Optimization of CUT&Tag for genome-wide profiling in Mtb
To map the location of G4s in Mtb genome-wide, we initially applied a chromatin immunoprecipitation (ChIP) protocol with BG4, the most used anti-G4 antibody17, followed by either qPCR or sequencing. Experiments were conducted under standard laboratory growth conditions and under oxidative stress conditions to compare the G4 genomic distribution in Mtb cultures exposed to environments mimicking host infection.
ChIP is a multi-step antibody-based technique which allows the identification of specific features within chromatin18. The process involves harvesting bacterial cells, formaldehyde cross-linking to preserve biological interactions, and cell lysis before chromatin shearing. The latter is a critical step that requires fragment distribution between 100-500 bp19. Immunoprecipitation with the target antibody follows, and enrichment of the genomic sequences is assessed via quantitative PCR (qPCR) or next-generation sequencing (NGS). In our set-up, each ChIP experiment consisted of three samples: IP (immunoprecipitated DNA), MOCK and input. The IP was treated with the antibody of interest, while the MOCK sample underwent immunoprecipitation in the absence of the antibody, serving as control for non-specific background noise. The input sample, consisting of unprocessed sheared chromatin, enabled qPCR data normalization and target enrichment evaluation18. After ChIP, qPCR assessed the effectiveness of the applied protocol, selecting three genomic regions as positive controls: zwf1 and mosR, previously shown to fold into G4s in vitro12 and lpqS, as an additional G4-forming region (Supplementary data 1). The G4 folding of these selected positive regions was confirmed by CD analysis (Supplementary data 1 and Fig. S1). As negative control, we identified a genomic region in the Rv1507A gene that, based on its sequence, could not fold into G4 in vitro. ChIP-qPCR analysis showed remarkable enrichment of IP positive samples with respect to the MOCKs, both in standard growth conditions and upon treatment with H2O2, indicating high BG4 specificity. Replicates were highly consistent, with positive regions at least five-fold enriched over the negative control, aligning with successful ChIP-qPCR data reported in the literature (Fig. S2A)20. Encouraged by these results, we proceeded with sequencing and data analysis. However, identifying a clear signal above the background proved to be challenging. Although we observed a general enrichment in putative G4 positive regions (i.e. zwf1) and no enrichment in the negative region Rv1507A compared to the input sample (Fig. S2B-C), the signal was characterized by broad enriched domains. Commonly used peak calling tools, such as MACS221 and Epic222, struggled to identify true enriched regions above the background. Interestingly, while G4-ChIP is widely and successfully applied in eukaryotes23,24,25,26,27, it has never been applied in bacteria, possibly due to different chromatin organization and accessibility28 that would explain the observed low signal-to-noise ratio. Since the noisy background limited an efficient peak calling, which is fundamental for bioinformatic analysis of the sequencing results, we moved to CUT&Tag, a technique known for providing a significantly lower background.
In fact, CUT&Tag is an advanced genomic profiling method that improves ChIP-seq in several aspects. It offers higher sensitivity, specificity, and reproducibility while requiring reduced input material and sequencing depth. Unlike ChIP-seq, CUT&Tag allows native-state chromatin mapping29. Its efficacy in profiling histone modifications and transcription factor binding, combined with reduced cost-effectiveness, has contributed to its widespread adoption across various organisms, including humans26,30,31,32, mice33, zebrafish34, plants35, and yeasts36. However, its application in bacteria has never been explored so far.
In the CUT&Tag protocol, cells are initially bound to magnetic beads and then mildly permeabilized to facilitate access of the target antibody and Tn5 enzyme to the cell genome29. We optimized Mtb cells binding to concanavalin A (ConA)-coated beads by measuring optical density upon varying incubation times (10 and 20 min), achieving optimal bead saturation at 20 min (Fig. S3A). To determine the optimal bacterial cell number, we performed CUT&Tag on three different sample sizes, 1, 10, and 20 million cells, quantifying bacterial DNA before and after library preparation (Fig. S3B, C). The highest DNA yield was obtained using 10 million cells.
To validate the novel CUT&Tag protocol in Mtb, we initially targeted the RNA polymerase (RNA Pol) subunit β, for which genome-wide ChIP-seq data were previously reported37. Experiments were conducted under standard growth and oxidative stress conditions. Briefly, bacterial cultures underwent mild formaldehyde fixation, and the permeabilized cells were incubated with the antibody, tagmented with pA-Tn5 transposase, and the PCR-amplified fragments were subjected to NGS. Each CUT&Tag experiment consisted of three biological replicates, exhibiting high Pearson’s correlation coefficients, indicative of robust reproducibility (Fig. 1A). Principal component analysis (PCA) revealed proper clustering according to the growth conditions (Fig. 1B), indicating high data quality and reproducibility among replicates performed in the two conditions. All samples achieved appropriate sequencing depth, with FRiP values largely exceeding 0.01, a threshold associated with successful experiments (Fig. 1C)38. Peak size distribution showed that the median peak size across all samples was ∼200 bp, as expected20 (Fig. 1D). Analysis of CUT&Tag signal distribution at the center of the called peaks showed a consistent enrichment in density heatmaps, both in high confidence peaks (Fig. 1E), defined as those present in at least two of three biological replicates, and in each replicate (Fig. S4) in the two growth conditions, supporting the efficiency of immunoprecipitation.
A Correlation plot among CUT&Tag biological replicates (n = 3, R1, R2, R3) performed in standard growth (STD) and oxidative stress (OX) conditions. B Principal component analysis (PCA) for the three replicates of each condition. C FRiP analysis. Results are shown as mean ± s.d. of independent biological replicates (n = 3). D Density plot of peaks width distribution. E CUT&Tag mean signal distribution at high confidence peaks. Average plots (top panel) and density heatmaps (bottom panel) of CUT&Tag reads referring to peak center, within ± 1 kb distance. Data referring to STD and OX conditions are shown in blue and orange, respectively. Source data are provided as a Source Data file and supporting data 3.
We then conducted a comparative analysis between our CUT&Tag results and the previously published ChIP-seq data, where positive regions were identified by calculating an enrichment ratio over input at defined locations37. Applying an analogous analytical approach to our CUT&Tag data, using the IgG profile as background, we found that 94% (874/924) of ChIP-seq-identified regions were retrieved by CUT&Tag (Fig. S5). Furthermore, consistent with observations in eukaryotic systems29, CUT&Tag showed enhanced sensitivity, enabling the identification of additional enriched regions. As an example, we report the visual comparison of RNA Pol coverage signals from ChIP-seq and CUT&Tag at the rrl locus that showed enrichment in both techniques and in both CUT&Tag conditions (STD and OX) (Fig. S5B-C). Overall, these findings strongly support the efficacy and applicability of the CUT&Tag protocol in Mtb for genome-wide profiling studies.
Genome-wide RNA polymerase profiling by CUT&Tag in Mtb
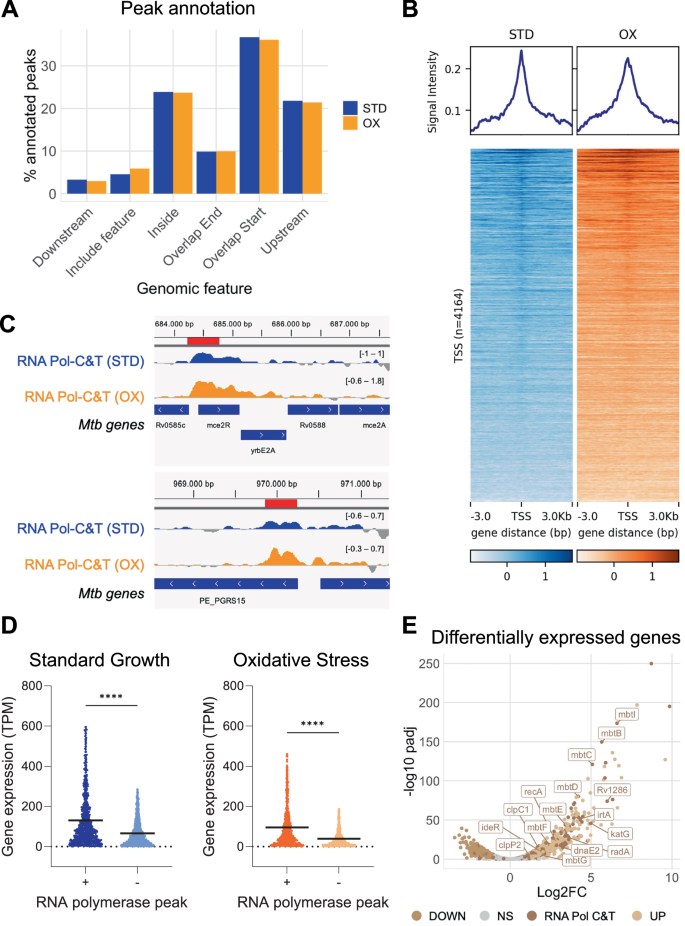
The sequencing profile of the RNA Pol-CUT&Tag revealed peaks across the entire Mtb genome: 1101 high confidence peaks were identified in standard growth, while oxidative stress yielded 986 peaks (Supplementary data 3). Regardless of the growth condition, RNA Pol-peaks showed a similar peak distribution in genomic feature annotation with a predominant location around gene start sites (36.7% and 36.1% in standard and oxidative stress condition, respectively), within genes (23.9% and 23.7%, respectively), and upstream of genes (21.8% and 21.4%, respectively) (Fig. 2A). As the β subunit constitutes an essential component of the holoenzymatic complex, the predominant location of peaks near gene start sites indicates that the enzyme is preparing to initiate transcription. The significant abundance of peaks residing within gene bodies or overlapping gene ends indicates that the RNA polymerase complex is either initiating or finishing the transcription process. This peak distribution was confirmed by density heatmaps of coverage at gene transcription start site (TSS) (Fig. 2B) and aligns with the pivotal role of the β subunit throughout the transcription process, in both conditions, from initiation to termination39. In addition, RNA Pol-CUT&Tag identified previously reported positive regions for RNA Pol binding in Mtb genome, rv0586/mceR2 and rv0872c/PE_PGRS1540 (Fig. 2C), further validating the protocol.
A RNA Pol-CUT&Tag high confidence peak annotation in standard growth condition (STD, blue bars) and upon oxidative stress (OX, orange bars). “Feature” stands for “gene”; “Include feature” indicates the peaks that extend over the whole gene coding sequence. B RNA Pol-CUT&Tag mean signal distribution at high confidence peaks. Average plots (top panel) and density heatmaps (bottom panel) of CUT&Tag reads in standard growth (STD, blue) and oxidative stress (OX, orange) conditions referring to peaks center, within ± 3 kb distance. C Visualization tracks of RNA Pol-CUT&Tag profiles, referred to known RNA Pol binding sites. Gene annotation is reported at the bottom of each track. D Expression levels of Mtb genes presenting (+) or not presenting (−) RNA Pol-CUT&Tag peaks in standard growth (left panel) and in oxidative stress conditions (right panel). Statistical differences were analyzed using unpaired t-test with two tails (STD: n (+) = 866, n (−) = 2649, p-value < 0.0001 (****); OX: n (+) = 740, n (−) = 2637, p-value < 0.0001 (****)); Mean values are reported as horizontal black lines. E Volcano plot illustrating differentially expressed genes upon oxidative treatment. Significant up/downregulated genes were identified using Deseq2 with p-value = 0.05 and Log2FC = 1. Dark brown dots indicate RNA polymerase-retrieved genes identified in oxidative stress conditions. Genes reported to be implicated in the response to oxidative stress are highlighted. Source data are provided as a Source Data file and supporting data 3.
Since RNA Pol is the primary enzyme involved in transcription, we integrated CUT&Tag results with Mtb gene expression levels from RNA-seq data. Raw read counts were converted into transcripts per million (TPM) for individual genes and associated with their corresponding genetic locus. Gene expression levels were then correlated with the presence or absence of RNA Pol-CUT&Tag peaks. A statistically significant increase in gene expression was observed in regions exhibiting RNA Pol-peaks versus those lacking them, both under standard and oxidative stress conditions (Fig. 2D). Genes were categorized into three groups based on gene expression interquartile (IQR) values (low, medium, and high). A substantial increase in the expression levels was observed for all RNA Pol-peaks across all categories (Fig. S6), consistently associating the presence of RNA Pol-peaks with enhanced transcriptional activity.
Differential gene expression analysis was subsequently performed between standard and oxidative stress conditions. It is established that Mtb treatment with 5 mM H2O2 substantially alters the bacterial expression profile41. Genes were considered differentially expressed when meeting criteria of log2-fold change exceeding 1 or falling below −1, with adjusted p-value < 0.05. Under these criteria, 732 and 650 genes were found to be upregulated and downregulated, respectively. Among these, 211 upregulated and 82 downregulated genes were associated with an RNA Pol-peak, constituting 28.8% and 12.6%, respectively, of the differentially expressed genes in response to oxidative stress (Fig. 2E). In these conditions, the induction of scavenging enzymes becomes also crucial for Mtb survival42,43. Indeed, katG, which efficiently reduces reactive oxygen species (ROS) concentrations, was strongly upregulated, along with cysN, responsible for sulfate activation in cysteine biosynthesis, and enzymes involved in the DNA damage response pathway. Other upregulated genes under oxidative stress conditions included iron-related genes such as the Mtb gene cluster encoding mycobactin (iron-chelating siderophore41), irtA and ideR (involved in iron import and regulation, respectively), DNA repair enzymes (recA, radA, and dnaE241,42), and protein repair or degradation systems (clpC1 and clpP2).
These results collectively validate the efficiency of the CUT&Tag protocol in profiling the Mtb genome, both under standard growth and oxidative stress conditions.
G4-CUT&Tag reveals a G4 landscape enriched in two-tetrad G4s
We then applied the CUT&Tag protocol using the BG4 antibody to define the G4 landscape in the Mtb genome under standard and oxidative stress growth conditions. The G4-CUT&Tag replicates showed high correlation values among them (Fig. S7A), expected fragment distribution44 (Fig. S7B), and exceeded the accepted threshold for library complexity and sequencing depth (Fig. S7C). Quantification of CUT&Tag signal at G4-peaks showed that the BG4 signal was significantly higher than the negative control (IgG-CUT&Tag): linear regression analysis of antibody coverage correlation yielded non-linear signal distribution with significant adjusted R2 values of 0.63 and 0.67 for standard and oxidative conditions, respectively, indicating that the BG4 antibody is specifically and reliably detecting the G4 structures in both conditions, with a slightly stronger correlation in oxidative condition (Fig. S7D). BG4 enrichment was further supported by positive values obtained from the difference of the log2 signal at the majority of G4 sites (Fig. S7D). Coverage distribution analysis revealed high signal density at the center of both high confidence peaks (Fig. S7E) and peaks separately identified in the biological replicates (Fig. S9), validating the G4-CUT&Tag approach in both tested conditions (Fig. S7E). Visual inspection of G4-CUT&Tag sequencing profiles showed remarkable improvement in signal-to-noise ratio (Fig. S7F) with respect to G4-ChIP data (Fig. S2C), enabling the identification of well-defined peaks distributed across the entire Mtb genome.
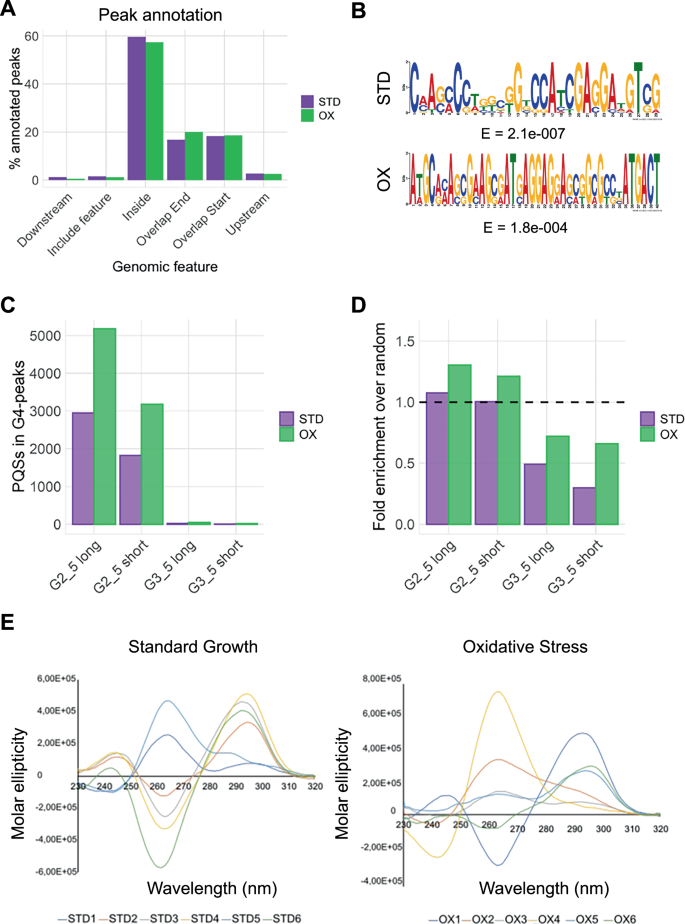
Notably, G4 peak annotation showed an unprecedented distribution of G4s, with almost 60% of peaks in both conditions located within gene bodies. The remaining peaks mostly overlapped with gene start and end sites (Fig. 3A). This distribution strongly differs from the typical eukaryotic pattern, where G4s predominantly accumulate at promoter regions23,25, thus suggesting different regulatory mechanisms in Mtb. Motif analysis of high confidence peaks confirmed G-rich patterns capable of G4 formation in both conditions (Fig. 3B). Consequently, we applied the G4 prediction algorithm Quadparser45 on high confidence peaks. We screened for motifs with the following characteristics: i) G-tracts of at least 2 or 3 guanines each; ii) loop length up to 7 (short) or 12 (long) nucleotides. Surprisingly ~99% of putative G4-forming sequences consisted of two-guanine tracts and short loops (Fig. 3C), generally associated with weaker G4s. Almost all G4 peaks contained this type of G4-forming sequences (Fig. S8). To assess the significance of our G4 prediction, we conducted a randomization analysis. We extracted random fragments from the entire Mtb genome, with abundance and average length comparable to that of CUT&Tag peaks (2000 fragments of approximately 260 bp), and applied the Quadparser algorithm to these random fragments, in accordance with the analysis on G4 peaks. We then calculated the fold enrichment of CUT&Tag samples over random fragments and observed that 2-tetrad G4s, except for those with short loops, exhibited a fold enrichment higher than 1, indicating that their identification is statistically significant and did not occur by chance. On the other hand, 3-tetrad G4s showed a fold enrichment much lower than 1, confirming the statistical significance of their absence from the pool of the identified peaks (Fig. 3D).
A G4-peak annotation in standard (STD, purple bars) and oxidative conditions (OX, green bars). “Feature” stands for “gene”; “Include feature” indicates the peaks that extend over the whole gene coding sequence. B MEME motifs obtained from G4-peaks in standard (STD) and oxidative conditions (OX). Motif logo and E-values are reported. C G4 motifs prediction of retrieved peak sequences, performed with the Quadparser mapper tool, in both tested conditions. G tracts composed of at least 2 or 3 Gs and up to 5 Gs were considered, with short (0-7 nucleotides) and long (0-12 nucleotides) loop lengths. D Fold enrichment of Quadparser-identified sequences over randomly generated fragments in the Mtb genome. E CD spectra of representative sequences derived from standard growth (left panel) and oxidative stress (right panel) conditions. Source data are provided as a Source Data file and supporting data 4.
CD spectroscopy validated G4 folding for six representative immunoprecipitated sequences from both conditions (Supplementary data 1, Fig. 3E). Most sequences showed the typical CD signature of a G4 with antiparallel topology, characterized by two positive peaks at λ ~ 240 and 290 nm and a negative one at λ ~ 260 (STD2, STD3, STD4, STD6, OX1, OX6). The remaining sequences showed a mixed G4 topology, with positive peaks at λ ~ 260 and 290 nm, with different intensities. Overall, all tested sequences were confirmed to fold into G4s, further validating the reliability of our G4-CUT&Tag in Mtb.
G4-bearing genes under oxidative stress are associated with lower transcript levels than in standard growth conditions
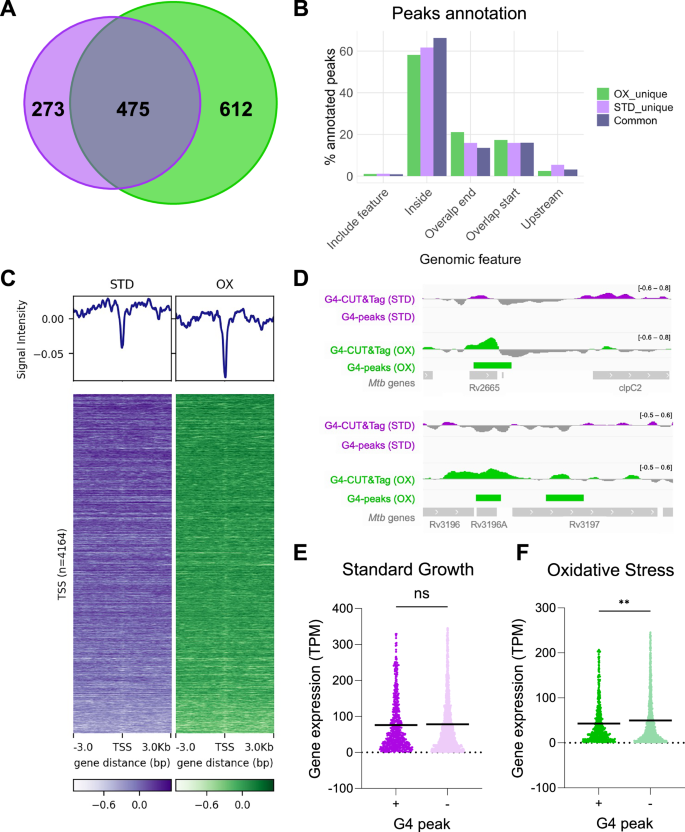
We compared G4 enrichment between standard growth and oxidative stress conditions. Remarkably, oxidative stress yielded more G4 high confidence peaks (1087) than standard growth (748) (Supplementary data 4). We identified 475 peaks shared between the two conditions, representing consistently folded G4s. Notably, over 50% of G4 peaks were unique to the oxidative condition, indicating that G4 folding was induced in response to the oxidative treatment. Conversely, less than 40% of peaks were unique to standard growth condition, suggesting that G4s may be more relevant in oxidative stress response (Fig. 4A). Genomic distribution analysis of unique and common peaks confirmed previous observations, with approximately 60% enrichment in gene bodies (Fig. 4B) and low abundance of BG4 signal at gene TSS (Fig. 4C). Visual inspection of unique G4-peaks in oxidative conditions further confirmed the high signal-to-noise ratio and efficiency of the CUT&Tag analysis (Fig. 4D).
A Venn diagram representing the G4-CUT&Tag peaks shared between standard (STD, purple) and oxidative (OX, green) conditions. B Genomic annotation of shared and unique peaks identified in standard and oxidative conditions. C Average plot (top) and density heatmap (bottom) of G4-CUT&Tag peaks in standard (purple) and oxidative (green) conditions. Peak signal distribution refers to gene TSS within ± 3 kb distance. D Visualization of G4-CUT&Tag profiles for oxidative unique peaks. Genomic tracks are reported as the difference between G4-CUT&Tag and IgG-CUT&Tag (grey) signals, both derived from the mean of three biological replicates. Expression levels of Mtb genes presenting (+) or not presenting (−) G4-CUT&Tag peaks in standard (E) and oxidative conditions (F). Statistical differences were analyzed using the unpaired t-test with two tails (STD: n (+) = 592, n (−) = 2906, p-value = 0.5028 (ns); OX: n (+) = 779, n (−) = 2590, p-value = 0.0014 (**)). Mean values are reported as horizontal black lines. Source data are provided as a Source Data file and supporting data 4.
Integration of RNA-seq analysis with G4-CUT&Tag data revealed no significant difference in transcript levels between G4-bearing and non-G4 genes under standard growth conditions (Fig. 4E). In contrast, oxidative stress induced an overall reduction in gene expression likely induced by the treatment41,42, with G4-bearing genes showing statistically significant lower levels of transcripts compared to non-G4 genes (Fig. 4F).
Detailed analysis revealed that high and medium-expressing G4-bearing genes underwent significant downregulation compared to non-G4 counterparts, while low-expressing genes showed no significant difference (Fig. 5A). To further investigate this finding, we analyzed the G4 peaks unique to the oxidative condition and compared their transcript levels in both standard and oxidative conditions: we found a significant downregulation upon G4 folding (Fig. 5B), therefore further supporting that G4 formation in the Mtb genome during oxidative stress is associated with gene expression downregulation.
A Expression levels of Mtb genes presenting (+) or not presenting (−) G4-CUT&Tag peaks in the oxidative stress condition grouped into three IQR categories according to their expression levels: High (left panel), Medium (central panel) and Low (right panel). Statistical differences were analyzed using the unpaired t-test with two tails (High: n (+) = 195, n (−) = 648, p-value < 0.0001 (****); Medium: n (+) = 390, n (−) = 1294, p-value < 0.0001 (****); Low: n (+) = 194, n (−) = 648, p-value = 0.9753 (ns)); Mean values are reported as horizontal black lines. B Expression levels in both standard (n = 542) and oxidative (n = 526) conditions of genes presenting G4s only in oxidative conditions and normalized to housekeeping sigA expression (p-value < 0.0001 (****)). Statistical differences were analyzed using the unpaired t-test with two tails and mean values are reported as horizontal black lines. C–E Expression levels of genes presenting G4s only in the oxidative stress conditions and grouped into three IQR categories according to RNA occupancy difference between oxidative and standard conditions. Statistical differences were analyzed using the unpaired t-test with two tails (High: n (+) = 38, n (−) = 33, p-value = 0.1010 (ns); Medium: n (+) = 100, n (−) = 98, p-value < 0.0001 (****); Low: n (+) = 45, n (−) = 46, p-value < 0.0001 (****)); Mean values are reported as horizontal black lines. F GO enrichment analysis of unique oxidative stress G4-bearing genes (FDR = 0.05). The horizontal axis represents the fold enrichment; the vertical axis represents the GO term or functional category; the size of the dot represents the number of genes in the GO term and the color of the dot represents the p-adjust value. Source data are provided as a Source Data file and supporting data 4.
We next analysed RNA Pol occupancy (Fig. 2) at genes presenting G4s only in the oxidative condition. We calculated the difference in RNA Pol coverage between oxidative and standard conditions at these G4 sites, and we defined three subsets according to the IQR method: Low, representing lower RNA Pol occupancy in oxidative condition; Medium, representing similar RNA Pol occupancy between the oxidative and standard conditions; High, showing higher RNA Pol occupancy in oxidative stress condition. Next, the genes included in the subsets were associated with their relative expression values in the two conditions. This analysis revealed that genes in the Low dataset (Fig. 5C) showed significant gene downregulation under oxidative stress. This behavior reflects genomic sites where G4 folding induced by oxidative stress is associated with a block in transcription. This result was obtained also with genes in the Medium dataset (Fig. 5D). Notably, genes in the High dataset (Fig. 5E) did not show an increase in gene transcription, even in the presence of high RNA Pol coverage, suggesting that the presence of G4s reduced RNA Pol activity also in this dataset.
We then investigated the possible role of genes presenting G4s only in oxidative conditions (Supplementary data 2) by performing GO enrichment analysis, which mostly showed association with metabolic and biosynthetic processes and cell wall biogenesis (Fig. 5F). Detailed analysis on expression-based subgroups (Fig. S10A) revealed that downregulated G4-bearing genes were mainly involved in cell wall organization and biogenesis, while non-differentially expressed genes were associated with biosynthetic pathways (Fig. S10B, C). The up-regulated group included too few genes for significant analysis.
In summary, our findings indicate that the G4-bearing genes recovered during Mtb stress response predominantly engage in various metabolic processes, as well as catalytic and binding functions. Interestingly, the identified enriched pathways are typical of Mtb stress-induced response, both at transcriptomic and proteomic levels46, suggesting that G4s might be constitutive elements for Mtb response to stress.