Protein profile of selected OPXV genus viruses
The proteins responsible for entry (A17L, A28L, A33R, H2R, L1R), exit (A27L, A35R, A36R, C19L), and both (B5R) of viruses from the OPXV genus were obtained in FASTA format from the NCBI Virus database. The “taxid:10242”, which covers all organisms of this genus, was selected. A total of 16 virus species were identified, including Orthopoxvirus Abatino, Akhmeta virus, Borealpox virus, Buffalopox virus, Camelpox virus, Cowpox virus, Ectromelia virus, Horsepox virus, Monkeypox virus, Rabbitpox virus, Racoonpox virus, Skunkpox virus, Taterapox virus, Vaccinia virus, Variola virus, and Volepox virus.
Among these, nine have RefSeq for all studied proteins, four have RefSeq for nine of the ten proteins, necessitating the generation of a consensus for one protein. Three viruses do not have reference sequences; thus, the available proteins were collected, and a consensus was generated. It is important to note that Rabbitpox virus was the only one with a protein (A27L) found by BLAST, and Mpox virus had two RefSeqs for each protein, corresponding to clades I and II, except for A27L, for which a consensus sequence was generated.
For sequencing of the Mpox clade Ib virus, the PQ220056.1 sequencing was used, with the respective proteins aligned and identified. The A27L protein was not found and thus was marked as absent. All data are detailed in Table 1.
Antigenic, non-allergenic, and stable epitopes identified in conserved regions of eight proteins
The IEDB server was used to find epitopes that can bind to MHC-I, MHC-II, and BCR, with all parameters set by the server. A global consensus sequence was generated for each of the ten proteins from each virus. The epitopes were identified, collected, and subjected to antigenicity analysis on the VaxiJen server, and their stability was assessed using the ProtParam tool from the Biopython library. It was noted that there were more epitopes for MHC-I and MHC-II receptors compared to BCR (Table 2).
The obtained epitopes were aligned using Jalview software and the ClustalW tool with the proteins from the sixteen viruses to identify conserved epitopes across all proteins, i.e., conservation regions that had at least one epitope were selected (Table 2).
Proteins A28L and B5R, although having conserved regions, did not show the presence of epitopes in these regions or, when present, the epitopes did not fully occupy these regions, failing to meet the minimum length of 8 amino acids. A total of 26 epitopes were identified and none of them presented evidence of allergenicity using the AllerCatPro 2.0. The amino acid sequences of each conserved epitope along with their respective antigenicity values are presented in Table 3. It is worth noting that the identified conserved epitopes are also present in the new Mpox variant, clade Ib. Although proteins A17L, A36R, C19L, and L1R have mutations in their protein composition, the identified epitopes remain conserved in this new variant (Fig. 1).
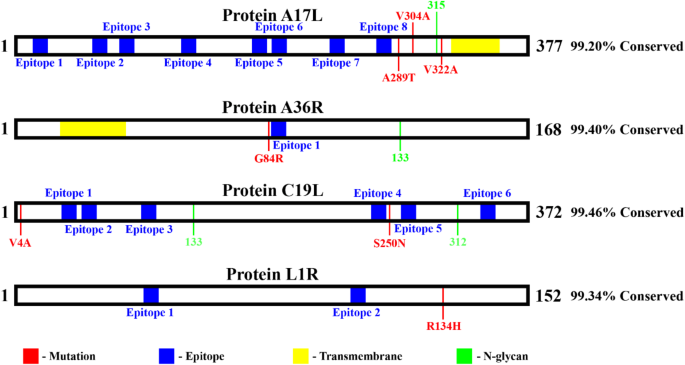
Analysis of the A17L, A36R, C19L, and L1R proteins from the new Mpox virus lineage (Clade Ib), sequenced in 2024. The conservation percentages of these proteins compared to Clade I are indicated to the right of each protein. Conserved epitopes are highlighted in blue, mutations in red, transmembrane regions in yellow, and N-glycosylation sites in green. (A) Protein A17L with 99.20% conservation, displaying multiple conserved epitopes but with mutations outside these regions. (B) Protein A36R with 99.40% conservation, showing one mutation and one conserved epitope. (C) Protein C19L with 99.46% conservation, revealing several mutations and conserved epitopes. (D) Protein L1R with 99.34% conservation, having two conserved epitopes and one mutation outside these regions. The identified mutations do not affect the conserved epitopes, suggesting that they remain preserved in this new virus lineage.
The new Mpox sublineage, declared a global emergency by the WHO due to the evolution of Clade Ib, has demonstrated increased transmissibility and the ability to spread to other continents3. Therefore, it is crucial to verify if conserved epitopes remain preserved in this new variant. Sequencing conducted in 2024 on patients from the Democratic Republic of Congo regarding the new Clade Ib lineage is available on NCBI under code PQ220056.125, and it was used for analysis. The A27L protein was not found, which is consistent with the results in Table 1, where A27L is the only analyzed Mpox protein without a RefSeq. Additionally, among the eight proteins with conserved epitopes, A33R, A35R, and H2R are 100% conserved, ensuring epitope conservation (Fig. 1A). The A17L, A36R, C19L, and L1R proteins show 99.20%, 99.40%, 99.46%, and 99.34% conservation, respectively, compared to Mpox Clade I proteins; however, mutations do not occur in the identified epitopes (Fig. 1B–D).
Characterization and evaluation of the constructed multi-epitope chimeric proteins indicate adequate physicochemical properties, absence of allergenicity, good antigenicity, and positive humoral and cellular immune responses
After identifying the epitopes for each protein, four multi-epitope chimeric proteins were constructed. Two adjuvants were selected and added to the beginning of the chimeric protein: PADRE, a synthetic epitope capable of binding to MHC-II, which enhances the immune response by increasing the activation and proliferation of helper T cells, and β-defensin, which stimulates both the innate and adaptive immune responses12. It is worth noting that the chimeric proteins related to A17L, C19L, and H2R were created individually as they were the only ones presenting more than one conserved epitope.
The chimeric protein constructs range from 146 to 173 amino acids, molecular weights varying from 16 to 19 kDa, and isoelectric points of approximately 9, indicating a basic nature for all proteins. Furthermore, all proteins meet the cut-off thresholds for stability, antigenicity, allergenicity, solubility, and autoimmunity, indicating that they are suitable for vaccination (Table 4).
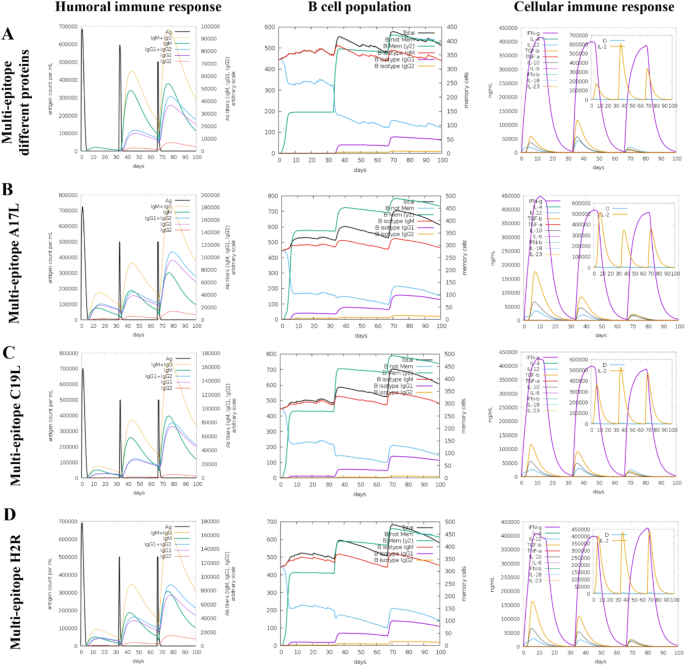
Subsequently, the simulation of the possible immune response to the four vaccine candidates was conducted on the C-ImmSim server. The multi-epitope protein containing epitopes from different proteins exhibited lower IgM + IgG titers compared to the vaccine candidates based on individual proteins (Fig. 2A). Additionally, the population of memory B cells was lower in the vaccine candidate containing epitopes from different proteins during the first 30 days, with an increase observed after the second dose administration. Comparing the cellular response, IFN-γ cytokine production was similar among the four vaccine candidates, while variations were observed in the levels of IL-2, TGF-β, IL-10, and IL-12 (Fig. 2A–D). All four vaccine candidates showed nearly null values for the “Danger” variable (Fig. 2A–D).
The constructed multiepitope chimeric proteins show satisfactory predictions for cellular and humoral responses after three doses. Cytokine and interleukin levels after 3 doses are displayed, with “D” in the graph indicating a potential danger signal. (A) Results for the chimeric protein containing epitopes from different proteins. (B) Protein with epitopes from A17L. (C) Protein with epitopes from C19L. (D) Protein with epitopes from H2R.
Structural stability and interaction with the TLR-2 receptor of the multivalent chimeric protein: modeling and molecular dynamics reveal good stability and fluctuation patterns
To determine the three-dimensional structure of the vaccine candidates and assess their interaction capability with the TLR-2 receptor, the AlphaFold 3 server was used. The predicted template modeling (pTM) score, interface predicted template modeling (ipTM) score, and per-residue measure of local confidence (pLDDT) were evaluated for each vaccine candidate. The modeling showed similarities, with TLR-2 exhibiting high pLDDT values, while the vaccine candidates showing low values. It was observed that the adjuvant region is responsible for anchoring to TLR-2 (Fig. 3A–D). Additionally, the pTM values ranged between 0.70 and 0.73, suggesting that the overall folds are close to the true structure, as values above 0.5 indicate good correspondence. However, ipTM values ranged from 0.20 to 0.22, indicating a potential failure in predicting the interaction. Therefore, validation of the structures through molecular dynamics was necessary.
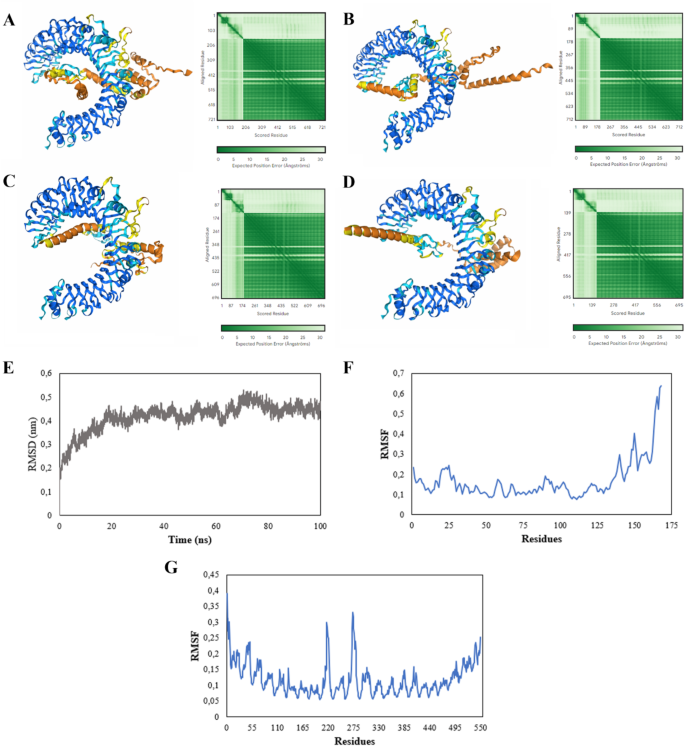
The constructed chimeric proteins interact with the Toll-like receptor 2 (TLR-2), with modeling indicating good results for the receptor. However, the multiepitope proteins show low reliability. The multiepitope chimeric protein from different proteins demonstrates stability after 20 nanoseconds, maintaining a good interaction with TLR-2. (A) Modeling and binding of the multiepitope protein with TLR-2. (B) Binding of the A17L epitope protein with TLR-2. (C) Multiepitope from C19L and TLR-2. (D) Multiepitope from H2R and TLR-2. (E) RMSD obtained after 100 ns of molecular dynamics of the TLR-2 complex with the multiepitope protein. (F) RMSF of the multiepitope protein. (G) RMSF of TLR-2.
Molecular dynamics was performed exclusively with the chimeric protein containing epitopes from different proteins due to its potential representativeness and comprehensiveness for immunization against the OPXV genus. Structural stability over time was assessed using the Root Mean Square Deviation (RMSD) graph, and the protein’s residual fluctuation was analyzed with the Root Mean Square Fluctuation (RMSF) graph. The RMSD graph showed that the protein reached stability after 20 ns and remained stable until the end of the simulation (100 ns), with a structural variation from 0.4 nm to 0.5 nm (Fig. 3E). The RMSF graph indicated low variation in the initial part of the chimeric protein, with an increase in fluctuation occurring in the last amino acids of the protein (Fig. 3F). In contrast, the TLR-2 receptor exhibited a characteristic fluctuation pattern, with two fluctuation peaks occurring between amino acids 200 and 300 (Fig. 3G).
Additionally, analyses of hydrogen bonds (H-bonds) and the radius of gyration (Rg) were conducted, as presented in Supplementary Fig. S1. The data show that the interaction complex between the TLR-2 receptor and the multi-epitope protein, composed of epitopes from different proteins, maintained between 450 and 550 hydrogen bonds throughout the simulation (Supplementary Fig. S1A). The radius of gyration ranged from 3.00 to 3.14 nm (Supplementary Fig. S1B), corroborating the RMSD values presented in Fig. 3E. These results indicate the preservation of structural integrity and stability of amino acid interactions during the 100 ns simulation, demonstrating the structural viability of the complex.