LNP X enables delivery of mRNA to resting T cells
We first assessed whether an LNP similar to an existing, FDA-approved LNP formulation (patisiran) was able to transfect non-stimulated primary CD4+ T cells by delivering a reporter mCherry mRNA. Our patisiran-like LNP (hereafter referred to as patisiran LNP) is formulated with DMG-PEG2000 instead of PEG2000-C-DMG. At the highest dose tested of 500 ng per 105 cells, mCherry expression was only detected in 2.1 ± 0.4% (mean ± SEM) of live cells (Fig. 1a and Fig. S1a) after 72 h incubation. Pre-stimulation of the CD4+ T cells with anti-CD3/anti-CD28 led to substantially higher transfection efficiencies of up to 51 ± 5.1% compared to non-stimulated T cells (Fig. 1a), though the toxicity associated with patisiran LNP treatment was greater in pre-stimulated compared to non-stimulated CD4+ T cells at higher LNP doses (Fig. S1b). These data show that successful mRNA delivery to primary CD4+ T cells using patisiran LNPs is dependent on the activation state of the T cell.
a CD4+ T cells from HIV-negative donors were rested or pre-stimulated with anti-CD3/anti-CD28, then treated with LNPs (similar to the patisiran lipid composition) encapsulating mCherry mRNA for 72 h at the indicated doses. Highest dose of mCherry-LNP X corresponds to 2.5 µg/mL. Transfection efficiency was determined by measuring mCherry expression using flow cytometry. Mean ± SEM, n = 6 donors. b Schematic representation of patisiran-like LNP formulation versus novel LNP formulation X. Mol %, relative molar percentage of indicated lipid in lipid mixture. Created in BioRender: Cevaal, P. (2025) https://BioRender.com/ypaicaz. c Size distribution of patisiran LNP (yellow) versus LNP X (green) encapsulating mCherry mRNA. Lines represent individual LNP batches. d Relative mCherry expression levels in Jurkat T cells treated for 24 h with indicated doses of patisiran LNP (yellow) or LNP X (green) encapsulating mCherry mRNA. MFI, median fluorescent intensity. Mean ± SEM, n = 2. e CD4+ T cells from HIV-negative donors were rested or pre-stimulated with anti-CD3/anti-CD28, then treated with LNP X encapsulating mCherry mRNA for 72 h at the indicated doses. Highest dose of mCherry-LNP X corresponds to 2.5 µg/mL. Transfection efficiency was determined using flow cytometry. Mean ± SEM, n = 6 donors.
We therefore modified the lipid composition of the LNP to enhance potency. First, the ionisable lipid DLin-MC3-DMA (MC3) was replaced with SM-102, an ionisable lipid previously shown to lead to greater cytosolic mRNA delivery through enhanced endosomal escape30. Second, the SM-102-LNPs were further modified using ß-sitosterol, a naturally-occurring cholesterol analogue associated with enhanced mRNA delivery31, to create a formulation referred to as LNP X (Fig. 1b). LNP X encapsulating reporter mCherry mRNA was formulated reproducibly and did not significantly differ in size, polydispersity or mRNA encapsulation efficiency compared to patisiran LNP (Fig. 1c and Table 1).
We assessed the potency of LNP X by treating Jurkat T cells, an immortalised T cell line, with increasing doses of reporter mRNA encapsulated in either patisiran LNP or LNP X. Treatment with LNP X compared to patisiran LNP resulted in up to 6-fold higher mCherry expression (Fig. 1d), confirming its superior potency. To assess the potency of our LNP X formulation further, we treated pre-stimulated CD4+ T cells with LNP X and observed mCherry expression in >75% of cells, in all doses tested, including a low dose of 6.25 ng mCherry-LNP X per 105 cells (Fig. 1e and Fig. S1a). Impressively, and in stark contrast to patisiran LNP, LNP X was able to achieve transfection of up to 76 ± 3.8% CD4+ T cells in the absence of pre-stimulation (Fig. 1a–e). Moreover, toxicity was negligible compared to baseline in both pre-stimulated and non-stimulated CD4+ T cells (Fig. S1c). To our knowledge, this is the first demonstration of potent transfection of primary, resting T cells in vitro in the absence of cellular toxicity.
LNP X was able to transfect both naïve and all memory T cell subsets (Fig. S2a–d), with effector memory T cells expressing higher levels of mCherry than naïve T cells (p = 0.007), potentially reflecting the higher basal translational activity in this memory T cell subset. Even in the context of peripheral mononuclear cells (PBMCs), LNP X was able to deliver mRNA to CD4+ T cells (Fig. S2e, f). Expression of mRNA was also detected in most other PBMC subsets, specifically monocytes, demonstrating that LNP X is T cell tropic, but not T cell specific.
Superior potency of LNP X is not related to endosomal escape
To understand the mechanism behind the superior potency of LNP X, we employed a SNAPswitch reporter system to quantify sub-cellular localisation of therapeutics in Jurkat T cells32 (Fig. 2a). SNAPswitch was recently used to probe the role that ionisable lipids play in delivering mRNA to the cytosol. This work showed that SM-102 had similar cytosolic delivery compared to DLin-MC3-DMA, the ionisable lipid found in patisiran LNP, however the protein expression induced by SM-102 was significantly higher33. To probe LNP X’s superior potency, we formulated SNAPswitch LNP X and compared these to LNPs formulated with SM-102, DSPC, DMG-PEG2000 and cholesterol. SNAPswitch LNP X did not exhibit different physicochemical characteristics compared to LNP X encapsulating a single reporter mRNA (Fig. S3). The LNP X formulation resulted in 10.5-fold enhanced mRNA expression in SNAPswitch Jurkat T cells (p = 0.0029, Fig. 2b). This enhanced transfection was in part explained by a 2.5-fold increase in LNP association, representing LNPs bound to the cell surface or internalised (p = 0.0025, Fig. 2c), yet LNP X still exhibited 4.1-fold higher protein expression relative to the amount of LNP association (p = 0.0032, Fig. 2d).
a The SNAPswitch assay simultaneously tracks the degree of nanoparticle association, endosomal escape and mRNA expression of LNPs co-encapsulating an AF488 oligo, Cy5-SNAPswitch oligo and mRNA encoding mScarlet. Figure adapted from Liu et al.33 and created in BioRender: Cevaal, P. (2025) https://BioRender.com/3ajxnam. b–g SNAPswitch-reporter Jurkat T cells were incubated with LNPs containing SM-102, DSPC, DMG-PEG2000 and either Cholesterol (Chol) or ß-sitosterol (Sito; LNP X) for 4 h. LNPs encapsulated a reporter mScarlet mRNA, AF488-tagged oligo and a Cy5-SNAPswitch oligo or constitutively active Cy5-oligo. Fluorescence was determined using flow cytometry. b Protein expression of the mScarlet reporter mRNA. c LNP association as determined by the fluorescence of the AF488-tagged oligo. d mScarlet protein expression (as in b) relative to LNP association (as in c). e Amount of cytosolic delivery of nucleic acid cargo as quantified by the Cy5-SNAPswitch oligo fluorescence. f Efficiency of endosomal escape based on the Cy5-SNAPswitch oligo fluorescence (as in e) normalised to a constitutively fluorescent Cy5-tagged oligo. g mScarlet protein expression (as in b) relative to the amount of cytosolic cargo delivery (as in e). b–g Bars represent aggregate mean of n = 3 independent experiments, each symbol representing the average of triplicate technical replicates. Significance was determined using a one-tailed paired ratio t test in (b–e,g) to allow comparisons of MFI values between experiments, or one-tailed unpaired t test in (f), ns non-significant.
While LNP X yielded an overall higher level of cytosolic cargo delivery (Fig. 2e), this effect correlated with the higher levels of LNP association to cells observed with LNP X. Indeed, the efficiency of endosomal escape was not improved by replacing cholesterol with ß-sitosterol (Fig. 2f). Importantly, the protein expression relative to the amount of mRNA delivered to the cytosol was enhanced by 5.3-fold when using LNP X (p = 0.0012, Fig. 2g), suggesting that the superior potency of LNP X involved processes downstream of endosomal escape and cytosolic delivery of the mRNA cargo. Our findings regarding the role of ß-sitosterol are consistent with previous studies comparing DLin-MC3-DMA/cholesterol to DLin-MC3-DMA/ß-sitosterol LNPs33, indicating the observed effects of ß-sitosterol are independent of the ionisable lipid.
LNP X delivers mRNA encoding HIV Tat to reverse HIV latency
We next aimed to assess whether LNP X was able to deliver a small, therapeutic mRNA expressing the 72 amino acids of the first coding exon of HIV Tat (Tat-LNP X, Fig. 3a and Table S1). This form of HIV Tat is normally expressed from the late-phase, Rev-dependent 4 kb viral Tat mRNA34. We first validated the potency of Tat-LNP X in J-Lat 10.6 cells, a T cell line model of HIV latency that expresses green fluorescent protein (GFP) under the control of the HIV LTR promoter. Treatment with Tat-LNP X, but not control mCherry-LNP X, resulted in an increase in GFP expression consistent with potent reactivation of LTR-mediated transcription (Fig. 3b). No impact on cellular viability was observed (Fig. S4). Similarly, Tat-LNP X was able to induce productive infection in primary CD4+ T cells infected with an HIV reporter virus (pMorpheus-V535; p = 0.0025) similar to levels seen after T cell activation with the mitogens phorbol 12-myristate 13-acetate (PMA) and ionomycin (Fig. S5).
a Schematic overview of the mechanism of action of HIV Tat on promoting transcription elongation. TAR; trans-activation response element. Created in BioRender: Cevaal, P. (2025) https://BioRender.com/cz40qwv. b J-Lat 10.6 cells were treated for 24 h with indicated doses of LNP X encapsulating codon-optimised mRNA expressing the 72 amino acids of the first coding exon of HIV Tat (Tat-LNP X) or mCherry (mCherry-LNP X) as control. Reactivation of HIV LTR-mediated transcription was determined after 24 h by measuring GFP expression. Highest dose of Tat-LNP X corresponds to 1 µg/mL. Treatment with PMA/ionomycin was included as a positive control. Mean ± SEM, n = 3 independent experiments. c–i CD4+ T cells from people living with HIV on suppressive ART were treated with 200 ng Tat-LNP X or mCherry-LNP X per 105 cells (4 µg/mL) or PMA/PHA as a positive control. After 48 h (squares) or 72 h (circles), expression of HIV transcripts TAR (c) LongLTR (d), Pol (e), PolyA (f) and Tat-Rev (g) representing transcription initiation, proximal elongation, distal elongation, completion and splicing, respectively, was determined using digital RT-PCR. Data were normalised to RNA input, then presented as fold-change induction compared to the corresponding non-treated (NT) control. Horizontal dashed line (c–g) represents no change relative to untreated cells. h Cellular toxicity was determined using flow cytometry. Where datapoints are missing in (h), cell input was insufficient to perform an accurate measurement. i After 72 h, the number of copies of HIV RNA per mL of supernatant was quantified using RT-PCR and normalised to the non-treated control. c–i Short horizontal line represents the median of n = 7–8 donors. Significance in (c–g and i) was determined using a one-tailed Wilcoxon signed-rank test, ns non-significant. After 120 h, the number of intact (j), 5’ defective (k) or 3’ defective (l) HIV DNA copies were determined using digital PCR. Copy numbers were normalised to cell input. Datapoints represent individual donors (n = 7). Note different donors were used in (j–l and c–i). Significance was determined using a two-tailed Wilcoxon signed-rank test, ns non-significant.
To assess latency reversal ex vivo, we treated CD4+ T cells from people living with HIV on suppressive ART with an equivalent dose of 200 ng per 105 cells Tat-LNP X or control mCherry-LNP X for 48–72 h. The expression of cell-associated HIV RNA was measured as described previously36,37, whereby we simultaneously quantified transcription initiation, elongation, completion and splicing37. After a single dose of Tat-LNP X, all measured HIV transcripts were significantly upregulated compared to non-treated control cells suggesting Tat-LNP X induces, and overcomes blocks in, transcription initiation (TAR transcript), proximal and distal elongation (Long-LTR and Pol transcripts, respectively), completion of transcription (Poly(A) transcripts) and splicing (Tat-Rev transcripts; detected by a primer/probe set that spans the junction between the first and second coding exon of tat and rev) (Fig. 3c–g and Fig. S6a–e). The induction of multiply-spliced Tat-Rev transcripts, an important predictor of virion production ex vivo38, was induced by 112-fold (p < 0.01 compared to untreated control). Treatment with Tat-LNP X or control mCherry-LNP X resulted in minimal overall toxicity (Fig. 3h). Importantly, the induction of HIV RNA transcription following Tat-LNP X significantly exceeded that observed in fully activated T cells stimulated using two mitogens, PMA and phytohaemagglutinin (PHA) (considered the gold-standard to reverse HIV latency in vitro and ex vivo37,38) (p < 0.01 for all transcripts). However, in contrast to PMA/PHA treatment, Tat-LNP X did not induce any change in cellular activation as observed by minimal changes to the expression of activation markers CD25, CD69 and HLA-DR compared to non-treated control cells (Fig. S6f–h). Furthermore, treatment with Tat-LNP X, but not mCherry-LNP X, resulted in a 17.2-fold increase in supernatant HIV RNA, indicating virion production following latency reversal (Fig. 3i). However, the reversal of latency by Tat-LNP X did not result in a decline in intact proviral DNA after 5-day culture (Fig. 3j–l). Combined, these findings demonstrate that Tat-LNP X is a highly potent latency-reversing agent that can overcome HIV RNA transcription and processing blocks to elongation, splicing and completion, and induce viral protein expression, all in the absence of T cell activation.
CRISPR activation machinery can be co-encapsulated by LNP X
We next assessed whether LNP X was able to encapsulate and deliver CRISPR activation (CRISPRa) machinery as a more complex and larger RNA-based, highly HIV-specific therapeutic. The dCas9-synergistic activation mediator (dCas9-SAM) CRISPRa system consists of a catalytically inactive (dead) Cas9 (dCas9) fused to a multimer of C-terminal herpes virus transcriptional activation domain 16 (VP64), which is guided to the genomic target site by a guide RNA (gRNA). The gRNA is modified to contain minimal hairpin aptamers that allow the recruitment of introduced p65 and heat shock factor 1 (HSF1) transcription activators through binding of the bacteriophage protein MS239 (Fig. 4a). The recruitment of multiple copies of the MS2-p65-HSF1 fusion protein, which synergise with dCas9-VP64, greatly enhances the potency of CRISPRa39, but can result in non-specific promoter activation if present in excess. We therefore first optimised the relative dose of the three RNAs that comprise the dCas9-SAM CRISPRa machinery. We found that a relative mass ratio of 0.8: 0.00625: 1 (dCas9-VP64 mRNA: MS2-p65-HSF1 mRNA: gRNA) yielded the optimal balance between on-target potency and non-specific background reactivation in cell lines (Fig. S7).
a Schematic overview of the dCas9-synergistic activation mediator (SAM) CRISPR activation system, consisting of a catalytically inactive (dead, d)Cas9 fused to transcriptional activator domain VP64, a gRNA and a further transcriptional activation MS2-p65-HSF1 fusion protein that is recruited to the Cas9-gRNA complex via MS2-binding to stem-loop structures in the gRNA scaffold. Created in BioRender: Cevaal, P. (2025) https://BioRender.com/p8j24y6. b–d LNP formulation X was used to encapsulate the CRISPR activation machinery (CRISPRa-LNP X) including a gRNA targeting the endogenous gene encoding CD25 (CD25 CRISPRa-LNP X) or a scrambled control gRNA (scr CRISPRa-LNP X). b Jurkat T cells were treated for 24 h with CRISPRa-LNP X, control mCherry-LNP X or PMA/ionomycin as positive control. Induction of CD25 expression was measured by surface stain using flow cytometry. CD4+ T cells from HIV-negative donors were treated with indicated doses of CRISPRa-LNP X per 105 cells or mCherry-LNP X as a control for 72 h (c) or 6 days (d), after which CD25 expression was determined using flow cytometry. Highest dose of CD25 CRISPRa-LNP X corresponds to 4 µg/mL. Treatment with PMA/ionomycin was included as a positive control. Dotted line represented average baseline CD25 expression in the absence of treatment. Mean ± SEM, n = 4 independent experiments (b) or n = 4–6 donors (c, d). Significance was determined using a two-tailed Student’s t-test.
We next developed a model system in which CRISPRa was targeted to the promoter of an endogenous gene, IL2RA (cd25), to induce overexpression of the CD25 receptor on the cell surface. CRISPRa-LNP X was formulated by co-encapsulating the two mRNAs and a CD25-targeting or a scrambled control gRNA. The CD25-targeting gRNA was validated in Jurkat T cells, which express negligible CD25 at baseline. We found that 24 h following treatment with CD25 CRISPRa-LNP X, there was an increase in surface CD25 expression in up to 29.6 ± 8.6% of Jurkat T cells, while minimal to no background expression of CD25 was observed with scrambled gRNA (scr) CRISPRa-LNP X (Fig. 4b and Fig. S8a). We demonstrated a similar level of potency in non-stimulated primary CD4+ T cells from HIV-negative donors, yielding a significant increase in cells expressing CD25 from 8.7 ± 0.7% with scr CRISPRa-LNP X to 14.6 ± 0.9% after 72 h treatment with 800 ng CD25 CRISPRa-LNP X per 105 cells (p < 0.01) (Fig. 4c and Fig. S8b).
We then extended the incubation time after a single dose of CD25 CRISPRa-LNP X, scr CRISPRa-LNP X or control mCherry-LNP X from 72 h to 6 days. The percentage of cells expressing CD25 after 6 days was increased to 24.6 ± 5.1% of live cells (Fig. 4d and Fig. S8c), suggesting the peak response of CRISPRa-induced protein expression occurs later than 72 h post-treatment. In contrast, the expression of mCherry was reduced at 6 days compared to 72 h post-treatment (Fig. S8d, e), consistent with transient expression of LNP-mediated delivery of exogenous mRNA. The enhanced CD25 expression at day 6 was therefore likely not the result of increased expression of the CRISPRa machinery (dCas9-VP64 and MS2-p65-HSF1) at day 6, rather the extended incubation is required to capture the translation of the induced CD25 mRNA and its trafficking to the cell surface. Importantly, these findings confirm, the first to our knowledge, successful delivery of the dCas9-SAM CRISPRa machinery to non-stimulated T cells in vitro.
HIV CRISPRa-LNP X activates HIV transcription ex vivo
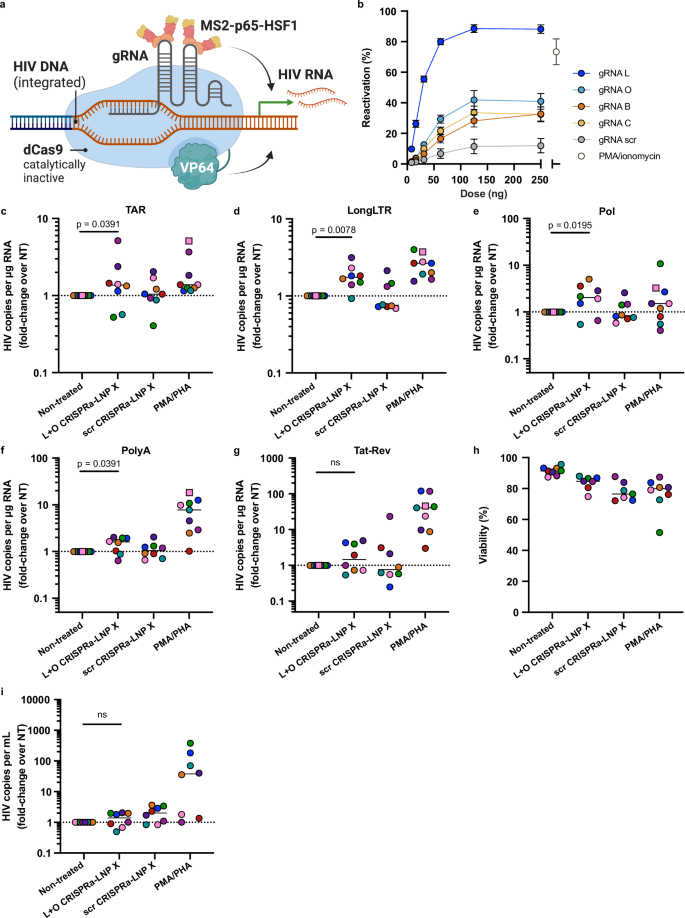
Finally, to generate HIV LTR-targeted CRISPRa-LNP X, we designed four MS2-p65-HSF1 recruiting hairpin-modified gRNAs (B, C, L and O) targeting different HIV LTR regions upstream of the transcription start site based on previous work23,40 (Fig. 5a, Table S1 and S2). We found that all four HIV LTR-targeting gRNAs, compared to a scrambled gRNA, could induce GFP expression in J-Lat 10.6 cells (Fig. 5b). To assess the potency of HIV LTR-targeting CRISPRa-LNP X ex vivo, we treated CD4+ T cells from ART-suppressed people living with HIV for 72 h with an equivalent dose of 200 ng CRISPRa-LNP X per 105 cells, analogous to what we used to deliver Tat mRNA. We used CRISPRa-LNP X co-encapsulating a 1:1 mass ratio of gRNA L and gRNA O (L + O CRISPRa-LNP X), which was found to exceed the potency of either gRNA alone (Fig. S9) and was hypothesised to increase the sequence coverage of HIV subtype B LTR sequences, the most common subtype in our cohort. L + O CRISPRa-LNP X treatment significantly increased all measured HIV transcripts except multiply-spliced Tat-Rev (Fig. 5c-g and Fig. S10a–e). The potency of latency-reversal was lower compared to Tat-LNP X treatment, with a maximum fold-increase over non-treated of 2.0 (IQR 0.87-3.39) for Pol transcripts, and did not exceed that of PMA/PHA treatment. No HIV transcripts were upregulated after treatment with a scrambled gRNA CRISPRa-LNP X, indicating the activation of transcription was gRNA-mediated. Treatment with HIV LTR-targeted CRISPRa-LNP X resulted in minimal toxicity, similar to treatment with Tat- or mCherry-LNP X, and did not result in generalised cellular activation (Fig. 5h and Fig. S10f–h), demonstrating the specificity of CRISPRa-LNP X as a next-generation LRA. However, treatment with L + O CRISPRa-LNP X did not lead to an induction of supernatant HIV RNA, as a measure of virion release (Figs. 5i and S10i).
a Schematic overview of the CRISPRa system targeting the HIV LTR. Created in BioRender: Cevaal, P. (2025) https://BioRender.com/p8j24y6. b J-Lat 10.6 cells were treated for 24 h with indicated doses of CRISPRa-LNP X containing one of four HIV LTR-targeting gRNAs (L, O, B, C) or scrambled control gRNA. Reactivation of HIV LTR-mediated transcription was determined after 24 h by measuring GFP expression. Highest dose of CRISPRa-LNP X corresponds to 2.5 µg/mL. Treatment with PMA/ionomycin was included as a positive control. Mean ± SEM, n = 3 independent experiments. c–i CD4+ T cells from people living with HIV on suppressive ART were treated with 200 ng CRISPRa-LNP X containing gRNAs L and O (L + O CRISPRa-LNP X) or a scrambled gRNA control (scr CRISPRa-LNP X) per 105 cells (4 µg/mL) or PMA/PHA as a positive control. After 48 h (squares) or 72 h (circles), expression of HIV transcripts TAR (c) LongLTR (d), Pol (e), PolyA (f) and Tat-Rev (g) representing transcription initiation, proximal elongation, distal elongation, completion and splicing, respectively, was determined using digital RT-PCR. Data were normalised to RNA input, then presented as fold-change induction compared to the corresponding non-treated (NT) control. Horizontal dashed line (c–g) represents no change relative to untreated cells. h Concurrently, cellular toxicity was determined using flow cytometry. Where datapoints are missing in (h), cell input was insufficient to perform an accurate measurement. i After 72 h, the number of copies of HIV RNA per mL of supernatant was quantified using RT-PCR and normalised to the non-treated control. c–i Short horizontal line represents the median of n = 7–8 donors. Significance in (c–g) and (i) was determined using a one-tailed Wilcoxon signed-rank test, ns non-significant.