Clinical cohort description
To study early events during the immune response to Mtb, we used biobanked samples from a cross-sectional study performed at GHESKIO Centers, Port-au-Prince, Haiti to assemble a clinical cohort of household contacts of active pulmonary TB cases and source community controls between 2015 and 2018 (NIH U19 AI111143, completed). A total of 78 active TB cases, 92 household contacts, and 605 community controls were recruited. We randomly sampled an age-, sex-, and IGRA-matched subset of 50 TB household contacts, 50 active TB cases from the same households, and 50 source community controls without household exposure, based on peripheral blood mononuclear cell (PBMC) availability and quality control (total n = 150). The overall cell viability was similar across all three patient cohorts, active TB (M = 69.4%, SD = 25.0%), household contacts (M = 75.8%, SD = 22.6%), and community controls (M = 81.2%, SD = 14.6%). All available specimens from IGRA− contacts (n = 14) were analyzed.
Age, sex, IGRA status, and relevant sociodemographic variables are summarized in Table 1. No significant variations were observed in the number of household members, household income, smoking status, or alcohol consumption between groups, consistent with previous work18. Among all recruited participants, 75.8% of household contacts were IGRA+ while 51.6% of community controls were IGRA+, consistent with increased transmission within the household. Fifty-percent of contacts slept in the same room as the active pulmonary TB index case. Among the 14 IGRA− household contacts selected in our study, only two IGRA conversions were detected during 6-month follow-up IGRA testing, supporting the premise that IGRA− contacts may resist Mtb infection without developing Mtb peptide-specific immunoreactivity.
IGRA+ community controls have elevated frequency of mature CD56− NK cells
We first compared the frequency of NK cell subpopulations between clinical cohorts using flow cytometry. Human NK cells were defined by the absence of lineage markers for T cells (CD3), monocytes (CD14), and B cells (CD19), and gated on surface expression of neural cell adhesion molecule (NCAM), CD56, and Fc receptor γRIIIA, CD16 (Fig. 1A). We stratified NK cells into three subsets based on expression of CD56: CD56−, CD56dim, and CD56bright40,41.
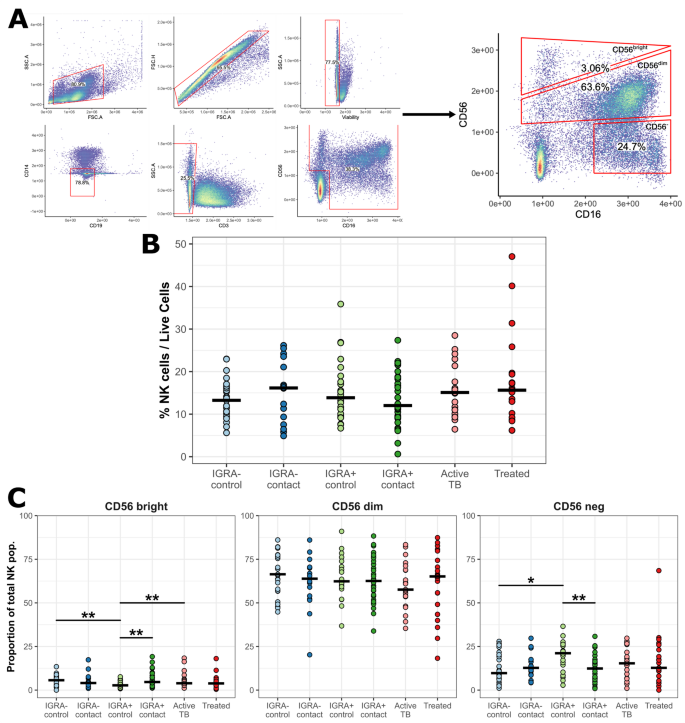
Frequencies of NK cell subpopulations are altered after Mtb infection. (A) Gating strategy employed for NK cells from a representative sample. NK cells were defined as CD19−, CD3−, CD14− lymphocytes that expressed any level of either CD16 or CD56. NK cells were further subdivided into three subsets based on CD56 expression. (B) Proportion of total NK cells among live cells in clinical groups. (C) Proportion of each NK cell subset in clinical groups. Frequency calculated as percent of each subset among total NK cells. Brackets represent Wilcoxon statistical tests, with unadjusted p values. *p ≤ 0.05, **p ≤ 0.01.
We subdivided the community controls and household contacts based on IGRA status. The active TB cohort was stratified by completion of TB treatment. The statistical analysis plan included a priori hypothesis tests as pairwise comparisons between two clinical groups: IGRA− contacts vs. controls, IGRA+ contacts vs. controls, IGRA− contacts vs. IGRA+ contacts, IGRA− controls vs. IGRA+ controls, IGRA+ controls vs. Active TB, IGRA+ contacts vs. Active TB, and Active TB prior to vs. after therapy completion. This enabled testing the overarching hypothesis that specialized NK cell populations responded during asymptomatic household exposure and infection. There was no difference in the frequency of total NK cells or CD56dim NK cells between clinical groups (Fig. 1B, C center). IGRA+ community controls had fewer CD56bright NK cells (3.11% vs. 5.45%, p = 0.006) and more CD56− NK cells (19.1% vs. 12.6%, p = 0.002) compared to IGRA− controls or IGRA+ contacts, indicating an accumulation of mature NK cells in immunoreactive donors within the source community (Fig. 1C, right and left).
IGRA+ contacts demonstrate a loss of circulating CD16+ NK cells
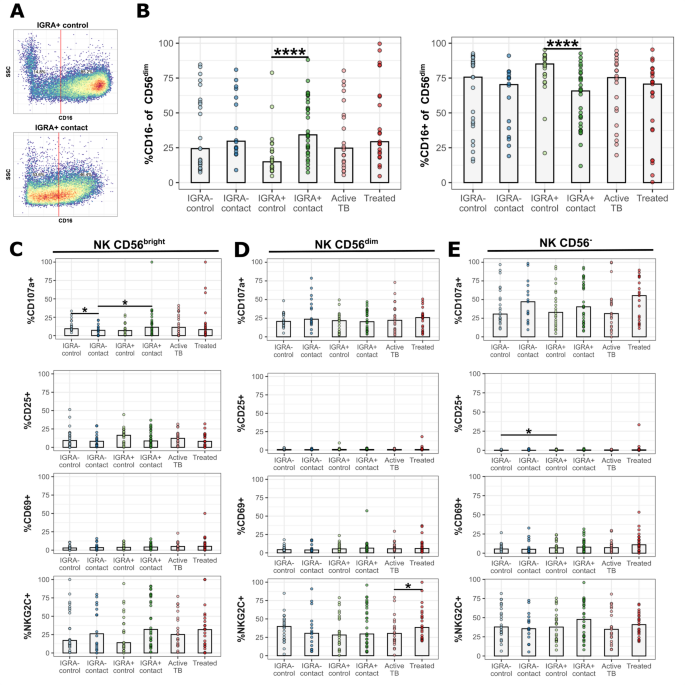
NK cell activity against Mtb depends upon complex combinatory signaling transduced by activating and inhibiting receptors42,43,44. We next investigated markers of NK cell activation ex vivo. We analyzed representative receptors, including CD16a (FcγRIIIA), which binds the Fc region of antibodies and can induce antibody-dependent cellular cytotoxicity (ADCC). Within the major CD56dim subset, IGRA+ contacts had decreased circulating CD16+ NK cells compared to IGRA+ controls (60% vs. 80%, p < 0.0001) with a reciprocal increase in CD16− population, representing downregulation or shedding of CD16 surface expression in IGRA+ household contacts (Fig. 2A, B)45. We found no remarkable differences in canonical markers of NK cell activation ex vivo including CD69 and CD25 (Fig. 2C–E). We observed decreased staining for degranulation marker CD107a in CD56bright NK cells of IGRA− contacts, potentially representing recent activation events during household exposure (Fig. 2C). Expression of the HLA-E restricted receptor NKG2C significantly increased in active TB cases that completed treatment relative to donors actively receiving anti-TB therapy (Fig. 2D).
NK cells demonstrate loss of CD16 expression in IGRA+ contacts. (A) Representative flow plots of CD16 gating in IGRA+ contact and IGRA+ control. (B) Frequency of CD16+ in CD56dim NK cells. Brackets represent Wilcoxon tests with unadjusted p-values. Percent positive staining for four activation markers in (C) CD56bright, (D) CD56dim, and (E) CD56− NK cells. Brackets represent Wilcoxon tests with unadjusted p values. *p ≤ 0.05, ****p ≤ 0.0001.
NK cells of IGRA− contacts have a depressed functional phenotype
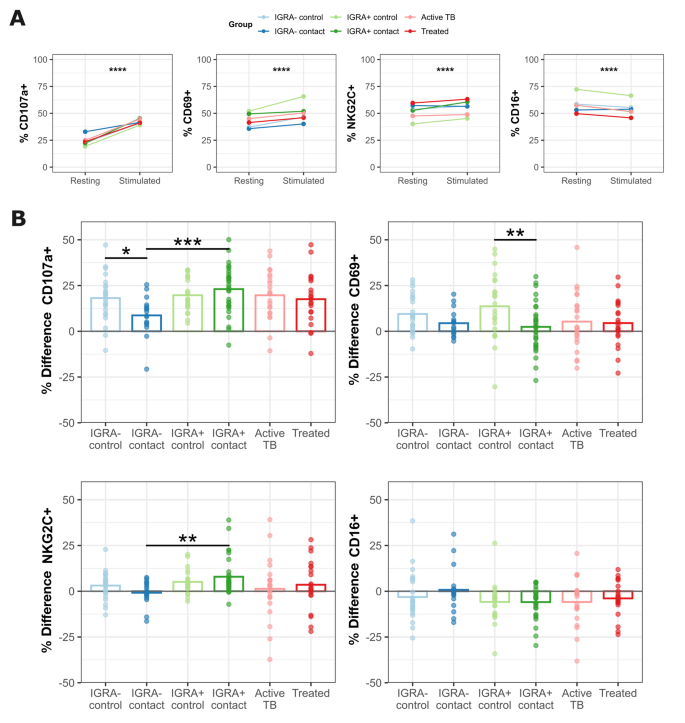
To assess functional activity of NK cells in vitro, we cultured PBMCs in media alone or with K562 cells—a leukemia cell line with low levels of HLA I expression known to activate NK cells—co-incubated at 1:20 K562:PBMC for 15 hours. Cells were then stained, and percent difference was calculated for CD69, CD107a, CD16, and NKG2C (Fig. S1A). Our positive control analyses in Fig. 3A display results from the major CD56dim subset and demonstrate CD107a, CD69, and NKG2C were significantly increased with K562, while CD16 was significantly decreased, across all clinical groups.
Next, we directly compared NK cell response to K562 between clinical groups. IGRA+ contacts demonstrated significantly more degranulation (CD107a) and NKG2C expression after stimulation relative to IGRA− contacts (Fig. 3B, top left and bottom left). IGRA+ contacts had an attenuated CD69 response relative to IGRA+ controls (Fig. 3B, top right). No significant differences in CD16 expression were detected. The same analyses applied to CD56− and CD56bright NK cells demonstrated similar responses to stimulation, but no significant differences between clinical groups, save an attenuated CD69 response in CD56bright NK cells of active TB cases (Fig. S1B–E). Taken together with our ex vivo data, these in vitro restimulation results indicate that CD56dim NK cells of putative Mtb resistors (IGRA− contacts) have depressed function, potentially representing sequelae of household-associated Mtb exposure.
IGRA− contacts demonstrate depressed NK cell function in vitro. PBMCs stimulated overnight by co-culture with K562 cell line compared to plain media alone, gated on CD56dim NK cells. (A) Overall results of activation assay for CD56dim NK cells for four activation markers. Y-axis represents mean percent positive for each activation marker. P values calculated as a global paired Wilcoxon test including all clinical groups. Each clinical group is displayed separately indicated by line color. (B) Percent difference of four activation markers (stimulated minus resting) compared between clinical groups. Brackets represent Wilcoxon tests with unadjusted p values. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
Loss of circulating CD8α+ NK cells during household-associated Mtb infection
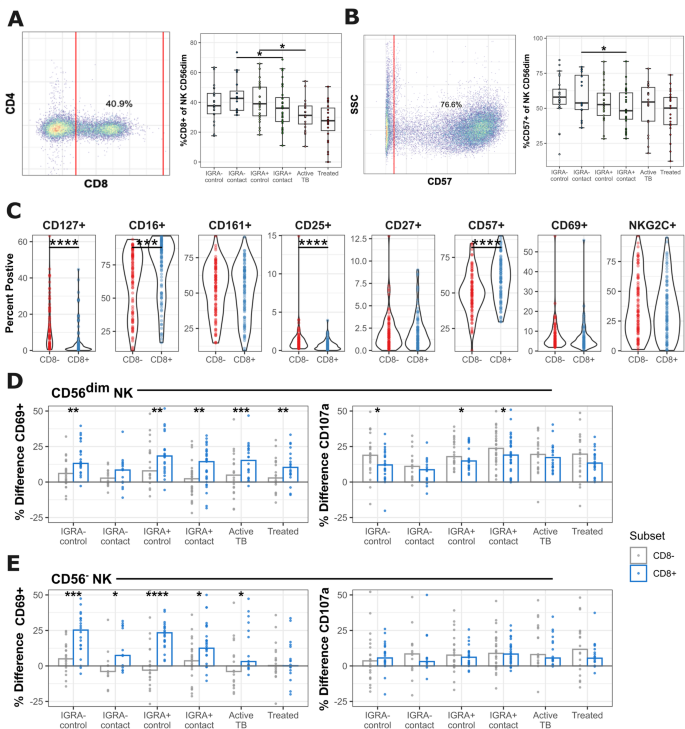
Despite their canonical classification as short-lived innate lymphocytes, NK cells are increasingly recognized for their marked diversity and ability to differentiate into longer-lived effectors19,21,31,46,47,48,49, including after mycobacterial stimuli50. The most extensively studied is an NK cell memory population co-expressing NKG2C and CD57 that expands in blood during human cytomegalovirus (CMV) infection31,51. Multiple putative markers of NK cell memory during Mtb infection have been suggested, including CD57, CD8α, CD161, and CD2732,39,50. We compared expression of these markers across clinical groups and found a loss of CD8α expression in IGRA+ contacts and active TB cases among CD56dim NK cells (Fig. 4A). A similar trend was observed with the marker of terminal differentiation, CD57 and there were significantly fewer CD57+CD56dim NK cells in IGRA+ contacts compared to IGRA− contacts (Fig. 4B). The frequency of CD8α+ T cells was also reduced in IGRA+ contacts relative to controls (Fig. S2A). CD8α+ NK cell frequency increased with age but was not significantly associated with sex (Fig. S2B). CD8α expression among CD56bright NK cells was also significantly decreased in IGRA+ vs. IGRA− controls (Fig. S3A). CD57+CD56bright NK cells expanded in IGRA+ contacts relative to IGRA+ controls (S3A). CD57 expression among CD56− NK cells significantly decreased in TB cases who completed treatment vs. those on treatment (Fig. S3C). We did not observe any significant differences in the expression of CD27, CD127, or CD161 between clinical groups (Fig. S3A-C). Together, these results indicate that donors who develop household-associated immunoreactivity to Mtb antigens or progress to active TB disease demonstrate decreased CD8α surface expression on CD56dim NK cells. In turn, CD8α+ NK cell frequency significantly correlates with resistance to primary household-associated infection as measured by IGRA immunoreactivity.
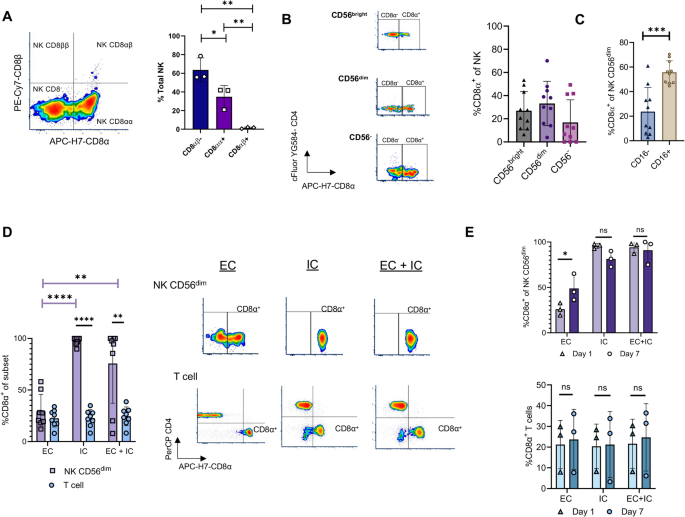
CD8α+ NK cells are depleted in the blood of cases with household-associated Mtb infection. (A) Representative gating and frequency of CD8α+CD56dim NK cells in clinical groups. (B) Representative gating and frequency of CD57+CD56dim NK cells in clinical groups. (C) Expression of select surface markers in CD8α+ vs. CD8α− CD56dim NK cells of healthy donors, with active TB excluded. (D) Activation assay with overnight K562 co-culture versus no K562 comparing CD8α+ vs. CD8α− CD56dim NK cells. Y-axis displays the percent difference for CD69 and CD107a. Asterisks represent paired Wilcoxon tests. (E) As in panel (D), instead displaying CD56− NK cells. P values represent unadjusted Wilcoxon testing. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
CD8α+ NK cells represent a mature population with high functional potential
Despite the identification of cytotoxic CD8α+ NK cells over 30 years ago52,53,54, the function of CD8α on NK cells during health and disease is not well-characterized and is of emerging interest as a cellular target against immune-mediated diseases52,54,55,56. We next used flow cytometry to define co-expressed receptors of CD8α+ NK cells in our cohorts. In the Haitian cohorts, a mean of 27% of NK cells expressed CD8α on their surface. Among NK cell subsets, CD56dim NK cells had the largest proportion of CD8 positivity (median: 36%, range: 1–74%) (Fig. S4A). We then defined co-expressed receptors on CD8α+ NK cells among all healthy Haitian donors, excluding active TB cases with chronic symptomatic disease. CD8α+ NK cells expressed significantly higher levels of CD57 and CD16, and lower levels of CD25 and CD127 (Fig. 4C). This indicates that CD8α+ NK cells represent more mature effectors compared to their CD8α− counterparts. When compared to T cells, median fluorescence intensity (MFI) of CD8α surface expression was significantly lower in NK cells (Fig. S4B).
To compare CD8α+/− NK cell function, PBMCs from the Haitian cohorts were co-incubated with K562 cells for 15 hours and compared to resting conditions as described above. CD8α+ NK cells were more responsive to K562 induction measured by CD69 upregulation compared to CD8α− NK cells but demonstrated lower levels of degranulation as measured by CD107a staining (Fig. 4D). The propensity for CD8α+ NK cells to upregulate CD69 upon K562 stimulation was more pronounced among the CD56− compartment, where the CD8α− fraction had little to no response (Fig. 4E). Overall, our data suggest that CD8α expression among NK cells marks a population with higher activation potential, specifically among terminally differentiated CD56− NK cells, that are reported to have depressed effector functions57.
CD8α protein is ubiquitously expressed in NK cells
We next interrogated the immune phenotype and function of CD8α+ NK cells in a distinct biorepository of healthy human PBMCs (IRB2021-00478; PI: Vorkas) to validate our findings and perform additional functional assays. First, we confirmed the presence of CD8α expression in NK cells by extracellular staining with CD8α and CD8β antibodies. About 60% of NK cells did not express surface CD8, CD8αα was found in ~ 40% of NK cells, while rare CD8αβ NK cells were detected (Fig. 5A). In blood, CD8α was expressed in all three compartments of NK cells, and most frequently among CD56dim cells, with median expression comparable to Haitian donors (Fig. 5B). We also validated our prior result from the contact cohort (Fig. 4C) demonstrating CD16 and CD8α were significantly co-expressed (Fig. 5C). As our TB contact studies only assessed surface CD8α on NK cells, we next analyzed intracellular CD8α expression by flow cytometry. We observed that nearly all CD56dim NK cells expressed CD8α protein intracellularly, contrasting with T cells whose CD8 expression did not significantly differ between surface and intracellular staining (Fig. 5D). CD56bright and CD56− NK cells also demonstrated ubiquitous expression of CD8α protein intracellularly (Fig. S5). To test the stability of surface CD8α on NK cells over time, we incubated PBMCs for 1 or 7 days in pro-survival cytokine IL-2. We observed significantly increased CD8α surface expression following prolonged incubation, indicating that surface presentation of NK cell CD8α may be more dynamic than in T cells where it is a constitutively expressed surface receptor (Fig. 5E).
CD8α+ NK cells are prevalent in healthy US donors with ubiquitous intracellular CD8α expression. (A) Flow cytometry plot and percentage of CD8αα−, CD8αα+ and CD8αβ+ among total NK cells (n = 3). (B) Percentage of CD8α+ NK cells in different subsets based on CD56 expression (n = 10). (C) Frequency of CD16 expression in CD8α+CD56dim NK cells. (D) Frequency and representative flow cytometry plots of CD8α expression comparing CD56dim NK cells and T cells with extracellular (EC), IC (Intracellular), and EC+IC CD8α staining. (E) Percent CD8α+ after EC, IC, and EC+IC staining at day 1 and day 7 in CD56dim NK cells and T cells. Statistical significance was measured using unpaired t tests, ns—no significance, *p < 0.05, **p < 0.005, ***p < 0.001, ****p < 0.0001.
CD8α+ NK cells have enhanced responses to cytokine stimulation
We next hypothesized that surface CD8α was associated with enhanced responsiveness to inflammatory signals, following prior reports of increased cytotoxicity against leukemic cells relative to CD8α− NK cells53,56. We stimulated PBMCs with either the potent IL15-IL15Rα sushi domain fusion construct (IL15-15RαSu)58, IL15-15RαSu/IL12/IL18, IL12/IL18 alone, or virulent Mtb Erdman whole cell lysate and assessed intracellular IFNγ expression by flow cytometry. Cytokine stimulation led to decreased CD8α+ surface staining while Mtb stimulation did not alter CD8 surface expression, suggesting that cytokine stimulation induces downregulation of the CD8α receptor or cell death (Fig. 6A). Both IL15/IL12/IL18 and to a lesser degree Mtb lysate conditions induced IFNγ responses in CD56dim NK cells compared to resting (Fig. 6B). When stratified by CD8α, we observed increased intracellular IFNγ with cytokine stimulation in both CD16−/+ CD8α+ NK cells relative to CD8α− cells, but no differences upon co-incubation with Mtb lysates (Fig. 6C). Responses were more robust among the CD8α+CD16− NK cell population, likely representing highly activated cells downregulating the Fc Receptor. Percentage of CD25, CD69, and intracellular Granzyme B responses were also enhanced in cytokine-induced CD8α+ NK cells across CD56-stratified subsets (Fig. S6).
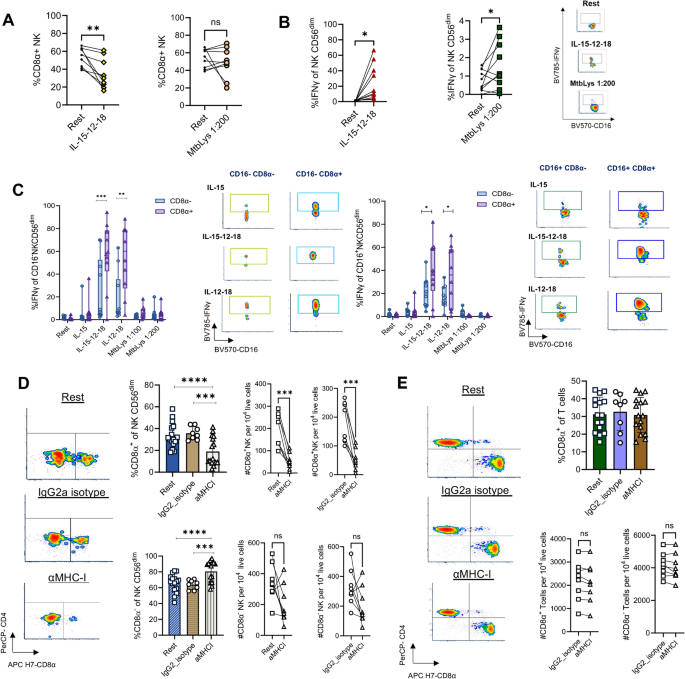
CD8α+ CD56dim NK cells have enhanced response to cytokine stimulation. (A) Pairwise analysis of percent CD8α+ NK cells upon stimulation with IL15/12/18 and MtbLys 1:200 relative to resting condition (n = 9). (B) Pairwise analysis and representative flow plots of intracellular IFNγ in CD56dim NK cells after stimulation with IL15/12/18 and MtbLys at dilution 1:200 relative to resting condition (n = 9). (C) Percent intracellular IFNγ+ in CD8α+ (triangles) versus CD8α− (circles) CD56dim NK cells comparing IL-15-Su, IL15-Su/12/18, IL-12/18, or MtbLys conditions (n = 9). Both the CD16+ (right) and CD16− (left) CD56dim NK cell subpopulations are displayed. (D) Representative flow cytometry plots, with %CD8α+ (top) and %CD8α−(bottom) of CD56dim NK cells alongside absolute count per 104 live cells after anti-MHC class I blockade compared to no antibody (rest) or IgG2a isotype controls. Statistical significance was measured using paired t-test, with threshold of *p < 0.05, **p < 0.005, ***p < 0.001. (E) Same experimental setup as D, gated on CD8+ T cells. IFNγ Interferon γ, MtbLys Mtb whole cell lysate.
CD8α surface expression on NK cells is MHC class I-dependent
To determine a mechanism by which CD8α may modulate NK cell function, we hypothesized that CD8α on NK cells engages MHC class I molecules like CD8+ T cells59,60. To test this, we incubated whole PBMCs with MHC class I blocking antibody (clone: W6/32) compared to IgG2 isotype and no antibody controls and measured surface CD8α expression. Strikingly, surface CD8α expression on NK cells was significantly reduced after MHC class I blockade (Fig. 6D). As MHC class I blockade may activate NK cells due to loss of inhibitory Killer-like immunoglobulin Receptor (KIR) engagement and potentially induce apoptosis, we measured frequency and CD107a staining to assess degranulation of CD8α+/− NK cells during MHC class I blockade. We found that MHC class I blockade led to a significant decrease in the frequency and absolute numbers of total NK cells (Fig. S7A), accompanied by a significant increase in CD107a staining among both CD8α+/− NK cells (Fig. S7B). These effects were unique to MHC class I blockade conditions and not observed with isotype control antibodies. Blockade was further validated by loss of surface staining of anti-HLA I antibodies with MHC class I blockade, but not isotype control (Fig. S7C). NK cell depletion with anti-MHC class I was explained by absolute loss of CD8α-expressing NK cells (Fig. 6D), without a consistent reciprocal increase in CD8α− absolute numbers (Fig. 6D). These results may represent preferential induction of CD8α+ NK cell death or CD8α surface downregulation after MHC class I blockade. Importantly, CD8α surface expression on CD8+ T cells was not affected (Fig. 6E). Together, our results indicate that MHC class I engagement of NK cells may stabilize CD8α surface expression.
CD8α+ NK cells have overlapping transcriptomes with CD8α− NK cells
The transcriptional programs associated with surface expression of CD8α in NK cells and whether these relate to homologous functions observed in CD8α+ T cells during disease is an area of emerging interest52,53,54,56,61. To interrogate the transcriptional profile of CD8α+/− NK cells, we sorted innate lymphocyte subsets from PBMCs including NK cells, γδ T cells, Mucosal-associated invariant T (MAIT) cells, and invariant Natural Killer T cells (iNKTs) by fluorescence-activated cell sorting (FACS) from two healthy donors. We performed cellular indexing of transcriptomes and epitopes single cell sequencing (CITE-seq) of these populations directly after sorting or after PBMC coincubation for seven days in IL2 supplemented media +/− Mtb lysates to define surface protein and transcript expression in CD8α+/− NK cells. We discriminated NK cells from surrounding innate lymphocytes using surface antibody barcodes for CD56 and CD16 together with expression of canonical NK cell gene transcripts CD3E, NCR1, and NCAM1 (CD56) (Fig. S8)52,62,63. Dimensionality reduction with UMAP identified four clusters of NK cells with distinct transcriptional signatures (Fig. 7A–D). To assess pathogen-specific responses, we first performed differential expression between the ‘rest’ and ‘Mtb lysate’ conditions and display the top eight most differentially expressed genes in Fig. 7E. The cytotoxic serine protease GZMB (Granzyme B) and killer immunoglobulin-like receptor, KIR3DL1, were the most upregulated genes in the Mtb lysate condition.
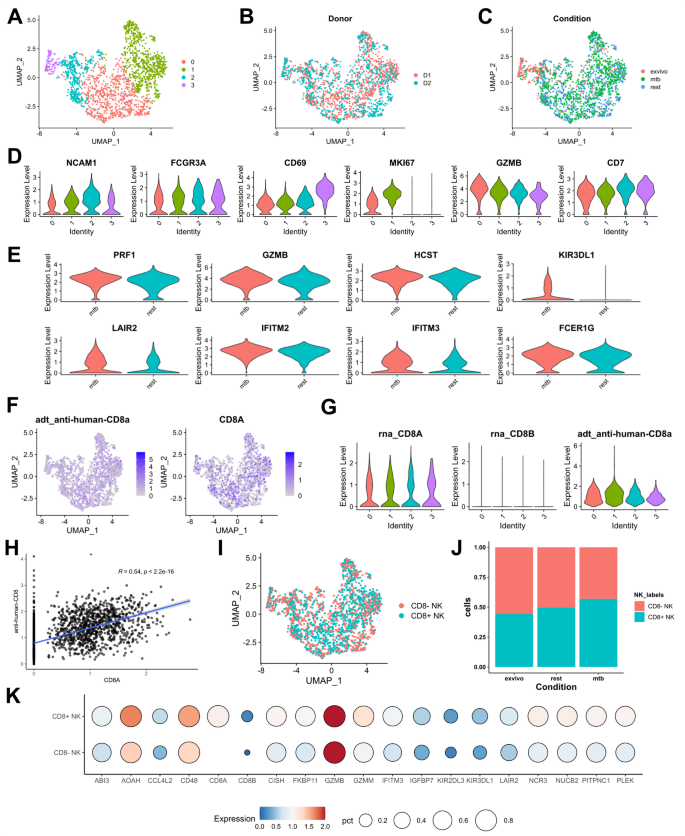
CITE-seq analysis of CD8α+ vs. CD8α− NK cells from two healthy donors reveals overlapping transcriptomes. (A) Results of clustering and uniform manifold approximation (UMAP) projection for all NK cells. (B) UMAP reduction colored by Donor. (C) UMAP reduction colored by condition. (D) Violin plots representing top genes that were differentially expressed between the four NK cell clusters. (E) Violin plots of the eight differentially expressed genes between resting and Mtb lysate condition. (F) Distribution of NK cell expression of CD8 using barcoded antibodies against CD8α protein and CD8A transcript. (G) Violin plot of the expression of CD8A and CD8B transcripts, as well as CD8α surface expression in the four NK cell clusters. (H) Correlation of CD8α expression between transcript and protein measurement. (I) Annotated UMAP showing CD8α+ and CD8α− NK cells based on CD8A transcript expression. (J) Percent of NK cells expressing CD8A transcript in stimulation conditions. (K) Dot plot of genes differentially expressed in CD8A+/− NK cells.
We next analyzed the distribution of CD8A transcript and CD8α protein expression in NK cells using both transcript and surface antibody barcodes for CD8α. NK cells did not cluster independently based on the presence of CD8A/CD8α (Fig. 7F). The expression of the CD8A gene and CD8α surface protein was distributed throughout all NK cell clusters and a positive correlation was observed between protein and transcript expression (Fig. 7G–I). Low amplification of surface antibody barcodes for CD8α was observed in NK cells, in contrast with other CD8α-expressing immune subsets sequenced (e.g., MAIT cells; Fig. S9) consistent with CD8α MFI differences detected by flow cytometry (Fig. S4B).
To define transcriptional signatures associated with CD8A transcript expression, we stratified NK cells expressing greater than zero CD8A transcripts and performed differential expression relative to CD8A− cells. Half of all NK cells expressed CD8A transcripts (Fig. 7J). Differential gene expression revealed few statistically significant differences between CD8A+/− cells. Notable differentially expressed genes in CD8A+ NK cells include cytotoxicity-associated proteins natural cytotoxicity receptor 3 (NCR3) and the serine protease granzyme M (GZMM) (Fig. 7K).