Further details of methods are given in the Supplementary Materials.
Estimating the transmissibility of MERS-CoV in camels
We estimated two different measures of the transmissibility of MERS-CoV in camels: the Force of Infection (FoI, λ) defined as the rate at which susceptible animals become infected, and the reproduction number (R0) defined as mean number of individuals infected by a single infected individual in an entirely susceptible population.
FoI
We fitted catalytic models of seroconversion to age-stratified seroprevalence estimates from across Africa, South Asia and West Asia collated previously through a systematic review8. Since the catalytic modelling approach assumes seroprevalence estimates are derived from a random cross-sectional sample of individuals, we excluded 3 of the 19 reviewed studies based on their sampling strategies (please see Supplementary Materials for more details on study inclusion). To make the geographical range of the FoI estimates as comprehensive as possible, we used seroprevalence measures from one additional study published after the systematic review. This allowed us to include camel populations in Senegal and Uganda which were not previously represented in the literature16. The tests used to determine seropositivity varied between studies and included both neutralisation tests (NTs) and non-neutralising Enzyme Linked Immunosorbent Assays (ELISAs) (Table 1). NTs are shown to be highly specific to MERS-CoV antibodies with little cross-reactivity with other camel coronaviruses26,27,28,29. MERS-CoV IgG ELISAs have been measured to be 99% specific when correlated with NTs30,31. Whilst ELISAs are considered more sensitive than NTs as they can pick up non-neutralising antibodies32, the seroprevalence measured by studies using NTs often approaches or reaches 100% in adult camels suggesting that – assuming they are indeed highly specific – they must also be highly sensitive. Therefore, we assumed a high sensitivity (98% for both test types) and specificity (99.5% for NTs and 98.5% for non-NTs) in our central results. We then conducted a sensitivity analysis assuming NTs to have a lower sensitivity of 85%.
We compared the fit of four models of seroconversion. In model 1 we assumed that all animals are born seronegative and become seropositive at a constant rate λ, as originally conceptualised by Muench and now regularly applied to epidemiological data33,34. Since MERS-CoV reinfection has been documented in camels6,13,14, in model 2 we extended model 1 to allow for seroreversion – with protective antibodies waning at rate σ. In model 3 we extended model 1 to allow a proportion of calves to be born seropositive due to protective mAbs which wane at rate ω, as evidence suggests that calves born to seropositive mothers are shown to acquire MERS-CoV specific mAbs through colostrum11,12. Finally, our fourth model allowed for both mAbs and seroreversion. Please see the Supplementary Materials for equations describing the solutions used for each model. We fit the models within a Bayesian framework using Hamiltonian Monte Carlo (HMC) sampling algorithm implemented in the R software package rstan version 2.32.735. Whilst we estimated the FoI per study to account for potential true differences between the FoI across husbandry systems, we assumed antibody waning rates to be constant, estimating them globally across all the datasets. We assumed that the seroprevalence data was beta-binomially distributed and re-parameterised the beta-binomial distribution in terms of the mean probability of being seropositive and the overdispersion parameter k where k > 0 and a k approaching zero would indicate negligible overdispersion. A detailed reparameterization available in the Supplementary Material. In order to evaluate which of the models was best supported by the data, we compared their fit using the Deviance Information Criterion36.
The reproduction number (R0)
We estimated R0 of MERS-CoV in each study population by calibrating a dynamic model of MERS-CoV transmission (see next section) to the modal FoI, by varying the transmission intensity parameter, β, under different potential immunity scenarios. R0 was approximated as the product of β and the infectious period, γ. The one-to-one relationship between β and the FoI meant that the credible intervals (CrIs) around the FoI could be used to propagate the uncertainty into the R0 estimates.
Development of a dynamic model of MERS-CoV transmission in camels
Infection
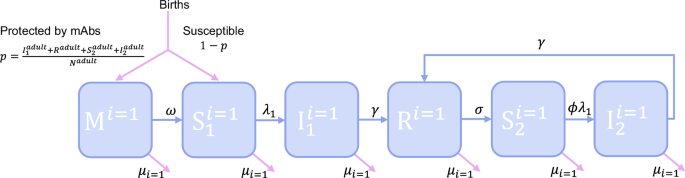
Based on what we know about camel demography from the literature, and our estimates of transmissibility and maternal antibody waning, we developed a stochastic, age-structured model of MERS-CoV transmission in camels. The model structure is represented schematically in Fig. 5, with a single age class shown for clarity. All symbols used are defined in Table 3 alongside the parameter values and their sources. Camels are born either entirely susceptible to MERS-CoV infection (state S1) or with complete protection by mAbs (state M) which wanes at a rate ω with calves becoming susceptible after an exponentially distributed period with a mean of ~2 months as estimated from the age-stratified seroprevalence data. The proportion of calves born in state M is dictated by the proportion of animals of reproductive age (>4 years) which have been previously infected. Animals in S2 become infected and transition to state I1 with the FoI, λ1, defined as the product of the effective contact rate, β, and the proportion of individuals in the population which are infectious:
$$\begin{array}{c}{\lambda }_{1}=\beta \frac{{I}_{1}+r{I}_{2}}{N}\end{array}$$
(1)
where I2 is the number of reinfected individuals, r < 1 and represents the relative infectiousness of reinfections compared to first infections, and β is varied to calibrate λ1 to our FoI estimates from age-stratified seroprevalence data. We chose to model transmission as frequency-dependent rather than density-dependent. In many countries camel farming is still overwhelmingly extensive25, with herds living and moving within large areas. In such settings, transmission is likely to depend more on the proportion of individuals infected rather than the absolute number. Modelling frequency dependence also enabled us to consider transmission at a wide range of scales and population sizes without requiring detailed data on animal stocking densities. To do this in a meaningful way whilst incorporating density-dependence would have required additional data on the relative sizes or densities of camel farms, grazing areas and markets and movement between these settings that is not currently available. No data is available on the potential latent period following MERS-CoV infection in camels. Infected animals are assumed to be instantaneously infectious. The period spent in state I1 is exponentially distributed around a mean of 14 days in agreement with the duration of shedding reported in longitudinal studies6,11,13.
A schematic of the model is shown for the youngest of i age classes (i = 1). Animals are born either protected by mAbs or susceptible, with a probability p, of being protected proportional to the fraction of adults previously infected at the time of birth. The states represented are defined as follows M – maternally acquired immunity, S1 – susceptible, I1 – first-time infected, R – recovered, S2 – susceptible again following infection and I2 – reinfected. This structure is repeated for each of the further 48 age classes. The symbols used to represent rates are defined in Table 3.
Immunity
Whilst our inference from age-stratified seroprevalence suggests that under catalytic model assumptions antibodies may be long-lasting following infection, documented reinfection of seropositive animals and rapid reinfection in high transmission intensity environments indicates that MERS-CoV seropositivity is not a good proxy for protective immunity in camels6,13,14. We therefore explored multiple reinfection scenarios (Table 3). Following a short period of complete immunity in state R, individuals become susceptible to reinfection in state S2. Most animals found to be shedding MERS-CoV in field surveys are calves and naïve animals, suggesting there is some long-term protection offered by past infection20,37,38,39. To reflect this, the degree of susceptibility in state S2 is modelled to be less than that experienced by individuals in state S1, meaning individuals in state S2 experience a reduced FoI, φλ1. Reinfected individuals in I2. are modelled to be less infectious than individuals in I1. This is based on measures of viral load collected in the control arm of the ChadOx1 MERS vaccine field study in camels6. We digitally extracted the daily mean viral load for seronegative calves (which we assumed to be infected for the first time during the study) and seropositive calves (which we assumed to be reinfected during the study) in the unvaccinated control group in from Fig. 4A of the online publication using PlotDigitizer version 2.240. We then calculated the difference between the area under the viral load curve for each of the two groups. Reinfected animals were approximately 1% as infectious as first-time infected animals when assuming a linear relationship between viral load and infectiousness. The relationship between viral load and infectiousness is not well characterised. A trial of the MVA-based vaccine candidate in camels measured a similar decrease in a measure of infectious viral particles and a measure of viral RNA shedding, following vaccination of four calves7. However, the study was not designed to have the power to reliably define the relationship between infectious virus particles and infectiousness. Therefore, whilst our central results assume a linear relationship, with reinfected individuals 1% as infectious as first-time infections, we include a sensitivity analysis assuming that the relationship between viral load and infectiousness is logarithmic, with a relative infectiousness of 50% for reinfected animals.
Age structure
Inclusion of age structure is vital given the strong dependence of infection status and seroprevalence on age, as well as for simulating age targeted interventions. Fine age structure is especially important up until the age of four years to enable accurate representation of age within the window where first infections are happening and accurate, age-targeted intervention modelling. For this reason, the model is stratified into month-wide classes, with camels moving to the next age-strata every 30 days in a 360-day year. From the 48th month wide class, camels enter a class aged >4 years where they remain until death.
Births
Camel calving is reported to be strongly seasonal19,41,42,43,44,45. Studies in KSA report most calves being born between October and March, with one study quantifying this at 83% during the high season19. The calving season is very similar in Egypt where it is reported between October and April45 and in Nigeria where surveyed pastoralists identified the calving peak to occur in the early dry season between October to December41. To capture this seasonality, the number of births per day is drawn from a Poisson distribution with a mean of αΝ0 where α varies annually as a function of cosine (Eqs. (2) and (3)) and N0 is the initial population size. The strength of seasonality can be weakened by setting δ < 1 during sensitivity analyses. However, when δ = 1, 82% of births fall between October and March which is in line with the 83% reported for camel births in Qassim, KSA19.
$$\begin{array}{c}{births} \sim {Pois}\left(\alpha \left(t\right) \,*\, {N}_{0}\right)\end{array}$$
(2)
$$\begin{array}{c}\alpha \left(t\right)=\overline{\alpha \left\{1+\cos \left(\frac{2\pi t}{360}\right)\right\}}\end{array}$$
(3)
Deaths
Since MERS-CoV causes very mild disease in camels, infection is modelled to have no bearing on mortality. Camels die off from each disease state compartment at the same age dependent average rate μi, with the mean number of deaths per day being equal to the size of the compartment multiplied by 1 – e -μi. The model assumes a higher probability of calf death in the first two years of life than in adulthood, as reported in KSA19,46. The modelled mortality rates are calibrated to the birth rate to give a stable population size and are equivalent to ~40% mortality in the first two years of life and ~12% afterwards, similar to overall mortality estimates for populations in KSA which are described in the literature as 10–26%, depending on herd type46.
Structure
For large populations, it becomes unrealistic to assume populations are well mixed. For example, in the population of ~10,000 camels in Laikipia county, Kenya47, an individual camel is far more likely to have contact with individuals in its own herd or grazing area than with animals in other areas of the county. The movements and interactions between herds of camels are not well documented. To explore the effect population structure has on dynamics, we developed a rudimentary structured population model where sub-populations or patches are arranged over a grid (Supplementary Fig. S5). Individuals are most likely to be in contact with other individuals in the same patch, less likely to meet individuals in neighbouring patches, and do not meet individuals in distant patches. Until better data on population structure allows a more accurate representation of networks and movements of camels within a region, the grid serves as a naïve representation of this reality.
We coded the model in R48 version 3.5.3, using the package odin version 1.5.1149 and ran stochastic iterations using odin.dust version 0.3.1350,51.
Estimating the Critical Community Size (CCS)
To evaluate the CCS of MERS-CoV in camel populations, we estimated the size of the population required for transmission to be sustained for at least 25 years in a closed population with no external sources of infection. The CCS was defined as the population size at which transmission was sustained in at least 50% of stochastic model runs. We ran the model using population sizes ranging from 500 to 1,000,000 and estimated the precise population at which 50% persistence was achieved using linear interpolation. For our central results we fixed the number of patches at 25, varying the patch size to represent different total population sizes. We conducted a sensitivity analysis to understand the effect patch size had on transmission persistence varying the number of patches from 9 (a 3 × 3 grid) to 49 (a 7 × 7 grid).
Evaluating the periodicity of infections
To determine the average time between peaks in infections we estimated the autocorrelation between each simulated time series of infections and lagged versions of itself using Pearson’s correlation test implemented through the acf function in the R “stats” package. The lag that maximised the autocorrelation coefficient was used to estimate the periodicity, for example if the lag that maximised the autocorrelation coefficient was between 350 and 370 periodicity was classified as annual. Very short lags of <100 days and any acf below the significance level using 95% confidence interval (CI) were excluded.
Estimating vaccine impact
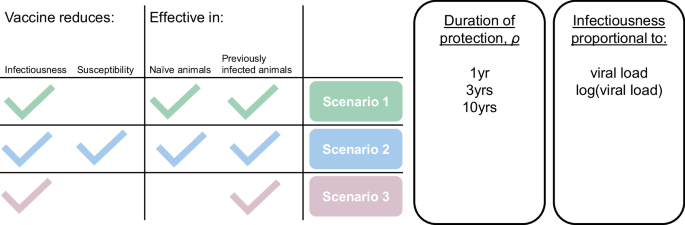
We extended the transmission model to simulate vaccination by duplicating the set of disease states to create a parallel set of vaccinated states. Although two vaccine candidates have been shown to reduce viral shedding in camels, uncertainty remains around their ability to reduce susceptibility and around the effectiveness of the ChAdOx1 MERS vaccine in naïve animals. Due to these uncertainties, three main scenarios are modelled (Fig. 6). In our central scenario 1, the vaccine reduces infectiousness but not susceptibility to infection for all vaccinated animals. This scenario reflects the finding that all previously naïve vaccinated animals became infected when challenged. Challenge doses administered intranasally or by confinement with multiple infectious animals could be much higher than the average natural exposure, and 1/5 of the previously infected ChAdox1 vaccinated animal did not become infected despite challenge so we also explored an alternative scenario 2 in which the vaccine reduces both infectiousness and susceptibility for all vaccinated animals. Finally, we explored a third scenario in which the vaccine reduces both infectiousness and susceptibility but only in animals that have been previously infected. Although the MVA study measured a large reduction in infectiousness of previously seronegative vaccinated animals, the ChadOx1 vaccine was only measured to reduce shedding in previously infected vaccinated animals. Authors suggest the low efficacy in this group could be due to the naive animals being very young, but their age was comparable with those used in the MVA study. Parameters used in vaccination simulation are presented in Supplementary Table S5.
Due to the scope of the efficacy studies, it is not possible to estimate the rate of waning of vaccine-induced effects, 1/ρ. Instead, for each main scenario, three options are explored with effects lasting one, three and ten years. The relative infectiousness of vaccinated infected animals and of vaccinated reinfected animals compared to unvaccinated naïve animals was parameterised using viral RNA shedding data6, assuming that infectiousness is either proportional to viral RNA shedding or to the log of viral RNA shedding. Vaccination is implemented in an age dependent manner and occurs immediately at the point at which camels reach the age being targeted for vaccination. To evaluate the ideal age for vaccination under the model assumptions, the target age group was varied from one month old to four years old. The vaccine efficacy was not modelled to vary with age. In scenarios 1 and 2 vaccination was assumed to reduce the relative infectiousness of first-time infected animals (rv) by the same amount as natural infection reduces viral shedding in reinfected individuals. The relative infectiousness of reinfected vaccinated individuals (rinf_v) was estimated as 0.15% when a linear relationship between infectiousness and viral RNA shedding was assumed, and 33% when infectiousness was assumed to be proportional to the log of viral RNA shedding6. See Table 4 for the complete set of parameters used to model the effect of vaccination. Vaccine impact was measured as difference in incidence following annual vaccination over a ten-year period and potential to disrupt patch or population-level transmission.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.