Influenza epidemics consist of distinct co-circulating transmission lineages
First, we characterized the transmission lineage structure of US seasonal influenza epidemics, to investigate whether they tend to comprise many distinct co-circulating transmission lineages that independently emerged in different states, or rather consist of a single dominant transmission lineage that propagates across the country. We analyzed 30,508 whole-genome seasonal influenza virus sequences from the 48 contiguous states and the District of Columbia, collected during routine surveillance from 2014 to 2023. In this period, all four influenza A subtypes and influenza B lineages (henceforth, subtypes) caused epidemic activity, but patterns of subtype dominance differed from season to season (Supplementary Fig. 1). We phylogenetically grouped these viruses into clusters that exhibit a comb-like branching structure, suggestive of exponential spread33. Given the exponential nature of influenza epidemics, we posit that groups of viruses with such a rapidly expanding branching structure plausibly descended from a single ancestral virus in the United States that was introduced from abroad (Fig. 1a, Supplementary Fig. 2–5).
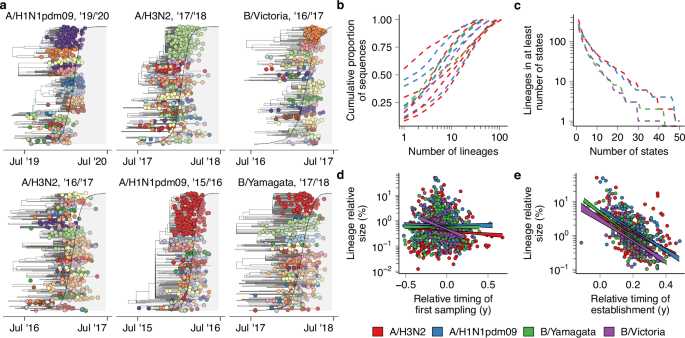
a Example phylogenetic trees with tips colored by transmission lineage. The shaded gray area corresponds to the cumulative proportion of the season’s nationwide positive tests in public health laboratories of the corresponding subtype that had been collected at each point in time. b Lineage size distribution by season and subtype. Each line represents the cumulative proportion of sequences that is accounted for by a number of lineages on the x-axis. c The number of subtype-specific lineages that accounted for >5% of sequences in a season-subtype in at least the number of states on the x-axis. d Relationship between the first collection date of virus in a lineage and the lineage’s nationwide size normalized by state (n = 1030). Lineage sampling dates were computed relative to the timing of nationwide epidemic onset, which was defined as the first week in which >5% of the season’s cumulative positive tests had been collected. Lines correspond to 50% CI given linear fit. e Relationship between the timing of establishment of substantial circulation of a lineage and its nationwide size (n = 569). Lineage establishment timing was computed relative to nationwide epidemic onset analogous to (d).
Using this procedure, we clustered 81.2% of sequences into 3842 lineages of at least two viruses. In most seasons, a relatively small number of transmission lineages accounted for the bulk of sequenced viruses (Fig. 1b), with median 5 lineages (range 1–13) cumulatively accounting for ≥50% of sequences across subtypes and seasons. Transmission lineage diversity differed substantially across seasons (Fig. 1b): in the 2015/2016 A/H1N1pdm09 epidemic, a single transmission lineage accounted for >50% of sequences, normalized across states, whereas in the 2016/2017 A/H3N2 season the largest lineage accounted for only 6.4%. Lineage structure was present for both circulating influenza A virus subtypes and both influenza B virus lineages, though transmission lineage clustering results are likely more error-prone for influenza B viruses given their lower evolutionary rate34, particularly in seasons that saw relatively little circulation and were less densely sampled. We found no relationship between transmission lineage diversity and metrics of epidemic timing (Pearson r = −0.16, P = 0.63) or intensity (Pearson r = −0.12, P = 0.73). Only a small proportion of lineages spread widely across the country (Fig. 1c): among the 1,104 identified transmission lineages that accounted for at least 5% of sequences in a season in at least one state, 144 (13.0%) lineages did so in at least 10 states, and 27 (2.4%) did so in at least 25. These results indicate that in most seasons, seasonal influenza epidemics are the result of the co-circulation of multiple independent chains of transmission, consistent with previous studies into individual seasons at the national and state level28,29. These conclusions are robust to the choice of parameters for the procedure used to cluster viruses into transmission lineages (Supplementary Fig. 6).
Lineage size correlates with timing of establishment of epidemic activity
We hypothesized that the first lineages to emerge in any season, for a given subtype, would be larger. Here, we defined lineage size as the proportion of sequences of the corresponding subtype that a lineage accounts for in a season across all states, where each state has an equal weight. However, across all subtypes and seasons, the relationship between time of first sampling of a lineage and (log) lineage size was weak (Spearman ρ = −0.08, P = 0.012) (Fig. 1d). Some transmission lineages that were first sampled a prolonged interval prior to onset of the national epidemic proliferated into peak periods (Fig. 1d). However, many of the most successful lineages were first sampled close in time to national epidemic onset (Fig. 1d). For example, by the time of first sampling of the largest lineage in the highly severe35 2017/2018 A/H3N2 season, >10% of all the season’s sequences had already been collected, but the lineage accounted for >40% of sequences around the epidemic peak following rapid expansion (Fig. 1a). The fact that a transmission lineage could rapidly sweep to national dominance despite emerging at a time when many other transmission lineages were already circulating suggests that influenza epidemic dynamics are strongly influenced by heterogeneous short-term epidemiological processes.
These results suggest that early-season transmission chains frequently go extinct before the onset of substantial epidemic activity. However, we hypothesized that if a lineage did cause substantial epidemic activity early on, it would likely be successful nationwide. We defined the timing of lineage establishment as the first week in which the lineage accounted for substantial epidemic activity in at least one state. When defining lineage establishment week as the first week a lineage had cumulatively accounted for >5% of the state’s total epidemic activity of that subtype, establishment week correlated strongly with nationwide lineage size for all subtypes (Spearman ρ = −0.46, −0.50, −0.70, −0.71 for A/H3N2, A/H1N1pdm09, B/Yam, B/Vic, respectively, P < 0.001 for all) (Fig. 1e). Statistical models indicated each week’s delay in establishment resulted in a decrease in lineage size of 16.9%, 15.1%, 14.6%, and 13.0% for A/H3N2, A/H1N1pdm09, B/Yam, and B/Vic, respectively (P < 0.001 for all). The fact that the lineages that first established substantial epidemic activity somewhere in the US were more likely to be successful suggests that the states with the earliest epidemic onset have outsized contributions to which viruses will circulate nationwide. The proportion of the earliest-establishing transmission chains that resulted in a successful transmission lineage decreased as the cumulative incidence threshold used to determine establishment week decreased (Supplementary Fig. 7). This suggests that while a first-mover advantage shapes lineage success, substantial levels of epidemic activity are necessary for a lineage’s early circulation to be predictive of substantial nationwide spread.
Transmission lineages are highly spatially structured
To quantify spatial structure in the circulation of transmission lineages, we computed the Bray-Curtis dissimilarity of transmission lineage compositions across states. Here, lower dissimilarity is indicative of more frequent circulation of the same lineages. Qualitatively, hierarchical clustering of the resulting similarity matrix recapitulated the geography of the United States, with relatively higher similarity for states within the same census region (Fig. 2a, Supplementary Fig. 8), suggestive of substantial spatial structure. Projecting the similarities among states onto a two-dimensional surface further recapitulated this spatial structure; for example, states belonging to the Northeast and Southeast appeared to form distinct clusters (Fig. 2b). However, the continuous distribution of states on the plane suggests states cannot consistently be classified into distinct communities, suggestive of substantial inter-regional mixing.
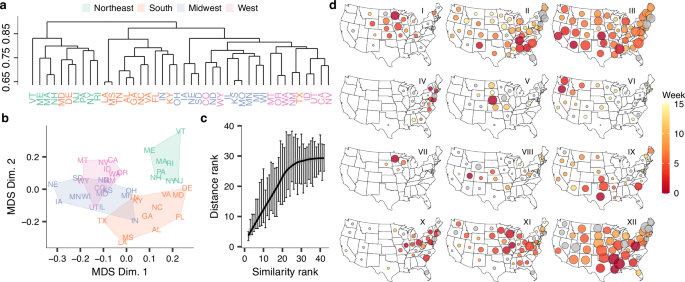
a Complete-linkage hierarchical clustering of pairwise state-to-state transmission lineage composition Bray-Curtis similarities across all subtypes, colored by census region. b Multi-dimensional scaling plot of the pairwise lineage composition Bray-Curtis similarity among states, colored by census region. c Relationship between pairwise transmission lineage compositional similarity rank and pairwise centroid distance rank (n = 42). Vertical lines show 50% CI of distance rank for each value of rank similarity; line corresponds to LOESS fit to medians. d Examples of lineage spatiotemporal spread for a geographically and temporally representative set of lineages. In each map, circle size and color correspond to the relative size and establishment timing of the corresponding lineage in each state, respectively. Gray fill corresponds to unknown lineage establishment timing.
Across all seasons and subtypes, epidemics in states in closer geographic proximity more frequently comprised the same transmission lineages (Mantel test, P < 0.001) (Fig. 2c), with the highest similarities found for adjoining states (highest: MS-LA, MO-KS, GA-AL, NH-MA, UT-ID). Stratifying by season and subtype, this correlation between distance and similarity was present in most, but not all, subtypes that accounted for >20% of nationwide positive tests in their respective season (Supplementary Fig. 9). Visualizing lineages’ spatial distribution and proliferation across states revealed a striking landscape of spatiotemporal spread. Individual lineages emerged from all regions of the US, each with distinct spatial signatures (Fig. 2d). Across seasons and subtypes, many lineages exhibited a radial spatiotemporal progression and were highly regional (e.g. I, V, VI, VII, Fig. 2d), whereas other lineages saw rapid cross-country spread (e.g. II, III, XII, Fig. 2d). Together, these results show that at the transmission lineage level, US seasonal influenza epidemics are highly spatially structured.
Consistent source-sink dynamics are absent across seasons and subtypes
Next, we reconstructed where in the United States successful transmission lineages initially established. US source-sink dynamics have been the subject of debate, with studies suggesting that the South represents a dominant source region16,30. Using BEAST36, we identified the Health & Human Services (HHS) region that represented the most likely region of initial expansion for each of the 262 transmission lineages that accounted for >0.5% of sampled viruses, normalized by state, in their respective season. To ensure our results were robust to sequence sampling effects, we performed these analyses with a population-weighted subsampling strategy and a subsampling strategy with a uniform number of sequences across regions. We found that the origin region of successful lineages differed substantially from season to season. Of the seven transmission lineages that accounted for >10% of sequences in a single season, normalized across states, three likely first established in the southern US (HHS regions 4 and 6; e.g., lineages II and XII, Fig. 2d), one likely emerged in the western US (HHS region 9; lineage III, Fig. 2d), one in the midwestern US (HHS regions 7 and 5; lineage XI, Fig. 2d), one in the northeastern US (HHS region 1), and one could not consistently be attributed to a single region across both sampling strategies.
To investigate if the dominant origin region of sampled viruses differed among states, we computed origin profiles that quantify, for each state, what proportion of viruses belonged to transmission lineages that initially expanded in each of the HHS regions. These profiles differed substantially across states (Supplementary Figs. 10, 11). For example, averaged across both subsampling strategies, the proportion of sequences that corresponded to transmission lineages initially expanding in HHS region 4, encompassing most of the southeastern US, ranged from 39.3% in South Carolina (HHS region 4) and 27.4% in Arkansas (HHS region 6) to 13.0% in Arizona (HHS region 8). Similarly, lineages initially expanding in HHS region 10, encompassing the Pacific Northwest, accounted for 15.3% of sampled viruses in Idaho (HHS region 10), 10.7% in North Dakota (HHS region 8), and only 2.7% in Arkansas (HHS region 6). States in closer geographic proximity saw more similar origin profiles, even if they corresponded to different HHS regions (Mantel test, P < 0.001). Across all states, a relatively limited proportion of viruses corresponded to lineages that originated from the state’s own HHS region (median 17.3%, range 11.4–29.8% for uniform subsampling strategy), suggesting a high degree of viral mixing at the national level. Importantly, origin profiles were strongly correlated across the two subsampling strategies (Spearman ρ = 0.81, P < 0.001). Together, these results suggest that influenza virus source-sink dynamics are highly heterogeneous, without consistent source regions of successful lineages, but are spatially structured.
Mechanistic simulations suggest commuting flows correlate with viral spread
Our analyses established a strong correlation between timing of lineage establishment and lineage size (Fig. 1e). However, this correlation does not account for a substantial portion of the observed variation in transmission lineage size. We hypothesized that differences among states in their connectedness through human mobility, coupled to competition between lineages for susceptible hosts, could account for further variation. In this hypothesis, lineages that established in more connected states have a greater potential to rapidly spread to other states, thereby outcompeting lineages that emerged around the same time in less connected states. This would explain why some lineages spread widely following local establishment, whereas other lineages remained highly spatially constrained (Fig. 2d).
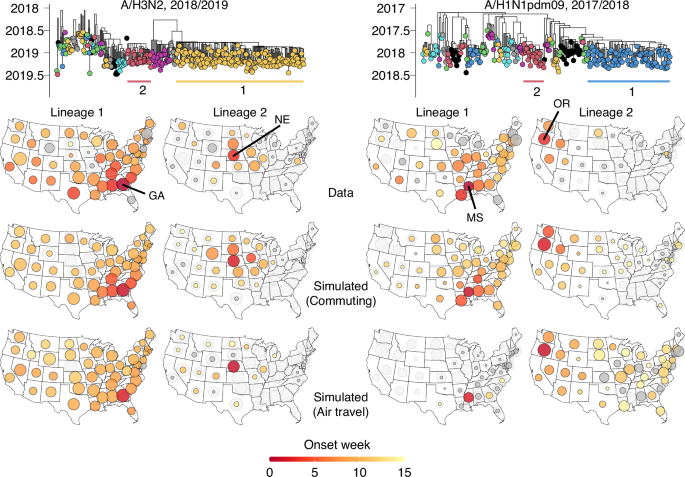
This hypothesis appears to explain lineage dynamics in the 2018/2019 A/H3N2 season. The beginning of this season was dominated by A/H1N1pdm09 viruses, but it also saw the rapid expansion of A/H3N2 viruses that were associated with reduced vaccine effectiveness37. Phylogenetic analyses, integrated with epidemiological data, indicate that A/H3N2 circulation in this season was dominated by two swiftly establishing lineages, appearing to emerge from Georgia (GA, lineage 1) and Nebraska (NE, lineage 2), respectively (Fig. 3, left). Spread of the Nebraskan lineage was regional and radial, causing substantial epidemic activity mainly in the immediately surrounding states (Fig. 3, left). Conversely, the lineage from Georgia quickly spread to almost all states with a less prominent temporal hierarchy, although it appeared to arrive in neighboring states first. We hypothesized that competition from the Georgian lineage explained why spread of the Nebraskan lineage remained so regional. In turn, this would explain why the Georgian lineage failed to spread substantially in Nebraska and immediately surrounding states. We used the reconstructed spread of these lineages as a ground truth to address long-standing questions regarding the underlying mobility determinants of influenza virus spread.
Phylogenies represent the 2018/2019 A/H3N2 season and 2017/2018 A/H1N1pdm09 season, with the two largest lineages labeled in each. For both seasons, the maps in the top row visualize the reconstructed spread of the labeled lineages, with size and color corresponding to lineage size and establishment timing, respectively. Middle and bottom maps show simulated spread of the two lineages for each of the two seasons, using commuting and air travel data, respectively. Light gray circles represent the total proportion of sequences in a state that was accounted for by simulated lineages. In each state, circle sizes for simulated lineages are scaled such that the sum of simulated lineages’ sizes is proportional to the proportion of sequences accounted for by the simulated lineages (i.e., the light gray area). Dark gray fill corresponds to absence of an establishment week (for top row, potentially due to missing data), or establishment after the 15th week.
To test the hypothesis that competitive interactions between lineages, coupled to mobility, drive lineage spread, we explored if we could reproduce the spread of these two lineages with mechanistic epidemic simulations. We used a metapopulation model to simulate the inter-state spread of multiple co-circulating lineages that compete for disease-susceptible hosts with perfect cross-immunity in a susceptible-infected-recovered (SIR) epidemic framework. We initialized the simulations in the lineages’ respective establishment weeks, in their respective onset states (Georgia and Nebraska) and simulated forward in time to model the spread of the two lineages. To ascertain the predominant mobility drivers of viral spread, we parameterized rates of state-to-state mobility using either commuting flows, extracted from the US Census Bureau, or air travel data, extracted from the US Department of Transportation.
When using commuting flows to parameterize rates of inter-state travel, we could reproduce the observed spread of the two lineages with striking accuracy: the simulations recapitulated the radial spread from Nebraska, and the relative success of the lineage emerging from Georgia (Fig. 3, left). The model of inter-lineage competition also explains why the lineage from Georgia failed to cause epidemic activity in Nebraska and the immediately surrounding states. The correspondence to the observed distribution of lineages across states was much poorer (Δℓ = −81.93) when using air travel flows, with an absence of substantial spread from Nebraska. Furthermore, simulated lineage establishment timings better matched observed establishment timings when using commuting data (MSE = 6.0 wk) than when using air travel data (MSE = 11.7 wk) Together, these mechanistic simulations suggest that commuting flows are a strong correlate of viral spread and suggest that competitive interactions between lineages, mediated by mobility flows, shape the distribution of lineages across states.
Spatial segregation and limited competition allow lineages from small states to spread widely
Lineage dynamics in the 2018/2019 A/H3N2 season are consistent with the conjectured gravity-like spread of seasonal influenza viruses12, with a lineage originating from a populous, highly connected state (in this case, Georgia, population ~11 million) spreading quickly through strong long-range connections, while spread from a smaller state (Nebraska, population ~2 million) was slower and more local. In this model, Georgia’s high degree of connectivity and earlier onset allowed lineage 1 to spread to other states more rapidly than lineage 2. Nevertheless, the substantial spread of the Nebraskan lineage shows that a lineage emerging from a small state can proliferate, even if it co-circulates with a lineage emerging from a more populous state, if it sees sufficiently early establishment and is spatially segregated from the lineage emerging from the larger state.
Our results suggest that by facilitating spread from less populous states, the short-range spatial coupling reflected in commuting flows is a key determinant of seasonal influenza virus spread. This notion is further supported in the 2017/2018 A/H1N1pdm09 season, in which the two largest lineages appeared to emerge in Mississippi (MS) and Oregon (OR), respectively (Fig. 3, right). When using commuting flows, the relative degree of spread of the two lineages could be reproduced. Despite its relatively small population, Mississippi’s high connectivity through commuting flows allowed lineage 1 to rapidly spread beyond local constraints. In contrast, due to Oregon’s relatively limited connectivity and the later establishment of lineage 2, competition from lineage 1 likely constrained the spread of those viruses to the Western United States. When using only air travel to parameterize inter-state mobility, the simulations overestimated the degree of spread from Oregon relative to Mississippi, with too slow spread from Mississippi, compared to the ground truth, resulting in poorer fit (Δℓ = −1.14 for observed sample counts, ΔMSE = 5.5 for observed establishment timings) (Fig. 3, right).
Using counterfactual simulations, we explored how mobility interacts with establishment timing to shape the spread of individual lineages. Simulations indicate that had lineage 2 established in Nebraska four weeks later (with lineage 1’s establishment timing unchanged), stronger competition would have constrained lineage 2 to Nebraska and immediately adjacent states. Conversely, if it had established four weeks sooner, lineage 2 would have been approximately equal in size to lineage 1, with earlier onset compensating for lower connectivity (Supplementary Fig. 12a). Similarly, later onset for lineage 2 in the 2017/2018 A/H1N1pdm09 season would have constrained it to the Pacific Northwest, whereas earlier onset would have facilitated substantially more expansive spread (Supplementary Fig. 12b).
Inter-lineage competition explains differences in lineages’ spread
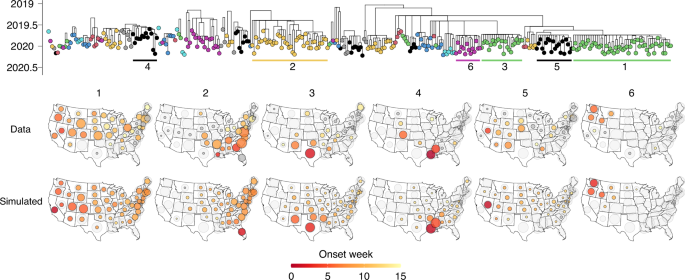
To further test the hypothesis that competition between lineages on the human mobility network explains individual lineages’ spread, we performed in-depth investigations into the 2019/2020 B/Victoria season, which was characterized by anomalously high levels of epidemic activity38. The lineage composition of this season was highly spatially diverse, with the largest lineages appearing to originate in or in the vicinity of California (lineage 1), Florida (lineage 2), Texas (lineage 3), Louisiana (lineage 4), Nevada (lineage 5), and Washington (lineage 6), respectively (Fig. 4). Lineage dynamics in this season highlight the heterogeneity of processes of lineage establishment. For example, the lineage emerging from Florida likely emerged in the spring of 2019 (posterior mean TMRCA May 5, 95% CrI March 14–June 12), seemingly persisting throughout the 2019 summer in Florida, potentially providing a counterexample to the general trend that viruses do not persist between seasons28. Conversely, the lineages from California, Nevada, and Texas spread widely following rapid establishment, despite much later emergence (e.g., lineage 1: posterior mean TMRCA August 29, 95% CrI July 3–September 25, Fig. 4). Some lineages spread to over half of all states (e.g., lineages 1 and 2 from Florida and California, respectively), whereas other lineages were more spatially constrained (Fig. 4).
Phylogeny represents the 2019/2020 B/Victoria season, with the six largest lineages labeled in order of size. Top row of maps represents the reconstructed spread and distribution of each of the six largest lineages. Bottom row of maps represents the simulated spread and distribution of the six lineages, initialized in the lineages’ likely onset state and onset week, using a combination of air travel data and commuting data. Circle sizes are scaled as in Fig. 3.
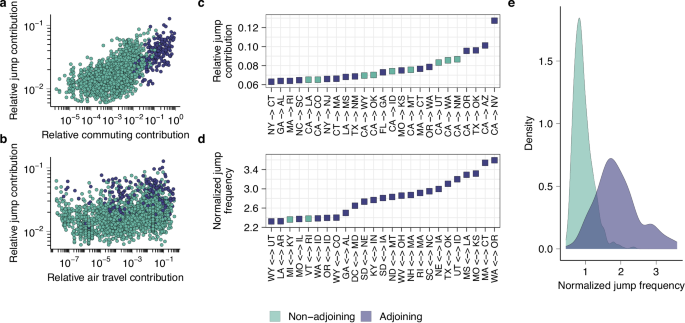
a Relationship between the relative contribution of each other state to a state’s inbound and outbound reconstructed viral migration events, and the other state’s relative role as a state’s commuting destination (n = 2352). b Analogous to (a), for air travel data (n = 2352). c Visualization of the 25 highest values of the relative jump contribution. d Visualization of the 25 highest values for the normalized pairwise jump frequency. e The distribution of normalized pairwise migration frequencies for pairs of adjoining and non-adjoining states.
Using a combination of commuting flows and air travel flows, the simulations reproduced the spread of individual lineages and their distribution across states (Fig. 4). Differences in mobility flows, in combination with competition for susceptible hosts, parsimoniously explain why the lineages emerging from California and Florida spread widely, whereas the lineages from Louisiana and Washington were more spatially constrained. Commuting flows in isolation also yielded a strong fit, but underestimated spread from Nevada, suggesting that residual air travel flows not captured by commuting could play a role in viral dissemination (Supplementary Fig. 13). Simulations using only air travel deviated from the ground truth particularly by underestimating short-range viral migration from Louisiana and Washington, and provided a worse fit to the observed distribution of lineages across states (ℓ = −347.7, −393.2, −400.0 for fit to lineage sample counts for combined, commuting, and air travel models, respectively) and the relative timing of lineage establishment in different states (MSE = 7.6, 7.9, 16.6, respectively, for the three models) (Supplementary Fig. 13).
Our metapopulation simulations demonstrate a strong model fit while directly leveraging data on mobility flows without fitting any mobility-related parameters, indicating these mobility metrics parsimoniously explain observed dynamics. To derive a more general quantification of the gravity-like nature of viral spread, we fit a model where the propensity pij for an individual from state j to travel to state i (conditional on traveling) depends on state i’s population size Ni and the distance between states dij as \({p}_{{ij}}\propto {N}_{i}^{\tau }/{d}_{{ij}}^{\rho }\). In this model, the daily probability for an individual from a given state to travel was informed by the commuting and air travel data. This general model accurately recapitulated observed lineage dynamics, with a strong fit to sample counts (ℓ = −329.7), but with moderately reduced fit to establishment timing (MSE = 9.5) (Supplementary Fig. 13). We estimated a rapid decay with distance (ρ = 1.4, 95% CI 1.2–1.5) with a statistically significant dependence on destination population size (τ = 0.5, 95% CI 0.3 − 0.8) (Supplementary Fig. 14). However, if we assumed the daily probability of traveling was constant across states, the model yielded a substantially worse fit than the models directly informed by commuting and air travel data (ℓ = −429.6, MSE = 11.2). This indicates that accounting for differences among states in outward travel rates is necessary to capture salient features of influenza virus spread.
Viral genomes suggest viral migration correlates with commuting
The mechanistic simulations suggest that commuting flows are the primary mobility drivers of influenza virus spread. However, these analyses could only be performed for the seasons with relatively low lineage diversity, as the large number of co-circulating lineages in some seasons rendered sample counts too low for individual lineages to yield a reliable ground truth for reconstructions of spread. We sought to confirm that commuting is the predominant correlate of viral spread across the full set transmission lineages in the study period. Hence, we probed whether the same mobility processes were reflected in the genetic relationships among viruses belonging to the same transmission lineage.
Using phylogeographic methods, we reconstructed the migration history of the sampled viruses in each transmission lineage, tracing the state-to-state jumps between the lineage root and each virus. Using these reconstructions, we quantified the relative viral jump contribution x → y, representing the proportion of reconstructed migration events to and from state y that was accounted for by state x. We found that the states with the greatest jump contribution to another state tended to be the states that were most strongly connected through commuting flows to that state (Spearman ρ = 0.63, P < 0.001) (Fig. 5a). The correlation with air travel was weaker (Spearman ρ = 0.32, P < 0.001) (Fig. 5b), as was the correlation with state centroid distance (Spearman ρ = −0.39, P < 0.001). This provides further support for the notion that commuting is a strong correlate of the mobility processes that disseminate seasonal influenza viruses across the United States.
The highest values of the relative jump contribution x → y were found when state x was highly populous and state y was in close geographic proximity to state x. For example, the highest values across all pairs were found for CA → NV, CA → AZ, TX → OK, CA → OR, and CA → NM (Fig. 5c). This is expected under classical gravity-like spread where, for any given state, the highest connectivity is expected to be to states that are in close geographic proximity and highly populous. However, this pattern could be confounded by higher sample counts for the most populous states, as higher sample counts in a spatial grouping in phylogeographic analyses will a priori be expected to lead to more reconstructed migration events even in the absence of any spatial signal in the data. Hence, we computed an alternative metric, the normalized pairwise jump frequency x ↔ y, which represents the proportion of migration events to/from state y that is accounted for by state x, normalized relative to the mean proportion of migration events that state x accounts for across all states. The highest values were for adjacent states that are strongly connected through commuting flows (highest: WA ↔ OR, MA ↔ CT, MO ↔ KS, MS ↔ LA, UT ↔ ID) (Fig. 5d), and this value was substantially greater for adjacent than non-adjacent states (Fig. 5e). This result suggests that when accounting for effects of population size and/or sampling, viral migration is strongly skewed toward short distances, further supporting the important role of short-range spatial coupling. To identify differences among subtypes in the degree of distance-dependence of viral spread, we computed the correlation between centroid distance and pairwise jump contribution for each season and subtype individually. While we established differences among subtypes and seasons, these differences were not consistently associated with subtype (Supplementary Fig. 15).