Here, we describe EpiGeoPop (our tool for constructing spatially accurate model populations) and discuss our improvements and extensions to Epiabm (the ABM we use for further configuring our model populations and then running epidemic simulations). The original release of Epiabm did not implement pharmaceutical or non-pharmaceutical interventions, and used a simplistic implementation of spatially mediated infections. Our extensions to Epiabm include both forms of intervention and an improved method for weighting spatial infections. Open-source code for both EpiGeoPop and Epiabm are available at https://github.com/SABS-R3-Epidemiology/EpiGeoPop and https://github.com/SABS-R3-Epidemiology/epiabm.
EpiGeoPop
A pre-requisite for spatially accurate epidemiological modelling is the integration of spatial data into the modelling framework. To simplify these initial stages of model set-up, we develop a software tool named EpiGeoPop (Fig. 1) which constructs a geographically accurate density map from global population density data34 and converts this into a human-readable population configuration file. Our tool allows this input file to be generated for any country or region of interest for which the border and population density data are available in the Natural Earth (https://www.naturalearthdata.com/downloads/10m-cultural-vectors/) and European Commission, Joint Research Centre Global Human Settlement Layer34 databases, respectively. We use the 2015 version of the population density dataset, which includes regions on the scale of cities, provinces and complete countries. The 2015 data, alongside datasets for other years, can be found at: https://human-settlement.emergency.copernicus.eu/download.php?ds=pop
The EpiGeoPop workflow
For ease of use, we present a complete workflow as a user-friendly Snakemake pipeline35 implemented in Python. This pipeline reads in the border and population density data described above, and generates a population configuration file that describes the distribution of individuals across a spatial grid with a resolution 30 arc seconds (approximately 1 km2). We label these \(\sim 1\) km2 regions ‘cells’. EpiGeoPop allows these cells to be optionally split further into ‘microcells’, should a smaller resolution be required. In the simulations presented herein, we use a grid size of 3×3 totalling nine microcells per cell, though the specific number of microcells can be specified by the user.
Additionally, users can specify parameters to add a number of places and households to each microcell in the population. Overall, the population configuration output file contains one line per microcell, stating the parent cell of that microcell and its location, as well as the attributes of the microcell, namely the number of households, places, and individuals that are contained within it. This structured hierarchy of people, places and households existing in microcells, which in turn exist within cells, is provided for in EpiGeoPop to ensure compatibility with Epiabm. This may not be applicable depending on the ABM being used, in which case the user can simply opt not to break cells into multiple microcells or to define a number of places and households.
EpiGeoPop also generates country-specific age distributions, taken from the UN 2022 Revision of World Population Prospects36. These are processed to represent the proportion of individuals in 5-year age groups from 0-80, with a final group of over 80 year-olds, resulting in an age distribution compatible with Epiabm.
Details on how to execute the workflow, tailored to the user’s specific needs, are provided in the Supplementary Information, Section S1.1.
Visualising epidemic spread
We also provide code to generate time-series animations of disease spread across the region of interest (examples of this are provided in Supplementary Videos 1 and 2). This code is run as a single Python script that creates a GIF visualising the infection counts as a heatmap changing over time. To use this script, the output data from the epidemiological simulation should be provided in the format matching that generated by the Snakemake workflow (details are provided at https://github.com/SABS-R3-Epidemiology/EpiGeoPop/#generating-animations).
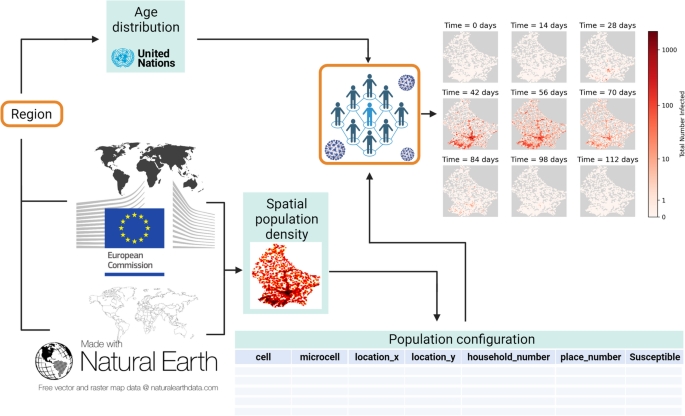
EpiGeoPop workflow. EpiGeoPop uses global population density data to generate population configuration files for any country, province, or city of interest. The configuration file contains one row per ‘microcell’ (the smallest defined spatial region) with columns recording the microcell and its parent ‘cell’, the location (in x and y coordinates) of that parent cell, and the number of households, places, and susceptible individuals in that microcell. In addition, age distributions are generated on a country-level basis. The choices of region and epidemiological model, highlighted in orange, represent the elements of the workflow determined by the user. As long as the overall format of the output file from the model matches that of the input file generated by EpiGeoPop (i.e., columns representing the location’s x- and y-coordinates and the number of individuals at that location), the visualisation module can be used. This allows geographically accurate visual summaries of disease dynamics to be generated in the form of GIFs. Created with BioRender.com.
Epiabm
We utilise Epiabm (v1.1.0) for all epidemic simulations, extending this software from the previously published version (v1.0.1). Epiabm includes algorithms for configuring age, household, and place structure within the model population. Households and places sit at the ‘microcell’ level, with microcells sitting within larger geographic regions termed ‘cells’ as above. The model uses a compartmental structure to simulate disease spread, with compartments for susceptible, infected (of varying severity), recovered, and dead individuals. Infected individuals are able to infect susceptible individuals through three transmission pathways: place-based, household-based, and spatial-based. Our extended version of Epiabm includes an altered weighting for spatial transmission, and implements a suite of time-limited events, including pharmaceutical and non-pharmaceutical interventions.
Improved weighting of spatial transmission
Spatial transmission across cells involves the random selection of one cell for each person a given infector should infect. The infectee is then selected at random from that cell’s population. In both CovidSim and the initial version of Epiabm (v1.0.1), this cell is selected with probability inversely proportional to its distance from the infector’s cell. Although travel between locations is not explicitly modelled in Epiabm, this method of spatial transmission has the effect of considering travel to two locations the same distance apart equally likely, irrespective of their population.
However, to accurately model the impact of heterogenous population density distributions, we adjusted the spatial infection mechanism such that an infectee is more likely to be selected from a higher-density cell than a lower-density cell the same distance away. In effect, this approximates the impact of transport networks with hubs centred on larger towns and cities37. The effect of this change is presented in Figs. 3 and 4 of the Results section ‘Population density influences epidemic dynamics’.
Non-pharmaceutical interventions
We additionally implemented the full suite of non-pharmaceutical interventions present in CovidSim4: case isolation, household quarantine, place closure, and social distancing. In our implementation of these interventions, we closely follow that used in CovidSim. Interventions can be activated at a specific simulation time, or when the total number of cases crosses a specified threshold, capturing some of the flexibility present in CovidSim. Multiple interventions can be active at the same time, and the same intervention can be implemented over different time periods with varying severity. All non-pharmaceutical interventions influence house, place, and spatial infections. When case isolation is active, symptomatic or positive-tested individuals will isolate within their household for a given period (for a description of the testing functionality please see the section ‘Pharmaceutical interventions’). Household quarantine is built upon case isolation and restricts members of the household of the isolated individual to also stay at home for a defined number of days. Place closure allows for the closure of specific place types, such as schools, workplaces, or outdoor spaces, to be simulated. Social distancing reduces the likelihood of house, place, and spatial infections. We allow for two strengths of this reduction, which can be applied to different age groups. Simulations including these interventions have the expected effect of delaying and reducing the peak of infections (Fig. S3)
Pharmaceutical interventions
In addition, we implement two pharmaceutical interventions: mass vaccination and infection testing to determine case isolation. Mass vaccination can be prioritised according to individual characteristics, and the protection afforded by vaccination can be parametrised by the user. Infection testing includes two testing streams for which parameters including sensitivity, specificity, testing capacity and type of user can be set. These two streams can therefore be adjusted to represent different testing modalities, such as PCR and lateral flow tests. Testing can then be used as a criterion to select an individual for case isolation instead of using their symptomatic status.
International travel
Finally, we develop a simple travel framework to capture international travel. Through this functionality, border closure and isolation of international arrivals (either within an existing household or in separate accommodation) can be activated as an additional intervention.
Combining EpiGeoPop and Epiabm to configure and run simulations
In the simulations presented herein, we generate population configuration files of real countries using EpiGeoPop, and simulate the spread of disease through these populations using Epiabm. Though we use the terminology of cells and microcells in both EpiGeoPop and Epiabm, the concept of these spatial regions is scale-agnostic in Epiabm, while in EpiGeoPop the resolution of the spatial data used means cells are considered to be \(\sim 1\) km2 regions, which can optionally be broken down into microcells.
We use the population configuration file output by EpiGeoPop to define the spatial distribution of the population. We then apply an algorithm included in Epiabm to assign ages and households to individuals according to the age and household distributions for the country of interest. The age distribution is also provided by EpiGeoPop, and we take the household distributions from data provided by the United Nations Population Division: https://www.un.org/development/desa/pd/data/household-size-and-composition. We verify this algorithm by comparing the age and household distributions of the configured population after application of the algorithm to the input distributions (Fig. S1).