Cohorts and features for training and testing
The cohort selection criteria for training the hospital model resulted in a total dataset size of 9,517 patient encounters with waveform recordings around the time of clinical suspicion of hospital-acquired infections (not including COVID-19). Of these patient encounters, 3,951 (3,665 controls and 286 HAIs; 51% Banner Health and 49% MIMIC-III) had overlapping PPG waveforms and impedance-based measurements with good data quality, and therefore had the full set of 18 demographics and vital sign features (see METHODS) available at 1-hour before clinical suspicion of infection. These 3,951 patient encounters were used to train the hospital model of hospital-acquired infection prediction.
The cohort selection criteria for testing the trained hospital model resulted in 301 COVID-19 positive subjects and 2,111 COVID-19 negative subjects from the wearable dataset. Within the 14-day windows prior to COVID-19 tests from these subjects, a total of 33,164 subject days and 31,269 subject sleep segments had vital sign measurements that passed our plausibility filter. From these subject days, we extracted the feature vectors comprising the same set of 18 features that was used to train the hospital model, using either all available vital sign measurements or those measured during sleep (“daily features” and “sleep-only features”, see METHODS), to quantify the performance of the trained hospital model in predicting COVID-19 infections.
Differences between training and testing datasets
The joint distribution of inputs and outputs of the infection prediction model differed between the training scenario in the hospital dataset and the testing scenario in the wearable dataset – a problem known as “dataset shift”16. Here we describe five sources of dataset shift in our study.
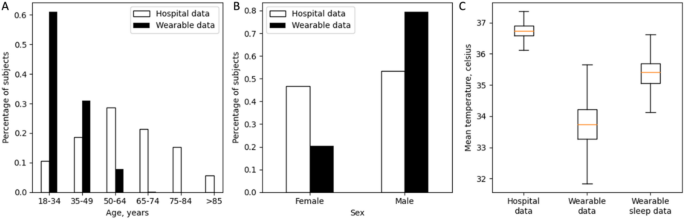
First, the demographics of the training and testing cohorts were different. The patients from the hospital dataset were older than the subjects from the wearable dataset (Fig. 2A), and the wearable dataset had an imbalanced sex ratio than the hospital dataset (Figs. 2B, 20% female in the wearable dataset versus 47% female in the hospital dataset). Both age and sex may result in differences in physiology24,25,26,27,28,29,30,31,32,33,34.
Second, the health states of the training and testing cohorts were different. Patients in the hospital dataset are those who developed hospital-acquired infections during their stays in general wards or in some cases intensive care units, and are likely older adults with comorbidities and under medical treatments, therefore the physiological measurements in the hospital dataset were more likely to be abnormal and unstable compared to the physiological measurements in the wearable dataset where healthy young military personnel performing their daily duty were monitored. We found that patients in the hospital dataset had higher heart rate and higher respiratory rate than the subjects in the wearable dataset (see the Average and Maximum statistic feature in Supplementary Table 1), which were consistent with an overall declined health state34,35,36,37. The hospital patients also had larger variations in heart rate and respiratory rate than the subjects in the wearable dataset (see the Standard Deviation statistic feature in Supplementary Table 1).
Third, the data sources where the physiological features were extracted from were different between the hospital dataset and the wearable dataset. Temperature features were extracted from core body temperatures in the hospital dataset, whereas in the wearable dataset skin temperatures measured at the fingers were used. We found that skin temperature had lower values and larger variance compared with core body temperature (Fig. 2C, Supplementary Table 1), which was consistent with the literatures38,39,40,41.
Fourth, the processing methods to extract physiological signals were different between the two datasets. Heart rate variability measurement RMSSD were computed based on pulse estimates of heart beats. However, the signal processing algorithms that Oura ring used could be different from ours in detecting the fiducial points on the pulse waveforms, and in the validation of the resulted inter-beat intervals. We suspected that differences in the signal processing algorithms to obtain RMSSD also contributed to the distribution differences in the RMSSD features between the hospital dataset and the wearable dataset (Supplementary Table 1), in addition to the demographics and health state differences mentioned above.
Finally, wearable physiological data is acquired in an unconstrained, real-world environment, which is influenced by everyday activities and other contextual factors. In contrast, hospital physiological data is typically acquired when the patient is sedentary. Daytime activity such as physical exercise increases heart rate and respiratory rate34,35 which is a confounding factor to infection prediction because infections cause similar changes in vital signs36,42. Skin temperature also changes dynamically upon physical exercise, and the directionality of change depends on the intensity level of the exercise and whether the skin temperature is measured over active or non-active muscles43. When limiting feature extraction to wearable physiology data acquired during sleep, we found that sleep-only features have different data distributions compared to the daily features (Supplementary Table 1). For example, the data distribution of the mean temperature feature was shifted towards higher values when restricted to measurements during sleep (Fig. 2C).
We included a full comparison of feature values in Supplementary Table 1.
Comparison between hospital dataset and wearable dataset. (A) Age distribution of hospital dataset (white) and wearable dataset (black). (B) Sex distribution of hospital dataset (white) and wearable dataset (black). (C) Boxplot of mean temperature feature value from hospital dataset (left), wearable dataset (middle), and wearable dataset during sleep (right).
Experiment design
We explored two approaches to correct for differences in data distributions between hospital and wearable datasets. First, we limited feature extraction to wearable physiological data from wearable sensors acquired when the subject was sleeping. This approach directly mitigated dataset shift by removing contextual confounders of daytime activities. Second, we explored a monotonic feature transformation method to convert the data distribution of physiological features in the wearable dataset to match the data distribution in the hospital dataset. This approach addressed covariate shift – one of the three types of dataset shift (see INTRODUCTION) – due to differences in demographics and health state between hospitalized patients and subjects in the wearable dataset, as well as differences in physiological measurements between COTS wearables and hospital grade devices. We compared model performances with or without using such correction techniques (Experiments I, II, III, IV in Fig. 3), and in addition benchmarked data requirements (Experiments V, VI, VII):
-
Experiment I: a baseline comparison where the trained hospital model was directly applied to the daily features from the wearable dataset. Physiological measurements during both awake and sleep were used to extract the daily features.
-
Experiment II: the trained hospital model was tested on sleep-only features from the wearable dataset. Sleep-only features were extracted from the same window and time interval as the daily features but only using measurements during sleep segments.
-
Experiment III: the trained hospital model was tested on sleep-only features after the sleep-only features were transformed to match the distribution of the hospital dataset.
-
Experiment IV: the trained hospital model was tested on daily features after the daily features were transformed to match the distribution of the hospital dataset.
-
Experiments V, VI, VII: benchmarking the amount and type of wearable data needed for the monotonic feature transformation.
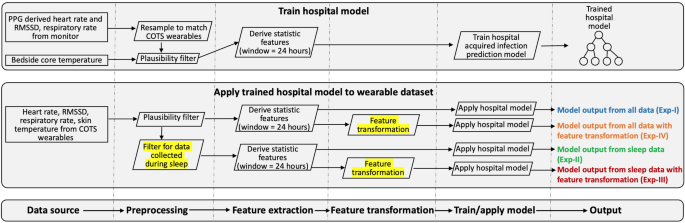
Schematic view of pipelines for training the hospital model (top box) and for testing the trained model in the wearable dataset (middle box). Similar steps of the two pipelines are aligned (bottom box). The trained hospital model was applied to the wearable dataset with or without the two dataset shift corrections (highlighted, middle box), which resulted in four experiments (Exp-I, II, III, IV in middle box) to compare model performance.
Baseline comparison (Experiment I)
We directly applied the hospital model trained for hospital-acquired infection prediction to the wearable daily features and quantified its performance in predicting COVID-19 infections. We hypothesized that the hospital model would not generalize well in predicting COVID-19 infections, due to the differences between hospital and wearable physiological feature spaces. We found that the hospital model performed at Area under ROC Curve (AUROC) = 0.527, Average Precision (AP) = 0.132, Sensitivity = 0.163 and Specificity = 0.866 at break-even point, Sensitivity = 0.193 and 0.113 respectively when Specificity was at 0.8 and 0.9. This performance was at chance level (95% confidence interval: [0.498, 0.566] for AUROC, [0.122, 0.153] for AP), suggesting that the hospital model failed to generalize when directly applied to wearable dataset.
Removing contextual confounders (Experiment II)
When controlling for contextual factors like daytime activity, we found that the hospital model using the sleep-only features performed at AUROC = 0.644 (95% confidence interval: [0.632, 0.701]), AP = 0.260 (95% confidence interval: [0.258, 0.351]), Sensitivity = 0.279 and Specificity = 0.897 at break-even point, Sensitivity = 0.402 and 0.269 respectively when Specificity was at 0.8 and 0.9. Thus, using sleep-only features resulted a 22% boosting of performance in terms of AUROC, suggesting the importance of controlling for contextual confounders when extracting the likelihood of infection from wearables physiological data.
Applying feature transformation after removing contextual confounders (Experiment III)
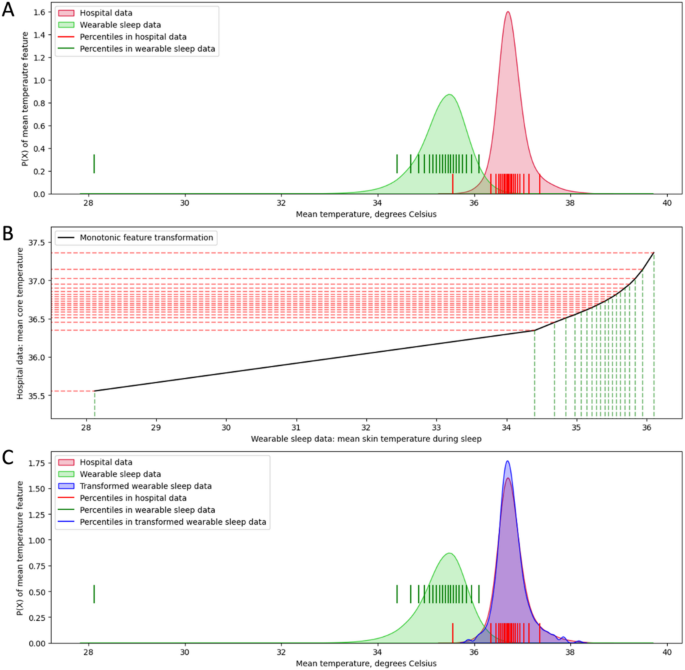
We hypothesized that a monotonic feature transformation procedure which transforms the wearable feature values to match the distribution in hospital dataset (see METHODS) could improve performance of the hospital model. Using mean temperature feature as an example (Fig. 4), the feature transformation procedure based on matching feature values that share the same 0-100 percentile value in their corresponding datasets resulted in an almost identical data distribution of the mean temperature feature between the two datasets, despite large discrepancies in the data distributions before transformation. Hence, we performed the same feature transformation procedure independently on each feature, and evaluated the performance of hospital model on the wearable dataset after all the features were transformed. We found that the hospital model performed at AUROC = 0.740 (95% confidence interval: [0.740, 0.808]; Fig. 5A, red solid line), AP = 0.330 (95% confidence interval: [0.330, 0.442]; Fig. 5B, red solid line), Sensitivity = 0.379 and Specificity = 0.910 at break-even point, Sensitivity = 0.588 and 0.409 respectively when Specificity was at 0.8 and 0.9, using transformed wearable sleep-only features. Applying feature transformation on the sleep-only features resulted an additional 15% boosting of performance in terms of AUROC (0.740 versus 0.643, red solid line versus green dashed line in Fig. 5A).
Monotonic feature transformation of mean temperature feature. Red, hospital dataset; green, wearable dataset (sleep-only features); blue, transformed wearable dataset (sleep-only features). (A) Data distribution of mean temperature feature: red and green shaded areas describe data distribution from hospital and wearable sleep data respectively. Vertical lines mark the 0-100 percentile values in 5% intervals on the x-axis corresponding to each dataset. (B) Monotonic feature transformation curve (black) where feature values with the same percentile value are mapped between two datasets. Dashed lines mark the 0-100 percentile values in 5% intervals on the x-axis for wearable sleep data (green) and on the y-axis for hospital data (red). (C) Data distribution of mean temperature feature: red, green and blue shaded areas describe data distribution from hospital dataset, wearable sleep dataset and transformed wearable sleep dataset respectively. Vertical lines mark the 0-100 percentile values in 5% intervals on the x-axis corresponding to each dataset; blue vertical lines are overlapped with red vertical lines.
Applying feature transformation without removing contextual confounders (Experiment IV)
We further investigated whether the same feature transformation procedure could improve the performance of the hospital model on wearable features without removing the contextual confounder of awake versus sleep. Similarly, we calculated percentile values of daily wearable features derived from awake and sleep data combined, replaced the feature value with the corresponding value from the hospital dataset, and evaluated the performance of the hospital model on the transformed features. The model had an AUROC = 0.566 (95% confidence interval: [0.554, 0.622]; Fig. 5A, orange dotted line), AP = 0.158 (95% confidence interval: [0.153, 0.205]; Fig. 5B, orange dotted line), Sensitivity = 0.256 and Specificity = 0.844 at break-even point, Sensitivity = 0.296 and 0.146 respectively when Specificity was at 0.8 and 0.9, when applied to the transformed wearable features without using sleep data exclusively. The model performance was slightly better than before feature transformation (AUROC: 0.565 versus 0.526, orange dotted line versus blue dash-dot line in Fig. 5A), but the improvement was not as substantial as when applying feature transformation to the sleep-only features (AUROC: 0.740 versus 0.643, red solid line versus green dashed line in Fig. 5A). These results suggested that both controlling for contextual cofounders and applying feature transformation to address dataset shift were important to enable good model performance.
Statistical analysis and comparison with previous work
We performed DeLong’s test22 to compare model performance between each pair of the four experiments and use Bonferroni method to correct for multiple comparisons23. We included the z-score, the p-value and the adjusted p-value in Supplementary Table 2. We found significant differences (p-value
We have shown that the hospital model trained for hospital-acquired infection prediction performed the best in detecting early signs of COVID-19 infection on wearable dataset when feature transformations were performed and when only sleep data were considered (AUROC = 0.740, Experiment III). Although this performance is viable for a system, it was lower than our previously reported solution using a model trained directly on wearable dataset with COVID-19 labels (AUROC = 0.82)6. This was expected because the hospital model was designed to be an economical minimal viable solution that uses no COVID-19 labels for training, and thus not capable of controlling for concept drifts and/or label shifts.
When overlaying risk scores with time from the Experiment III hospital model, on average subjects with positive COVID-19 test results showed risk score elevations around COVID-19 test time (Fig. 5C, black), whereas subjects with negative COVID-19 test maintained their baseline risk scores (Fig. 5C, blue). Based on a cut-off risk threshold of 15 (yielding 60% Sensitivity and 78% Specificity), we identified the days in which the model output exceeded the defined threshold within the 14-day window prior to COVID-19 testing to estimate the lead time of positive classification (see METHODS). We found that the Experiment III hospital model successfully predicted COVID-19 infection, on average, 2.2 days prior to testing (95% confidence interval: [1.95, 2.61] days). This lead time was slightly lower but comparable to our previously reported wearable solution of 2.3 days prior to testing6.
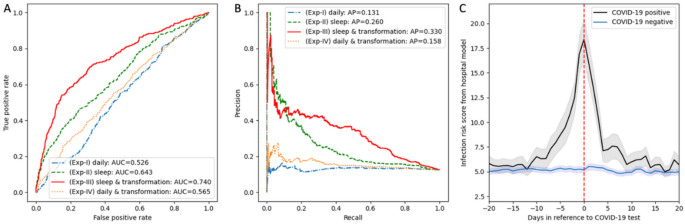
Hospital model performance and risk scores in detecting COVID-19 infection from wearable dataset. (A) Receiver Operating Characteristic (ROC) curves. Experiment I (blue dash-dot line): hospital model directly applied to wearable daily features. Experiment II (green dashed line): hospital model applied to wearable sleep-only features. Experiment III (red solid line): hospital model applied to wearable sleep-only features after feature transformation. Experiment IV (orange dotted line): hospital model applied to wearable daily features after feature transformation, without using sleep data exclusively. Area under the ROC curve (AUROC) for each experiment is included in the figure legend. (B) Precision-recall curve. Colors are the same as described in subplot A. Average Precision (AP) score for each experiment is included in the figure legend. (C) Mean infection risk score based on the output of the best generalized hospital model (Experiment III: sleep-only features + feature transformation) in 301 COVID-19 positive subjects (black) and 2,111 COVID-19 negative subjects (blue) as a function of number of days relative to the COVID-19 test time (red). Grey and light-blue shaded area depicts 95% confidence interval calculated from the standard error.
Data requirements for feature transformation (Experiments V, VI, VII)
Given that our best generalized hospital model (Experiment III) performed reasonable but inferior to our previous wearable model6, it is most sensible to use a hospital model for predicting COVID-19 in the absence of the wearable model, e.g. at the onset and during the early stage of the outbreak when data from COVID-19 positive cases were limited or unavailable to train a wearable model. Therefore, we investigated the data requirements of the generalized hospital model in Experiment III, in particular, the type and amount of wearable sleep data needed for the feature transformation. A favorable solution should require minimal COVID-19 positive instances. We performed three sets of additional experiments.
First, we asked whether illness data of COVID-19 were required for feature transformation (Experiment V). Interestingly, we found that baseline healthy data was sufficient because (1) using wearable sleep data from subjects that only reported negative test results for the feature transformation resulted in similar AUROC of 0.741 (Experient V-a, Supplementary Table 3), and (2) using wearable sleep data 4 weeks to 2 weeks before COVID-19 test – a time range when subjects were not infected – achieved similar results (AUROC = 0.741; Experient V-b, Supplementary Table 3).
Second, we asked whether wearable sleep data for feature transformation needed to be from the same subjects (Experiment VI, Supplementary Table 3). We randomly split the subjects in the wearable dataset into 5 folds, and for each subject we used subjects from the other four folds to transform the features of the given subject. The model performed with an AUROC of 0.741, suggesting that the wearable data used for feature transformation do not need to come from the same subjects used to test the model.
Third, we examined the minimum sleep data needed for feature transformation by benchmarking model performance against using sleep data from reduced number of days or from reduced number of subjects (Experiment VII). We gradually decreased the number of days from the 14 days prior to COVID-19 test where the wearable data were used for feature transformation (Experiment VII-a, Supplementary Table 3). We found that the model performed at AUROC of 0.74 when more than 2 days immediately preceding the COVID-19 test were used for feature transformation, and the model performed at AUROC = 0.73 when using data from the day before or two days before COVID-19 test for feature transformation. We also benchmarked against data from randomly selected days within the 14-day window prior to the COVID-19 test for feature transformation and found that the model performed at AUROC of 0.74 for all experiments – randomly selecting number of N days where N ranges from 1 to 13 days (Experiment VII-b, Supplementary Table 3). Further, we pooled all subject days and used random down-samples for feature transformation (Experiment VII-c, Supplementary Table 3). We found that the model performed at AUROC of 0.74 for all experiments of reduced subject days (number of reduced subject days: 25,000, or 20,000, or 15,000, or 10,000, or 7,500, or 5,000, or 3,000, or 1,000, or 500, or 300), even when only 300 subject days were used. Regarding the number of subjects needed, we used data from randomly down-sampled subjects for feature transformation and found that the model performed at AUROC of 0.74 for all experiments of reduced number of subjects (number of reduced subjects: 2,000, or 1,500, or 1,000, or 500, or 250, or 100, or 50, or 25), even when the number of subjects was reduced to 25 (Experiment VII-d, Supplementary Table 3).
Summarizing all the experiments (Supplementary Table 3), we concluded that healthy wearable data from 25 subjects collected in a period of 14 days for feature transformation would be sufficient to ensure the same model performance of AUROC = 0.74.