Clinical categorization of infected infants and changes in PBMC compositions
We collected blood samples and processed serum and peripheral blood mononuclear cells (PBMCs) from 26 infants (median, IQR age: 1.63 [0.93–7.59] months) hospitalized with SARS-CoV-2 infection (pediatric COVID; pCoV) and 14 matched pediatric healthy controls (pHC; median IQR age: 2.01 [1.86–4.28] months) (Fig. 1a). Patient demographics, clinical and laboratory data and treatment are summarized in Supplementary Data 1a, b. We categorized pCoV patients into three groups (G1, G2 and G3; Fig. 1b) based on (i) the time (days) since exposure (DSE) to a known COVID-19 case, (ii) disease severity, and (iii) semi-quantitative viral loads (VL) measured within a median of 17 [11–36] hours of hospitalization. Infants in G1 (n = 10) were diagnosed with COVID-19 as part of the diagnostic work-up for infants prompting hospitalization mainly for fever and/or respiratory symptoms. Infants in G1 had a median of 21 days since they were exposed to an index case and had lower SARS-CoV-2 viral loads (subacute group). Infants in G2 (n = 11) had moderate disease and high SARS-CoV-2 loads, but did not require significant medical interventions such as supplemental oxygen. One infant in G2 received intravenous fluids. G3 (n = 5) infants also had high SARS-CoV-2 viral loads but required a higher level of care, such as oxygen supplementation, intensive care unit (ICU) admission or mechanical ventilation (Supplementary Data 1a–d).
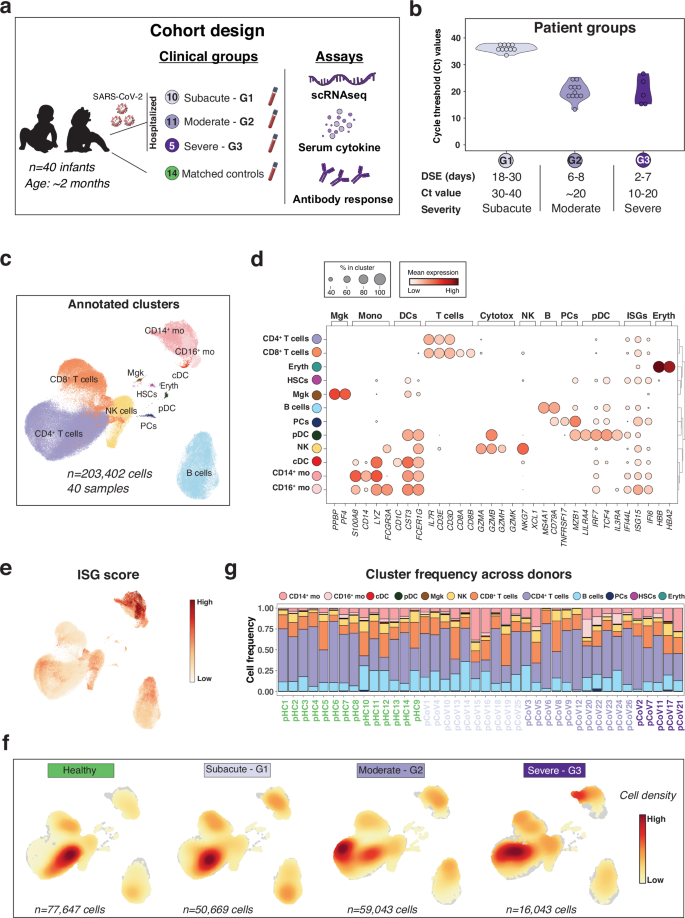
a Study design. b Viral load quantification across the three pCoV groups (Patient groups; G1, G2 and G3). Days since exposure (DSE), cycle threshold (Ct) values and disease severity are shown for each patient group (see Supplementary Data 1 for more details). c UMAP plot representing 203,402 cells colored by annotated immune subsets. Top 100 differentially expressed genes in each subset can be found in Supplementary Data 2. d Immune subset annotation. Dot plot representing expression values of selected genes (x-axis) across each cluster (y-axis). Dot size represents the percentage of cells expressing the marker of interest. Color intensity indicates the mean expression within expressing cells. e UMAP plot representing Interferon-stimulated genes (ISG) score (gene list available in Supplementary Data 3). f UMAP plots representing cell compositional changes (cell density) across healthy controls (pHC) and patient groups (G1, G2 and G3). g Bar plot showing the cell frequency of each cluster across the 40 individuals.
To assess the immune response to SARS-CoV-2 in these infants and to determine how these responses correlated with their varying clinical phenotypes, we performed: (i) single cell RNA-seq (scRNA-seq) of PBMCs, (ii) serum cytokine concentration assessment using the 92-Olink platform with a focus on inflammatory pathways, (iii) quantification of antibodies against coronaviruses (anti-Spike and anti-RBD) and anti-IFN activity, and (iv) semi-quantitative viral load measurement in nasopharyngeal (NP) swabs (Fig. 1a, b and Supplementary Fig. 1a).
For scRNA-seq data analysis, raw data from pHC PBMCs (38% of the pool) and pCoV PBMCs (62% of the pool) were integrated (Fig. 1a). Hybrid transcriptomes (multiplets) were identified using scrublet12 and excluded from the rest of the analysis. After filtering steps, pHC and pCoV samples yielded, respectively, a mean (±S.D.) of 6559 (±2016) cells and 5981 (±1807) cells per individual, and a mean of 1411 and 1603 genes per cell (Supplementary Fig. 1b, c, Supplementary Data 2). scRNA-seq profiles that passed quality control (n = 203,402 cells) were then corrected for batch effect using BBKNN13 (Supplementary Fig. 1d, e). Unsupervised clustering of the corrected data, followed by a two-dimensional uniform manifold approximation and projection (UMAP), yielded 25 clusters independently of 10x run (Supplementary Fig. 1e) and subject (Supplementary Fig. 1f) batch effects. Principle Variance Component Analysis which quantifies the contribution of different factors (e.g., biological groups, sex, age, technical batches, etc) into the total variance, showed that “patient groups” was the major contributor (Supplementary Fig. 1g). Using marker genes, clusters were first mapped to major immune cell types, including CD4+ and CD8+ T cells, B cells, NK cells, CD14+ and CD16+ monocytes, conventional dendritic cells (cDCs), plasmacytoid dendritic cells (pDCs), plasma cells (PCs), hematopoietic stem cells (HSCs) and Erythroblasts/Erythrocytes (Eryth) (Fig. 1c, d, Supplementary Fig. 1h, i). To analyze the cellular distribution of interferon stimulated genes (ISGs) within different immune subsets, we calculated an ISG score based on expression levels of genes in interferon modules14 (Supplementary Data 3). This calculation revealed increased ISG scores in most cell types, including monocytes, T cells and B cells (Fig. 1e). The comparison of PBMCs from acutely infected infants (combined G2 and G3) vs. healthy controls further confirmed a broad ISG upregulation across all cell subsets (Supplementary Data 5a). Cell compositional analyses in pHC, G1, G2 and G3 pCoV groups revealed comparable distribution of pHC and G1 groups, contrasting with striking differences observed in G2 and G3 groups (Fig. 1f, g). Overall, these high-level analyses revealed significant alterations in the PBMC composition of infants upon SARS-CoV-2 infection and a robust ISG signature in most immune cell types in response to the infection. To refine these alterations, we further subclustered and analyzed the different immune cell subsets.
Increased numbers of ISGhi IL-1B+ CD14+ monocytes in infants with mild, moderate and severe disease
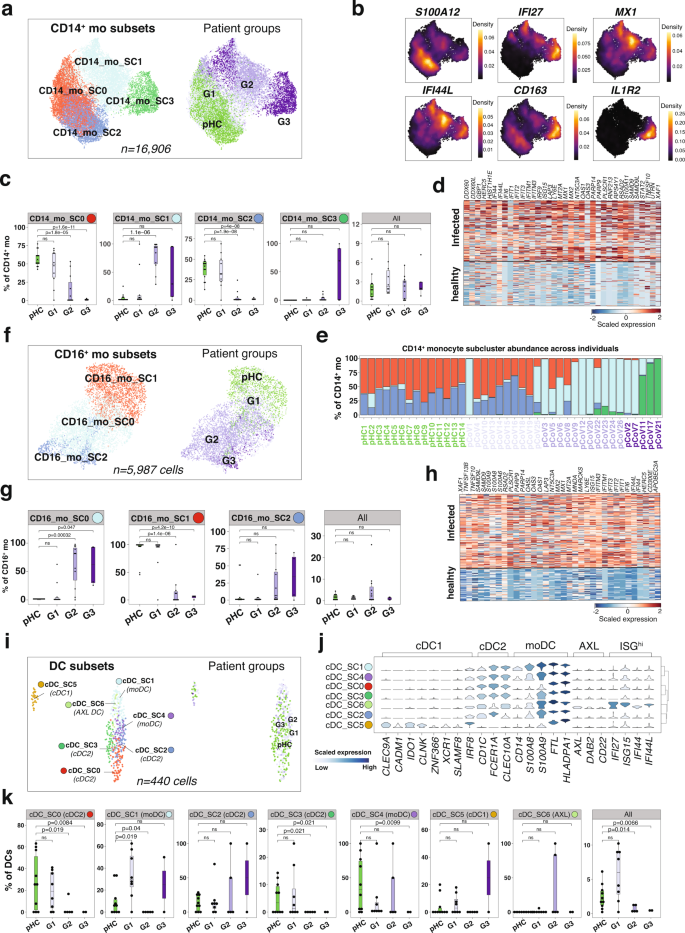
Previous studies have reported alterations in the CD14+ compartment of adults with severe COVID-19, including decreased HLA-DR surface expression and the accumulation of immature circulating myeloid cells, suggesting dysregulated myelopoiesis/innate immune response15,16. To investigate this in our infant cohort, we further clustered CD14+ monocytes (n = 16,906) into four subclusters (SCs) (Fig. 2a). CD14+ mo-SC0 (n = 5495) expressed antigen presentation-related transcripts (e.g., HLA-DRA and CD74) together with small and large ribosomal subunits (e.g., RPS3A and RPL3), while CD14+ mo-SC2 (n = 4590) expressed high levels of alarmins S100A8 and S100A9. CD14+ mo-SC1 (n = 5017) displayed an ISG signature, including ISG15, IFI6, and MX1. CD14+ mo-SC3 (n = 1804) was characterized by the upregulation of IL1R2, CD163, FKBP5, as well as ISGs (Fig. 2b, Supplementary Data 6a). CD14+ mo-SC1 was significantly increased in G2, while CD14+ mo-SC0 and CD14+ mo-SC2 were predominantly present in pHC and G1 (Fig. 2c). CD14+ mo-SC3 was almost entirely contributed by 3/5 patients with severe disease (G3), two of whom had received systemic steroids (Supplementary Data 1a). In fact, the CD14+ mo-SC3 transcriptional profile (Fig. 2b, c) was reminiscent of what was previously described in adults with SARS-CoV-2 infection receiving steroid treatment17,18,19. The comparison of CD14+ monocytes from acutely infected (G2 and G3) infants versus healthy controls revealed a clear upregulation of ISGs (Fig. 2d and Supplementary Data 5b). The ISGhi subcluster (CD14+ mo-SC1) was infrequent in pHC and significantly increased in 2/10 of G1, 11/11 of G2, and 2/5 of G3 patients (Fig. 2e). As both inflammation and type I IFN pathways were previously found activated in monocytes from adults with SARS-CoV-2 infection5, we analyzed the co-expression of inflammatory transcripts and ISGs at the single cell level in these infants. Approximately, 49% of SC1 monocytes co-expressed IL1B and ISG15 (Supplementary Fig. 2a, b).
a UMAP plots representing CD14+ monocyte subclusters (SCs; n = 4, left panel), patient groups (G1, G2, G3) or pHC. b Density plot, generated by the Nebulosa51 package, showing the expression of selected marker genes. c Boxplot comparing the proportion of each CD14+ monocyteSC (n = 4) across the individuals (n = 40). pCoV patient groups (G1, G2 and G3) are shown in purple, and pHC in green. P values were calculated using a two-sided t-test comparing the mean (G1, G2 or G3) with mean (pHC). ns: non-significant. The upper and lower bounds represent the 75% and 25% percentiles, respectively. d Heat map representing scaled expression values of the most significant genes in CD14+ monocytes from acutely infected (G2 and G3) infants, compared with healthy controls. e Bar plot showing the cell frequency of each CD16+ monocyte SC across the 40 individuals, as categorized by patient groups (G1, G2 and G3; in purple) and pHC in green. f As in (a) in CD16+ monocyte SCs (n = 3). g As in (c) in CD16+ monocytes. h As in (d) in CD16+ monocytes. i As in (a) in cDC SCs (n = 7). j Violin plot showing median expression values for selected marker genes across the cDC SCs. k As in (c) in cDC SCs.
Thus, infants with moderate and severe COVID-19 were separated in two groups: the first included 2/10 G1, all G2 and 2/5 G3 infants and was characterized by increased numbers of CD14+ monocytes co-expressing inflammatory molecules and ISGs with low HLA gene expression. The second group consisted of 3/5 G3 infants whose monocytes expressed ISGs and IL1R2 along with CD163, likely reflecting the influence of steroid treatment.
Infants with SARS-CoV-2 infection displayed increased interferon expression but normal frequency of CD16+ monocytes
Contraction of CD16+ monocytes has been previously described as a feature of severe COVID-19 in adults16. Subclustering of CD16+ monocytes in infants (n = 5987) generated three SCs (Fig. 2f). CD16+ mo-SC0 (n = 3009) expressed an ISG signature (e.g., ISG15, IFI6, and MX1). CD16+ mo-SC1 (n = 2217) expressed ribosomal-associated genes RPL5, RPL10A, or RPL3 and CD16+ mo-SC2 (n = 761) was characterized by the expression of inflammatory molecules (e.g., IL1B and CCL3) (Supplementary Fig. 2c, Supplementary Data 6b). CD16+ mo-SC0 (n = 3009) was significantly increased in G2 and G3; SC1 was enriched in pHC and G1, while CD16+ mo-SC2 showed a mild increase in G2/G3 (Fig. 2g). In contrast to the CD14+ population, only about 13% of CD16+ SC0 monocytes co-expressed IL1B and ISG15 (Supplementary Fig. 2d). The comparison of CD16+ monocytes from acutely infected (G2 and G3) infants with those from healthy controls revealed an enrichment in ISGs (Fig. 2h and Supplementary Data 5c). The ISGhi subcluster SC0 was rare in pHC and significantly increased in 9/11 of G2 and 4/5 of G3 infants (Supplementary Fig. 2e). Unlike severe COVID-19 in adults, which was characterized by CD16+ monocyte depletion20, infants with moderate to severe disease (G2/G3) exhibit normal frequencies of monocytes due to the increased numbers of ISGhi CD16+ monocytes.
To explore how our findings compare with previous studies, we integrated our scRNA-seq data with a recent study by Wimmers et al.11, which analyzed PBMCs from older infants with mild disease (average 9 months of age, not hospitalized and untreated) during both Omicron and non-Omicron variants of SARS-CoV-2 infection. This cohort included: (i) nine samples collected before (pre, n = 9), during (acute; n = 10) and after infection (convalescent or conv; n = 9) with non-Omicron variants, (ii) eight samples during acute Omicron infection (acute omicron) and (iii) seven matched controls. Our integrative analysis of CD14+ and CD16+ monocytes from 83 infants in total confirmed the presence of an ISG signature in infants with moderate to severe disease (10/10 G2 and 5/5 G3 patients in our cohort), which contrasted with only 5/10 acute non-Omicron and 1/10 acute Omicron patients in the Wimmers cohort (Supplementary Fig. 2f–i). These differences are likely attributable to variations in the time from exposure to the virus, the age of the infants, and, most importantly, the clinical disease severity between the two cohorts.
SARS-CoV-2 infected infants showed reduced frequency of conventional dendritic cells (DCs)
Similar to classical monocytes, DCs from SARS-CoV-2 infected adults exhibit lower levels of HLA-DR compared to healthy controls21. Subclustering conventional DCs (cDCs) in infants (n = 440) generated seven SCs (Fig. 2i and Supplementary Fig. 3a), which were further classified into (i) cDC1 (SC5; expressing CLEC9A and XCR1), (ii) cDC2 (SC0/SC3; expressing CD1C and CLEC10A), (iii) monocyte-derived cDCs (Mo-DCs; SC1/SC4; expressing CD14 and S100s) and (iv) AXL+ DCs (SC6; expressing AXL and DAB2) (Supplementary Data 6c). The AXL+ DCs (SC6) expressed the highest levels of ISGs (Fig. 2j and Supplementary Fig. 3b). G2/G3 groups showed a marked reduced numbers of cDC1, cDC2 and Mo-DCs, while two G2 patients showed an increase in AXL+ ISGhi SC (SC6) (Fig. 2k and Supplementary Fig. 3c). The comparison of cDCs from acutely infected (G2 and G3) infants with those from healthy controls revealed a pronounced upregulation of ISGs (Supplementary Data 5e). Overall, our data showed that profiles of G2 and G3 infants were characterized by a decrease in cDC1, cDC2, and Mo-DCs.
The frequency of pDCs in peripheral blood was reduced in adults with COVID-1922, possibly due to pDC apoptosis17. Subclustering of pDCs in infants (n = 386) generated two SCs (Supplementary Fig. 3d). pDC-SC0 (n = 226) was significantly reduced in G2/G3 patients, while pDC-SC1 (n = 160) was significantly increased in these infants and characterized by the upregulation of ISGs (Supplementary Fig. 3e, f). The comparison of pDCs from acutely infected (G2 and G3) infants with those from healthy controls revealed a pronounced upregulation of HLA related molecules and ISGs (Supplementary Data 5f). The depletion of ISGlow pDCs is consistent with reports in adults with COVID-19; however, a subset of ISGhi circulating pDCs was increased in G2/G3 infants.
Circulating megakaryocytes (MGKs) have previously been linked to COVID-19 outcomes in adults23. Two of the infant’s SCs (in C22, Supplementary Fig. 1h, i) were annotated as megakaryocyte (MGK; n = 408; expression of PPBP, CLU and PF4) and hematopoietic stem cells (HSCs; n = 305; expression of CD34 and ITM2C) (Supplementary Fig. 3g, h, Supplementary Data 6e). MGKs (n = 408 cells) were significantly increased while HSC (n = 305 cells) were significantly decreased in G1 and G3 groups (Supplementary Fig. 3i).
ISGhi naive CD4+ T cells are increased in SARS-CoV-2 infected infants
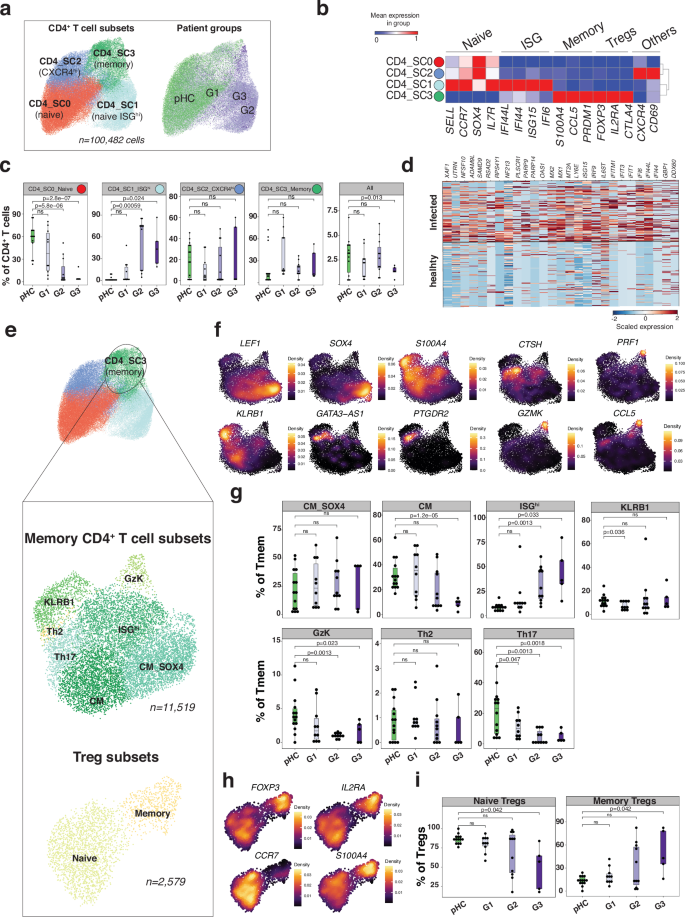
Activation and proliferation of both CD4+ and CD8+ T cells have been associated with COVID-19 disease severity4,17, though no specific T cell signatures have been identified as primary correlates in this context. In this infant study, subclustering CD4+ T cells (n = 100,482) yielded four SCs (Fig. 3a). CD4+T-SC0 (n = 43,567), expressing ribosomal protein-associated genes such as RPL5, RPL10A and RPL3, was significantly decreased in G2 and G3 infants. CD4+T-SC1 (n = 22,847), which exhibited an ISG signature (e.g., IFI44L, ISG15 and XAF1), was increased in G2 and G3 infants. CD4+T-SC2 (n = 14,383), characterized by expression of JUNB/JUND, CD69 and CXCR4 did not differ between infants with COVID-19 and pHC. Finally, CD4+T-SC3 (n = 14,383), expressing the memory marker S100A4, was increased in all three pCoV groups (Fig. 3b, Supplementary Data 6f). The ISGhi CD4+T-SC1 was present in 2/10 G1, 8/11 G2, and 5/5 G3 infants. Notably, more than 90% of CD4+ T cells from a fraction of G2 and G3 patients (e.g., pCoV11 and pCoV26) switched to an ISGhi state (Fig. 3c). Overall comparison of CD4+ T cells from acutely infected (G2 and G3) infants with those from healthy controls revealed an enrichment in ISGs (Fig. 3d, Supplementary Data 5g). CD4+T-SC0 and CD4+T-SC1 exhibited naive (CCR7, LEF1 and SELL) markers, while CD4+T-SC3 displayed memory markers (Fig. 3b). Our analysis also revealed the expression of the transcription factor SOX4 within the naive compartment, which is consistent with an immature/naive phenotype (Supplementary Fig. 4a). While the majority of ISGhi T cells originated from infected patients, we also detected these cells in pHC, indicating constitutive expression of ISGs in young infants (Supplementary Fig. 4b, c).
a UMAP plots representing CD4+ T cell subclusters (SCs; n = 4) and patient groups (G1, G2, G3 or pHC). b Heat map representing scaled expression values of selected marker genes. c Boxplot comparing the proportion of each CD4+ T cell SC (n = 4) across the individuals (n = 40). pCoV patient groups (G1, G2 and G3) are shown in purple, and pHC in green. P values were calculated using a two-sided t-test comparing the mean (G1, G2 or G3) with mean (pHC). ns: non-significant. The upper and lower bounds represent the 75% and 25% percentiles, respectively. d Heat map representing scaled expression values of the most significant genes in CD4+ T cells from acutely infected (G2 and G3) infants, compared with healthy controls. e UMAP plot representing memory CD4+ T cell subsets. f Density plot, generated by the Nebulosa51 package, showing the expression of selected marker genes. g As in (c) in memory CD4+ T cell SCs. h As in (f) in Tregs. i As in (g) in Tregs.
Further analysis of CD4+T-SC3, resulted in the separation of memory (Tmem, six subsets) and regulatory T cells (Tregs; two subsets). Tmem included seven subsets: (a) central memory (CM), (b) SOX4+ CM, (c) ISGhi, (d) GZMK+ Tmem (Gzk), (e) KLRB1+ Tmem, (f) TH2 (PTGDR2, GATA3-AS1) and (g) TH17 (CTSH) (Fig. 3e, f). Infected infants exhibited increased numbers of ISGhi Tmem, along with reduced numbers of GzK and TH17 cells (Fig. 3g). Treg were separated into naive and memory (Fig. 3e, h). G3 infants exhibited lower naive Tregs and higher memory Treg frequencies (Fig. 3i). Within the memory compartment, our analysis revealed the presence of cells expressing both GMZK and Temra-associated transcripts such as PRF1 and CCL5, an unusual finding in the adult CD4+ T cell memory compartment (Fig. 3f).
Analysis of granzyme (GZM) family expression revealed a broad upregulation of GZMM, contrasting with an absence of GZMB and restriction of GZMK to CD4+T-SC3.
The integration analysis with the Wimmers et al. dataset further confirmed an augmented ISG signature in all G2 and G3 individuals in our cohort, contrasting with 4/10 acute non-Omicron and 1/10 acute Omicron individuals in the Wimmers cohort (Supplementary Fig. 5a, b). Overall, these data indicate that the blood CD4+ T cells of COVID-19 infected infants fall into two main compartments: (i) naive, as expected the predominant population, that had switched to an ISGhi state, and (ii) memory, which encompassed a mixture/combination of central and effector memory CD4+ T cells, T helper subsets and Tregs that was altered in SARS-CoV-2 infected infants.
ISGhi CD8+ T cells are increased in infants with COVID-19
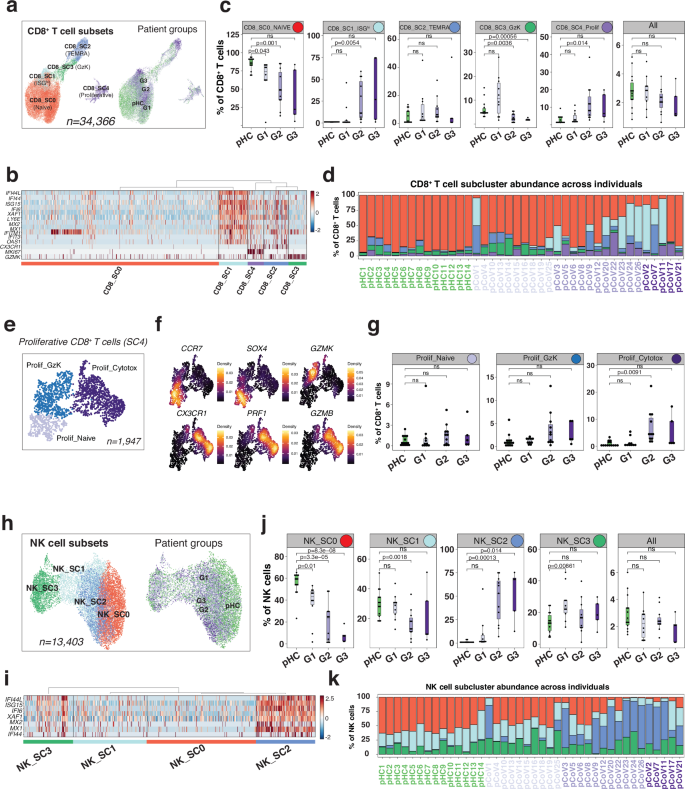
COVID-19 associated lymphopenia3 is reported to preferentially impact CD8+ T cells4, possibly due to T cell exhaustion24. However, the overall CD8+ T cell compartment (n = 34,366), was not decreased in infants with COVID-19 and was distributed into five SCs (Fig. 4a). CD8+ T-SC0 (naive, n = 23,768) expressed ribosomal-associated genes (e.g., RPL34 and RPL39), and was decreased in G2 and G3 infants; CD8+T-SC1 (ISGhi; n = 3483) exhibited an ISG signature (e.g., IFI44L, ISG15 and XAF1) and was increased in G2/G3 infants. CD8+ T-SC2 (n = 2967) displayed similar frequencies in infected infants and pHC and expressed cytotoxic genes, including PRF1, GZMA and GNLY, inflammatory chemokines (CCL4 and CCL5), S100A4 and CX3CR1 consistent with a TEMRA phenotype. CD8+ T-SC3 (GzK; n = 2201) expressed GZMK and CCL5 and was mostly present in G1, but reduced in G2/G3 infants. Finally, CD8+T-SC4 (Prolif.; n = 1947), which expressed proliferation markers such as MKI67 and HMGBs, was increased in G2 and G3 infants (Fig. 4b, c, Supplementary Data 6g). The CD8+ T ISGhi SC was rare in pHC but present in 2/10 G1, 8/11 G2, and 3/5 G3 infants. Notably, more than 50% of CD8+ T cells in some G2 (pCoV13, pCoV24, pCoV26) and G3 (pCoV2 and pCoV11) infants demonstrated an ISGhi state (Fig. 4d). A direct comparison of CD8+ T cells from acutely infected (G2 and G3) infants with those from healthy controls revealed an upregulation of ISGs (Supplementary Data 5h). The proliferative SC4 included a subset of cells expressing TEMRA markers, such as CX3CR1, FCGR3A and GZMB (Fig. 4e, f). This SC represented 15–20% of CD8+ T cells in 6/16 of the G2/G3 infants (Fig. 4d). Further clustering of proliferative CD8+ T cells revealed three subsets, including a naive (CCR7, SOX4), a granzyme K (GZMK) and cytotoxic (PRF1 and GZMB) subsets (Fig. 4e), the latter being significantly increased in G2 (Fig. 4g). Comparison with published data11 on infants with mild disease further confirmed the presence of a robust ISG signature in all G2 and G3 patients in our cohort, contrasting with 4/10 acute non-Omicron and 1/10 acute Omicron individuals in the Wimmers cohort (Supplementary Fig. 5c, d). In summary, infants with more severe disease exhibited increased numbers of ISGhi CD8+ T cells and induction of CX3CR1+ proliferating CD8+ T cells with a cytotoxic program, a finding that contrasts with previous reports in adults25.
a UMAP plots representing CD8+ T cell subclusters (SCs, n = 5) and patient groups (G1, G2, G3 or pHC). b Heat map representing scaled expression values of selected ISGs and marker genes across CD8+ T cell SCs. c Boxplot comparing the proportion of each CD8+ T cell SC (n = 5) across the individuals (n = 40). pCoV patient groups (G1, G2 and G3) are shown in purple, and pHC in green. P values were calculated using a two-sided t-test comparing the mean (G1, G2 or G3) with mean (pHC). ns: non-significant. The upper and lower bounds represent the 75% and 25% percentiles, respectively. d Bar plot showing the cell frequency of each CD8+ T cell SC across the 40 individuals, as categorized by patient groups (G1, G2 and G3; in purple) and pHC in green. e UMAP plot representing proliferative CD8+ T cell SCs. f Density plot, generated by the Nebulosa51 package, showing the expression of selected marker genes. g As in (c) in proliferative CD8+ T cell SCs. h As in (a) in NK cells. i As in (b) in NK cells. j As in (c) in NK cells. k As in (d) in NK cells.
ISGhi NK cells are increased in infants with severe SARS-CoV-2 infection
A recent study reported an increase of “adaptive” NK cells in the blood of adults with severe COVID-1926. However, little is known about the role of these cells in infants. Herein, NK cell (n = 13,403) subclustering yielded four SCs (Fig. 4h). NK-SC0 (n = 5013), reduced in G2 and G3 infants, expressed ribosomal-associated genes. NK-SC1 (n = 3393), reduced in G2 infants, expressed CD160 and KLRB1. NK-SC2 (n = 2721), increased in G2/G3 infants, exhibited an ISG signature. NK-SC3 (n = 2276) displayed similar frequencies in pHC and infected infants and upregulated transcripts such as XCL1, XCL2 and GZMK (Fig. 4i, j). While very rare in pHC, the ISGhi SC2 was present in 2/10 G1, 11/11 G2, and 5/5 G3 infants. Notably, more than 70% of NK cells from 9/11 G2 and all 5 G3 infants switched to an ISGhi state (Fig. 4k, Supplementary Data 6h). The direct comparison of NK cells from acutely infected (G2 and G3) infants with those from healthy controls confirmed the enrichment in ISGs (Supplementary Data 5i).
The comparison with previously reported cohorts confirmed the presence of a strong ISG signature in all G2 and G3 patients in our cohort, contrasting with 5/10 acute non-Omicron and 1/10 acute Omicron patients in the Wimmers cohort (Supplementary Fig. 5e, f).
SARS-CoV-2 infection in infants is associated with increased ISGhi transitional and naive B cells
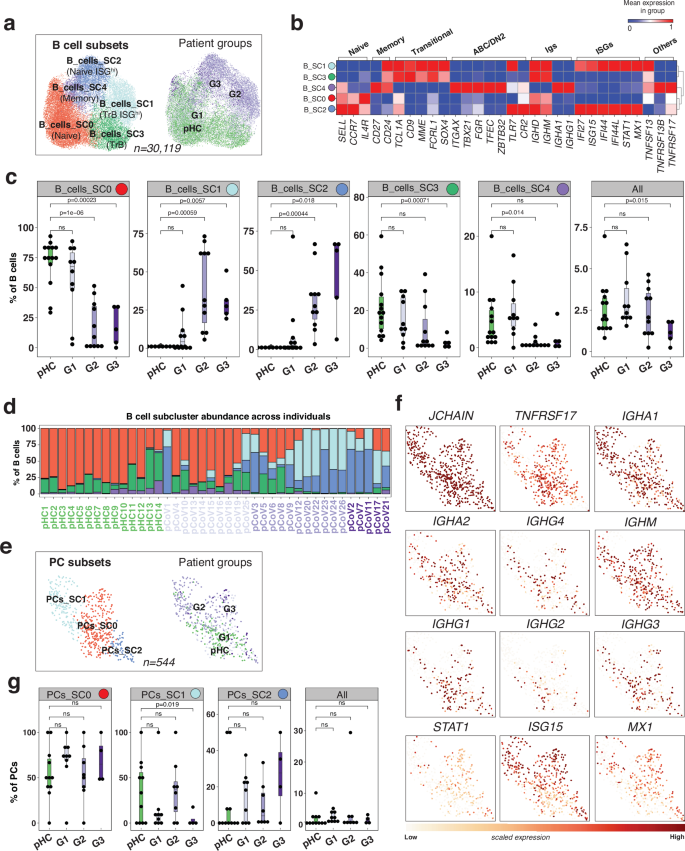
In adults with COVID-19, activated B cells and plasmablasts are expanded4,27, and an increased frequency of extrafollicular (Tbet+CD11c+CXCR5neg) B cells correlates with disease severity28. In our study, further clustering of infant’s B cells (n = 30,119) generated five SCs (Fig. 5a). Differential marker expression distinguished three compartments: (i) naive B cells (NBC; IGHD, CCR7 and SELL), (ii) Transitional B cells (TrBC; MME (encoding CD10), CD9, and CD24) and (iii) Memory B cells (MBCs; CD27, IGHA1, IGHG1 and S100A4). An ISG signature (e.g., IFI44L, ISG15 and IRF7) was detected in both the NBC and TrBC compartments (Fig. 5b). In G2/G3 infants, B-SC1 (n = 5228) and B-SC2 (n = 4985) were increased, while B-SC0 (n = 13,820), B-SC3 (n = 4857) and B-SC4 (n = 1229) were decreased (Fig. 5c). Thus, NBCs included B-SC0 (ISGlow; increased in pHC/G1) and B-SC2 (ISGhi; increased G2/G3); TrBCs encompassed B-SC3 (ISGlow; increased in pHC/G1) and B-SC1 (ISGhi; increased in G2/G3). B-SC4 included transcripts characteristic of double negative (DN)2 memory cells (e.g., ITGAX, TFEC and FGR) (Fig. 5b, Supplementary Fig. 4d, Supplementary Data 6i). The ISGhi naive B-SC2 expressed TLR7, a marker of both extrafollicular activated naive and DN2 cells29 (Fig. 5b). In addition to memory markers (conventional, switched and extrafollicular), B-SC4 also expressed B cell survival and plasma cell differentiation markers such as TNFRSF13B (encoding TACI) and TNFRSF17 (encoding BCMA), which could correspond to cells prone to differentiate into plasma cells. Although rare in pHC, the ISGhi SC was present in 2/10 G1, 11/11 G2, and 5/5 G3 infants. Overall, >70% of naive (B-SC2) and transitional (B-SC1) B cell compartments from 13/16 of the G2/G3 infants switched to an ISGhi state (Fig. 5d). The comparison of B cells from acutely infected (G2 and G3) infants with those from healthy controls revealed an enrichment in ISGs (Supplementary Data 5j). While the majority of the ISGhi B cells originated from infected patients, we also identified an ISG signature in both naive and transitional B cells from healthy control infants (Supplementary Fig. 4e, f).
a UMAP plots representing B cell subclusters (SCs; n = 5) and patient groups (G1, G2, G3 or pHC). b Heat map representing scaled expression values of selected marker genes defining each of the B cell SC (Supplementary Data 6i). c Boxplot comparing the proportion of each B cell-SC (n = 5) across the individuals (n = 40). pCoV patient groups (G1, G2 and G3) are shown in purple, and pHC in green. P values were calculated using a two-sided t-test comparing the mean (G1, G2 or G3) with mean (pHC). ns: non-significant. The upper and lower bounds represent the 75% and 25% percentiles, respectively. d Bar plot showing the cell frequency of each B cell SC across the 40 individuals, as categorized by patient groups (G1, G2 and G3; in purple) and pHC in green. e As in (a) in plasma cells. f UMAP plots representing scaled expression values of selected marker genes. g As in (c) in plasma cells.
The comparison between cohorts demonstrated the presence of a strong ISG signature in all G2 and G3 patients in our cohort, contrasting with 10% and 40% of acute Omicron and non-Omicron individuals, respectively, in the Wimmers cohort (Supplementary Fig. 5g, h).
Finally, plasma cells (PC, n = 544) subclustering generated three SCs (Fig. 5e), which were distinguishable based on IGH expression (Fig. 5f). Although the low number of cells is a limitation, cell compositional analysis did not reveal significant differences between COVID-19 infant groups and pHC (Fig. 5g), which contrasts with what was reported in adults with COVID-1930.
Patient stratification using cell frequencies
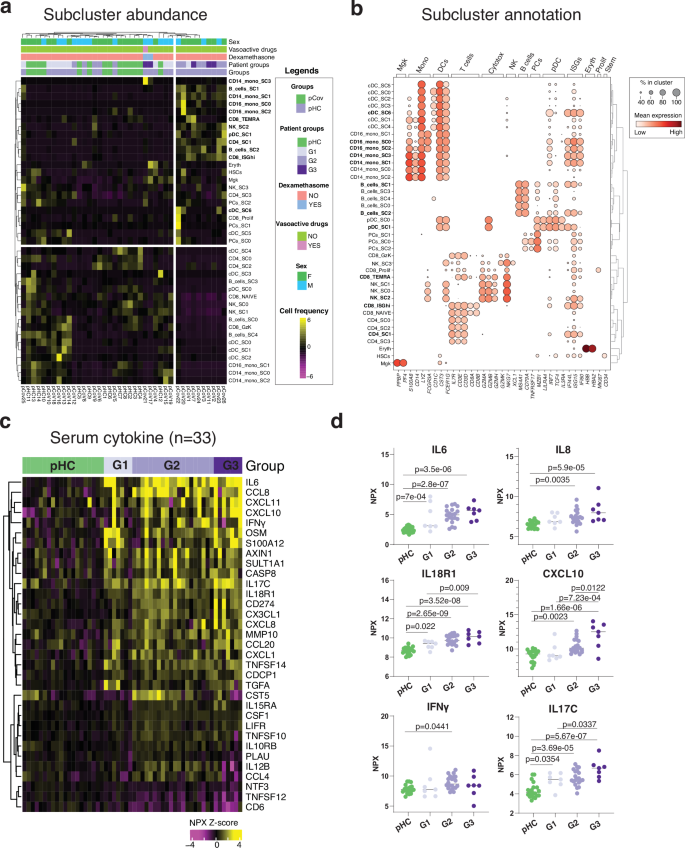
To integrate cell subcluster analysis data (Supplementary Fig. 6a–d) with clinical disease severity, we performed an unsupervised clustering based on the abundance of detected subclusters (n = 40) across infants with COVID-19 (n = 26) and pHC (n = 14). The study subjects were accordingly clustered into two main sets (Fig. 6a). The first set, which included 10 infants with COVID-19 (3/5 G3, 6/11 G2, and 1/10 G1 infants), was characterized by the expansion of ISGhi SCs. The second set, which included the remaining 16 COVID-19 infants together with the 14 pHC, displayed ISGlow SCs including naive CD4+, CD8+ T cells and B cells (Fig. 6b). Interestingly, the two G3 patients who clustered with pHC/G1 (pCoV17 and pCoV21) had received systemic steroids, consistent with the modulatory effect of corticosteroids on type I IFN expression31. These data therefore support that clinically severe SARS-CoV-2 infection in infants is associated with a robust ISG response.
a Heatmap representing the abundance of each of the subclusters (n = 40) across the individuals (n = 40). Clinical information are highlighted on top of the heatmap. b Subcluster annotation. Dot plot represents expression values of selected genes (x-axis) across each cluster (y-axis). Dot size represents the percentage of cells expressing the marker of interest. Color intensity indicates the mean expression within expressing cells. c Heatmap representing the most significant (two-sided ANOVA, padj < 0.05) inflammatory-related cytokine (CK) measurements (n = 33 CK) across the individuals (n = 54; G1 n = 7; G2 n = 20, G3 n = 7 and matched HC n = 20; Supplementary Data 7). d Dot plot representing selected CK across the individuals, as categorized per patient groups and healthy controls. *P < 0.05; **P < 0.01; ***P < 0.001: ****P < 0.0001.
SARS-CoV-2 infected infants had increased serum concentrations of inflammatory cytokines
To complement our transcriptional profiling studies, we measured serum concentrations of cytokines in 34 children with COVID-19 (7 G1, 20 G2, and 7 G3) and 20 age-matched pHCs using the Olink inflammatory panel (n = 92 analytes). This cohort included 24 of the 26 pCoV infants profiled with scRNAseq (Supplementary Fig. 1a, Supplementary Data 1e). Overall, 72 cytokines were consistently detected in serum samples (Supplementary Fig. 6e); and 33 of them showed significantly different concentrations between disease severity groups (Fig. 6c). IFNγ concentrations were increased in infants with COVID-19 across all disease groups. Cytokines and inflammatory proteins such as IL6, IL8, IL17C, IL18R1 and CXCL10 (IP-10) were particularly increased in G3 infants (Fig. 6d), while soluble CD6 and TNFS12/TWEAK were markedly decreased in this group of severe disease compared with pHCs. Patients in the Wimmers cohort did not show significant increases in plasma inflammatory cytokines possibly due to differences in time since infection and/or clinical disease severity. In summary, SARS-CoV-2 infected infants displayed increased serum concentrations of inflammatory cytokines, especially those infants with severe disease (G3).
Infected infants develop anti-SARS-CoV-2 IgG antibodies but not anti-seasonal coronavirus or anti-IFN responses
To assess the infants’ humoral response to SARS-CoV-2 and seasonal coronaviruses, we measured antibody levels against: (i) SARS-CoV-2 full-length spike, RBD and S2 subunit, (ii) seasonal coronaviruses full-length spike, (iii) S1 subunit of beta-coronaviruses OC43 and HKU1, and (iv) full-length spike of alpha-coronavirus 229E. Samples from 16 infants were obtained at the time of hospitalization, and 13 of those had paired follow-up samples four weeks later. The 13 infants with paired samples, who were also included in the scRNAseq cohort (Supplementary Fig. 1a, Supplementary Data 1f), predominantly (12/13) showed a significant increase in IgG antibody titers against SARS-CoV-2 antigens at follow-up, including spike, RBD and S2 subunit (Supplementary Fig. 7a, b). While 10 infants increased their anti-S2 and anti RB levels by 20 and ~8 fold, respectively, no increase in antibody titers against any of the seasonal alpha- or beta-coronaviruses was observed, despite the presence of detectable IgG levels at baseline (T0); (Supplementary Fig. 7c). Given the young age of this cohort (median 3.2 months), we hypothesized that the detected antibodies against seasonal coronaviruses are likely of maternal origin.
Finally, although anti-IFN autoantibodies have been identified in adults with severe COVID-1932, none of the patients (pCoV, n = 16) or pHCs (n = 6) in this pediatric cohort demonstrated anti-IFN activity in either the acute or convalescent serum samples (Supplementary Fig. 7d).
Lymphocytes from SARS-CoV-2 infected infants express a broader ISG signature compared to those from infected adults
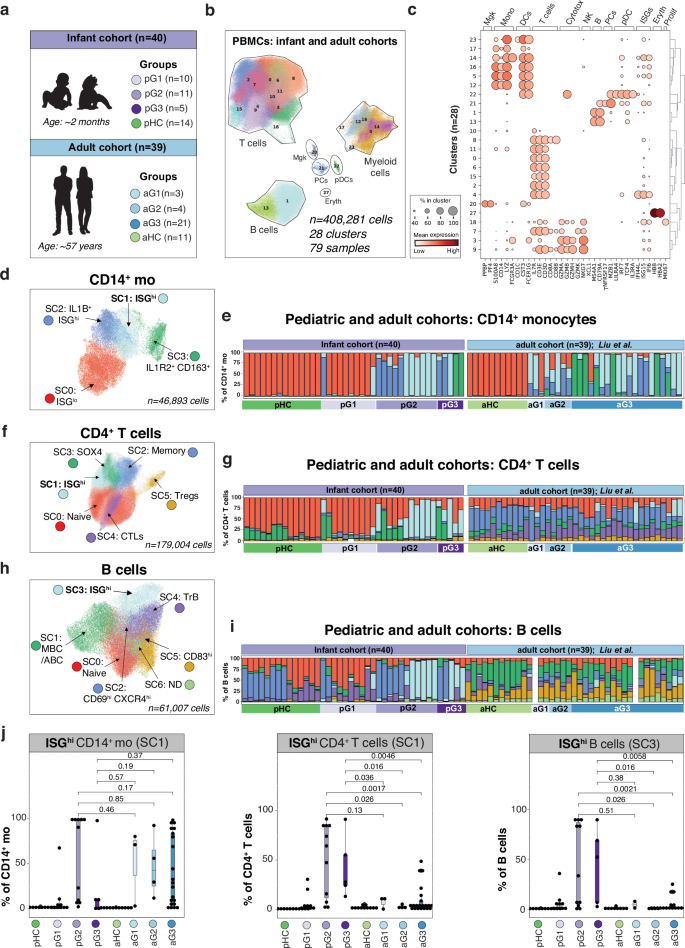
We next compared our infant data with a scRNA-seq PBMC dataset (GSE161918) from a reported cohort of 28 adults with COVID-19 and controls17. For this comparison, we included data from the adult patients’ first time-point (T0), as well as from 11 healthy matched controls (aHC)17. After concatenating adult (n = 39) and infant (n = 40) datasets, we applied our pipeline (see methods for details) on doublet-cleaned data, which included pre-processing, batch correction (using Harmony33), and unsupervised clustering. Based on the criteria we used for the infant cohort (excluding viral loads, which were not available for the adult dataset), we categorized adult patients into three clinical groups, from lower to higher severity (aG1, aG2 and aG3; Fig. 7a). The first round of clustering generated 28 clusters and seven cell-types (Fig. 7b, c) that were then analyzed separately.
a Pediatric and adult cohorts’ information. b UMAP plot representing clusters (n = 28) from 408,281 PBMCs and 79 samples (40 infants and 39 adults) c Cluster annotation. Dot plot represents expression values of selected genes (x-axis) across each cluster (y-axis). Dot size represents the percentage of cells expressing the marker of interest. Color intensity indicates the mean expression within expressing cells. d UMAP plots representing CD14+ monocyte subclusters (SCs; n = 4) and their respective annotations. e Bar plot showing the cell frequency of each CD14+ monocyte SC across the 79 individuals, as categorized by patient groups: pG1, pG2 and pG3; in purple, pHC in dark green, aG1, aG2 and aG3; in blue and aHC in light green. f As in (d) in CD4+ T cell SCs (n = 6). g As in (e) in CD4+ T cell SCs. h As in (d) in B cell SCs (n = 7). i As in (e) in B cell SCs. j Boxplot comparing the proportion of ISGhi CD14+ monocyte (SC1), CD4+ T cell (SC1) and B cell (SC3) subclusters across the individual (n = 79) in both pediatric (n = 40) and adult (n = 39) cohorts. P values were calculated using a two-sided t-test comparing the mean (G1, G2 or G3) with mean (pHC). ns: non-significant. The upper and lower bounds represent the 75% and 25% percentiles, respectively.
CD14+ monocytes included four subclusters (SCs), which were categorized into: (i) ISGlow (SC0), (ii) ISGhi (SC1), (iii) ISGhi inflam+ (SC2) and (iv) CD163+ IL1R2+ (SC3) (Fig. 7d and Supplementary Fig. 8a, b). Cell composition analysis of CD14+ monocytes showed a decrease of SC0 (ISGlo) and a switch to SC1 (ISGhi) and/or SC2 (IL1B+ ISGhi) in pG2/pG3 and aG2/G3 groups compared to their respective age-matched controls. Interestingly, we replicated the increase in SC3 (IL1R2+ CD163+) in patients receiving steroid therapy (Fig. 7e). The analysis of CD16+ monocytes showed a switch to an ISGhi state in both G2/G3 infants and G3 adults (Supplementary Fig. 8c–e).
Subclustering CD4+ T cells generated six SCs (Fig. 7f), annotated as: (i) naive (SC0 and SC1; SELL and CCR7), with SC0 being ISGlow and SC1 ISGhi, (ii) memory (SC2 and SC4, S100A4), (iii) SOX4+CD4+ T cells (SC3; SOX4) and (iv) regulatory T cells (SC5, FOXP3 and IL2RA). SC4 (CTLs) exhibited the cytotoxic cell-associated chemokine CCL5 (Supplementary Fig. 8f). Cell composition analysis showed: (i) an increase in numbers of naive cells in pediatric healthy controls (pHC) compared to adult healthy controls (aHC), (ii) a reduction of naive cells in pG2/pG3 compared to pHC/pG1, (iii) an increase of memory cells (SC2) in aHC compared to pHC, (iv) increase of CTLs (SC4) in aHC, and (v) an increase of SOX4+ CD4+ T cells in pHC. Total Tregs (SC5) did not significantly change upon infection in either infants or adults. The ISGhi CD4+ T cell SC (SC1) was present in both infected infants (2/10 pG1, 8/11 pG2, and 5/5 pG3) and adults (6/21 aG3 patients), although their proportion was higher in infants (Fig. 7g).
B cell subclustering generated seven SCs (Fig. 7h), annotated as: (i) naive (SC0, CCR7), (ii) activated (SC1, CD69) (iii) memory (SC2, S100A4) and (iv) transitional (SC4; CD9, MME (CD10)). SC3 encompassed ISGhi B cells and SC5 upregulated CD83 (Supplementary Fig. 8g). Cell composition analysis showed: (i) increase of naive B cells in pHC compared to aHC, (ii) an increase of memory B cells in aHC compared to pHC, and (iii) a subtle ISGhi increase in infected adults (aG3) contrasting with a near-complete ISGhi status in the infected infants with severe disease (pG2/pG3) (Fig. 7i).
CD8+ T cell subclustering generated five SCs, classified as: (i) naive (SC0; RPLs), (ii) TEMRA (SC1; GZMB), (iii) Gzk (SC2, GZMK), (iv) MAIT (SC3, KLRB1 and ZBTB16) and (v) proliferative CD8 T cells (SC4, MKI67) (Supplementary Fig. 9a). As expected, cell composition analysis showed increased naive CD8+ T cells in pHC and TEMRA/MAIT cells in aHC (Supplementary Fig. 9b). NK cell subclustering generated two SCs (Supplementary Fig. 9c). SC0 upregulated FCGR3A (CD16+ NK), and SC1 was XCL1+; no disease severity-associated differences were detected (Supplementary Fig. 9d).
Overall, these comparative analyses (Supplementary Fig. 10a–d) showed that CD4+ T and B cells from infants with severe disease almost entirely switch to an ISGhi state—while this is only observed in a minority of T and B cells in adults with severe disease (Fig. 7j).