We developed a fully automated HISCL system for high-throughput detection of binding site–specific antibodies in patient specimens. By measuring region-specific antibody titers, this system enables detailed analysis of how individual epitopes contribute to the clinical immune response. We defined two distinct regions, designated Region A and Region B, within the RBD of the SARS-CoV-2 spike protein. These regions were determined based on the binding footprints of REGN10933 and REGN10987, respectively, which predominantly target different sides of the RBM (Fig. 1). This design allows separate quantification of antibodies directed against each region, providing deeper insight into epitope-specific immune responses. We verified the competition effect by conducting a competition assay with a human monoclonal IgG antibody sharing the same variable region as the competing antibody, which produced > 80% signal reduction (Fig. S1).
Initially we evaluated region-specific antibody titers in sera from vaccinated and virus-infected individuals (Fig. 2). Overall, the vaccinated group displayed higher antibody titers across all assessed regions than the convalescent group. This result suggests that three vaccine doses cause more efficient antibody induction than antigen exposure due to a natural viral infection.
Epitope-specific antibody titers. Antibody titers were calculated as the difference between measurements obtained without and with competitive antibodies. The x-axis displays four regions: S1 (the Spike S1 subunit), Region A + B, Region A, and Region B (Regions A and B are detailed in Fig. 1). (A) Antibody titers in samples from virus-infected individuals. (B) Antibody titers in samples from vaccinated individuals.
Next, we focused on samples with an S1 antibody titer ≥ 20 BAU/mL to assess the distribution of antibodies targeting the RBD (Fig. 3). The median for both virus-infected and vaccinated individuals exceeded 50% in regions A and B (Fig. 3A), indicating that the RBM is a major target for SARS-CoV-2 antibody induction, regardless of the infection or vaccination status.
Ratio of antibody titers targeting defined regions of the S1 subunit. The y-axis in each panel represents the percentage of total S1-targeting antibodies that bind to the indicated region(s). P-values were calculated using the Wilcoxon–Mann–Whitney test with R (***p < 0.001). (A) The ratio of antibody titers targeted at A + B of the S1 subunit. (B) The ratio of antibody titers targeted at A of the S1 subunit. (C) The ratio of antibody titers targeted at B of the S1 subunit.
To evaluate the antibody titer ratio of A + B in S1 and A in S1, we revealed a trend in which infected individuals had significantly higher ratios than vaccinated individuals (Fig. 3B). This suggests that vaccination with the ancestral monovalent BNT162b2, which encodes a stabilized full-length spike protein23, might promote a broader epitope recognition compared to natural infection, resulting in more uniform coverage of both RBM and non-RBM epitopes.
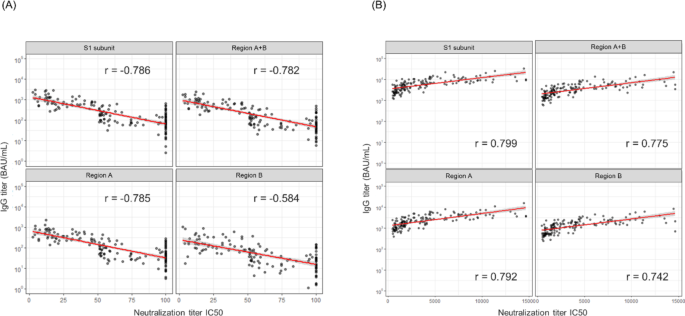
We further investigated the relationship between the activity of nAbs and epitope regions targeted by the antibodies. Spearman’s rank correlation coefficient revealed the strongest correlation between the antibody levels of the S1 subunit and neutralization activity in both the virus-infected and vaccinated individuals (Fig. 4A and B). Interestingly, focusing solely on a narrow antibody-binding domain did not necessarily enhance the correlation with neutralization. While RBM-targeting antibodies directly block ACE2 binding and exhibit potent neutralizing activity11, our findings suggest that non-RBM antibodies may also contribute to neutralization through indirect mechanisms.
Correlation between the antibody titer and neutralizing activity. The red and gray areas indicate the regression lines and 95% confidence intervals, respectively. Spearman’s rank correlation coefficient was calculated using R. (A) Correlations in the infected samples. (B) Correlations in the vaccinated samples.
Although several previous studies have reported such effects24,25,26, our study provides additional evidence from clinical samples, highlighting the importance of targeting multiple epitopes for effective SARS-CoV-2 neutralization. Future studies should investigate (i) cross-neutralization, (ii) targeting the N-terminal domain (NTD), (iii) targeting the S2 subunit, (iv) preventing S1 shedding, (v) steric hindrance, and (vi) preventing conformational changes to further elucidate the roles of non-RBM antibodies. Establishing an antibody screening system using HISCL could facilitate the discovery of such antibodies, contributing to the development of next-generation therapeutics.
Currently, the ACE2 competition assay is the primary platform for determining the neutralizing activity of antibodies in clinical use as competitive virus neutralization tests (cVNTs) against SARS-CoV-227. However, this method is limited to detecting only ACE2-blocking antibodies, overlooking those targeting other key epitopes. In contrast, our system employs two monoclonal antibodies that target complementary, non-overlapping epitopes within the RBM, enabling a more refined characterization of antibody responses. This approach can reveal how specific viral mutations might diminish antibody binding to one epitope while sparing the other, providing valuable insights into immune escape mechanisms. Additionally, using clinically approved antibodies (Casirivimab and Imdevimab) links our results directly to therapeutic strategies, illustrating how serum-derived antibodies may compete with or complement these treatments in real-world scenarios.
In conclusion, we developed a novel high-throughput system using a fully automated immunoassay to detect the antibody epitopes in specimens, focusing on two regions of the SARS-CoV-2 RBD. Clinical sample analysis revealed that the vaccinated individuals exhibited higher antibody titers across all epitope regions than the virus-infected individuals. Our findings suggest that antibodies binding to regions other than RBM also indirectly contribute to the neutralizing activity. Overall, our highly automated immunological assay system can facilitate the design of novel diagnostic antigens tailored to the specific objectives of antibody testing in clinical settings.