powerful predictors, whereas purely physicochemical descriptors as commonly used in QSAR modelling yielded least performant models. Similarly, functional protein-protein interactions (PPIs), protein sequence embeddings (which are known to capture molecular function and structural features), and protein annotated pathways were most predictive, while physical (not necessarily functional) PPI interactions detected in vitro in a yeast-two hybrid (Y2H) assay were the least informative in this setting (Fig. 2C).
We then carried out an exhaustive prediction experiment evaluating all pairwise combinations of drugs and genes. To simplify the analysis, we empirically derived a consensus Z-score averaged across the three models. For a majority of the drugs of interest (i.e., malaria and TB drugs), we could find significant predictions, including drugs such as rifapentine for which no information was available in PharmGKB (Fig. 2D). Likewise, ADME genes encoding enzymes such as NAT1 or ABCC2, which had no or little annotation in PharmGKB, appeared to be associated with a relatively high number of malaria and TB drugs.
In modern AI search algorithms, embedding-based search is often used as an initial ranking procedure that is then refined with a large language model (LLM). We therefore submitted our ranked list of pharmacogenes predicted for each drug to additional scrutiny using GPT-4, which has demonstrated competitive performance in biomedical domain settings18. We took the top-50 genes (ranked by Z-score) predicted for each drug, and asked GPT-4 to select ten of them. This selection was done using controlled prompt-engineering and including prior knowledge about drugs and genes as context for the LLM (Fig. 2E and Table S4). This can be viewed as an LLM-driven “feature selection” task19. In Fig. 2F, it can be seen that, while there was a correspondence between the top ten genes selected in the embedding-based search and the final LLM, some genes were favoured by the LLM, including genes encoding transporter proteins such as ABCB1, ABCC2, ABCG2, and the CYP2C9 enzyme. On the other hand, CYP2S1 was deprioritised by the LLM, as well as the segregating pseudogene CYP2D7.
Overall, we obtained, for each of the 32 malaria and TB drugs of interest, a list of top ten ADME pharmacogenes predicted by our pipeline (Fig. 2F). Some of the predicted associations were already known PGx associations, such as the relationship between rifampicin and NAT2, or the association between sulfadoxine and G6PD. These are marked in red in Fig. 2F. However, most of the associations have not previously been reported. For example, bedaquiline, a relatively new (2012) approved drug to treat multidrug resistant TB, is not annotated in PharmGKB and we predicted it to be associated with CYP3A4 and CYP3A5 (which, reassuringly, tend to co-occur as in the literature20) as well as ABCB1, among other pharmacogenes. Indeed, ABCB1 was predicted to be associated with several drugs, which might indicate that this is a gene worth exploring for these disease areas generally. ABCB1 has also been associated with malaria severity, both through effects on drug response and through affecting the host’s interaction with P. falciparum21. Interestingly, ABCB1 harbours AFR-abundant and AFR-specific variants, as do other ABC transporters like ABCC3, ABCC4 and ABCG2, also found in our panel of predictions. To make the results of this ML/AI-based prediction available to the community, we have released a web application as specified in the Code Availability section. In this web application, we include known and predicted associations (e.g., NAT2-sulfadoxine OR CYP3A4-delamanid) along with LLM-generated summaries for the drugs, genes and interactions, with the hope that they can guide researchers into interpreting the predictions and critically assessing them.
Integrating predicted pharmacogenes into PBPK modelling
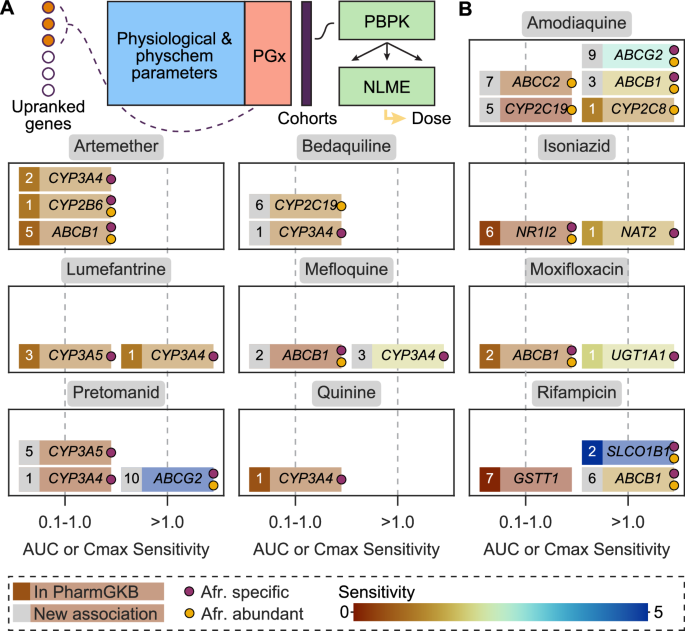
Having prioritised 10 pharmacogenes per drug, we then asked whether these predictions could be incorporated into PBPK modelling (Fig. 3A). To quantitatively assess this, we performed a comprehensive sensitivity analysis of ten of our drugs for which we could find evidence of variable PK and drug response in African cohorts (Table S5). In the sensitivity analysis, the AUC and Cmax PK parameters were evaluated for their fold-change if the predicted genes were incorporated into the model with respect to the absence of those genes (Methods). In Fig. 3B, it can be seen that, for all drugs, we found at least one sensitive pharmacogene, with those encoding for CYP3A4, CYP3A5 and ABCB1 being frequent across drugs. Importantly, of the 23 sensitive drug-pharmacogene pairs (Fig. 3B), 12 (52%) were not previously reported in PharmGKB, demonstrating the potential of our approach to quickly prioritise pharmacogenes, especially for drugs for which few or no PGx interactions are known. For amodiaquine, for example, only CYP2C8 of the five sensitive genes was reported in PharmGKB, while predicted genes such as ABCB1 and ABCG2 elicited even higher sensitivity scores. Moreover, when we randomly sampled ten genes from the ADME list, we could not identify any sensitive drug-gene pair, demonstrating the significance of our results. Of the identified PBPK-sensitive drug-pharmacogene associations, 61% involved genes harbouring at least 50 AFR-abundant variants, and 87% involved genes with at least one AFR-specific variant, suggesting that our findings are worth exploring more in depth in the context of African genetic studies. Notably, >80% of the variants in our dataset corresponded to intronic variants (annotated with SNPeff22, see Methods), which may be particularly relevant in the context of PBPK modelling, since intronic variants tend to be associated with gene regulation. This is coherent with the PBPK software PK-Sim® used for the analysis, which utilises gene expression data as a proxy for protein abundance to estimate the in vivo activity of enzymes and transporters that affect PK23.
A Graphical representation of the pipeline to include PGx information into PBPK and PK modelling. B Sensitivity analysis by PBPK of the top ten predicted PGx associations between selected drugs and ADME genes. Genes with a sensitivity between 0.1 and 1 inclusive are shown in the left column of each panel. Genes with sensitivity values greater than one are shown in the right column. Red-to-blue spectral scale is for the sensitivity values (low-to-high). The number indicates the rank in our predicted list. Numbers in a grey box indicate genes not reported in PharmGKB with any level of evidence for the drug. Numbers in a coloured box indicate that some degree of evidence is available in PharmGKB.
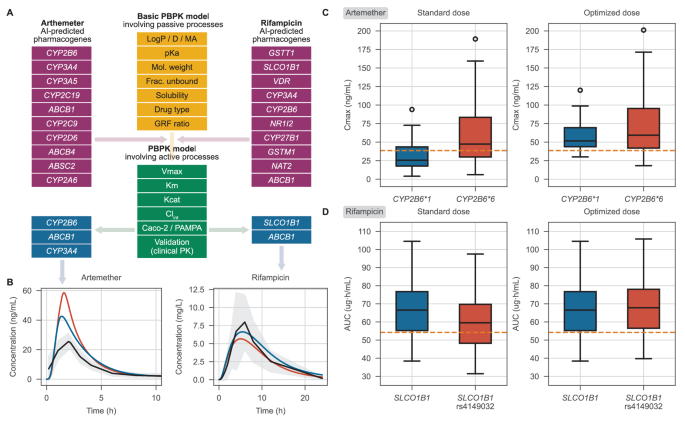
To illustrate how the pharmacometric analysis could proceed from this point, we chose the case of rifampicin and artemether (Fig. 4A). For artemether, the CYP3A4, CYP2B6 and ABCB1 pharmacogenes were sensitive. While association with CYP3A4 and CYP2B6 was previously reported in PharmGKB, for ABCB1 we found further support in the literature24 (there was only low-level evidence in PharmGKB via an automated annotation). The simulated concentration-time profile of artemether in the presence and absence of ABCB1 transporter protein is shown in Fig. 4B. ABCB1 was also found to be sensitive in the PBPK analysis of rifampicin, and has also recently been associated with rifampicin PK25. Another highly sensitive ( >1) gene, SLCO1B1 is known to play a role in rifampicin PK26 and was considered jointly with ABCB1 for further analysis. The full PBPK models files of artemether and rifampicin are included as supplementary Table S6.
A Building PBPK models for artemether and rifampicin involving integration of AI predicted pharmacogenes and model validation using clinical PK data. B Predicted concentration-time profiles involving active processes of sensitive pharmacogenes (blue and red lines) with a black line showing the mean observed averaged clinical PK data (grey shade shows standard deviation over three studies). C, D NLME modelling, assessment of standard dose of artemether and rifampicin for PK target attainment in African population harbouring enzyme/ transporter variants and Monte Carlo simulation of optimised doses. The boxplots in C, D show the median (line), interquartile range—IQR (box), and range within 1.5 × IQR (whiskers). Overlaid error bars denote the 95% confidence interval around the mean. Data for each boxplot was from 200 virtual patients. The horizontal dashed orange lines represent pharmacokinetic target value of 38.6 ng/mL for artemether and 54.2 µg*h/mL for rifampicin. Source data for C,D are provided as a Source Data file.
The PBPK modelling predicted secondary pharmacokinetic parameters (AUC, Cmax and Tmax) were comparable to those estimated from the observed clinical data as shown in Table S7. The bias was within the acceptable range, which was below ±15%.
NLME modelling and Monte Carlo simulation for dose optimisation
The final model for artemether was best described by a 2-compartment PK model with linear elimination which had consisted of CYP2B6 polymorphism as covariate. For rifampicin, the final model was best described by a 1-compartment model and linear elimination with SLCO1B1 polymorphism as covariate. The final models were used to perform Monte Carlo simulations (Table S8). Finally, we assessed the above-mentioned genes in the context of a dose optimisation analysis. The NLME modelling of artemether confirmed the CYP2B6*1 and CYP2B6*6 star alleles to be the significant covariates of artemether PK (Table S6). ABCB1 was not found to be a significant covariate of artemether PK. Virtual patients harbouring CYP2B6*1 and CYP2B6*6 variants were associated with ‘normal’ and ‘high’ plasma drug exposure, respectively. Monte Carlo simulation of artemether standard dose of 80 mg resulted in 41.8% of the population with CYP2B6*1 attaining the Cmax target of 38.6 ng/mL while in the population with CYP2B6*6, 78% attained the target Cmax. The optimum doses necessary for the patient populations with CYP2B6*1 and *6 variants to reach a probability of >91% target attainment were 152 and 85 mg, respectively (Fig. 4A). Similarly, the SLCO1B1 wild-type and SLCO1B1 rs4149032 transporter variants were confirmed as significant covariates of rifampicin pharmacokinetics but ABCB1 had an insignificant effect. The SLCO1B1 rs4149032 variant was associated with low rifampicin plasma exposure compared with the SLCO1B1 wildtype. Simulation of the rifampicin standard dose of 600 mg resulted in 43.5% of the virtual patient population with SLCO1B1 rs4149032 attaining the AUC target of 54.2 µg*h/mL while 91.3% of the virtual patient population with the SLCO1B1 variant attained the AUC target (Fig. 4). Monte Carlo simulations showed that the rifampicin dose of 750 mg was optimum in the population of patients with the SLCO1B1 rs4149032 transporter variant as it resulted in over 91% of the population attaining the target AUC. The proposed optimal doses are not for all African populations as the optimal dose will depend on the frequency of the alleles that are significant in the African subpopulation of interest.