Epidemiological Analysis of dengue virus infections relative to the climate-driven index
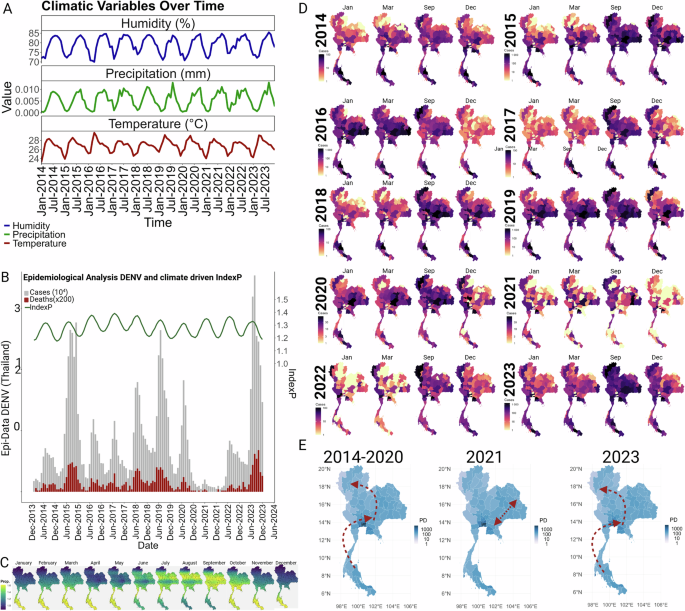
Before exploring the factors that determine dengue dynamics in Thailand, we first describe the magnitude of its burden and its typical yearly pattern. An analysis of dengue fever (DF) epidemiological trends in Thailand from 2014 to 2023 demonstrates a clear seasonality in case numbers, with yearly peaks coinciding with the rainy season and the periodicity of Index P, a climate-driven variable that reflects mosquito availability (Fig. 1A, B). Particularly high case counts were noted in 2015 and 2019, along with a significant decline in 2021. However, by 2023, the case numbers had surged to a maximum (Fig. 1B). Over the same 10-year period, deaths attributed to dengue fever were consistently low (0.1%). Indeed, mortality trends paralleled incidence rates, indicating that fluctuations in number of deaths were more indicative of changes in virus transmission (e.g. proportional to case counts) rather than alterations in clinical severity (Fig. 1B).
A Climatic variables (precipitation, humidity and temperature) per month obtained for Thailand from 2014 to 2023 were used as priors to estimate index. B Temporal trends and climatic influences on dengue virus transmission represented by monthly reported cases and fatalities associated with DENV in Thailand, alongside with a mosquito-viral suitability index (Index P). Index P integrates climatic data, anthropogenic factors, and the prevalence of the primary vector, Aedes aegypti. This index is computed monthly and averaged nationally, reflecting the combined impact of environmental and human variables on vector capacity and virus transmission (for detailed methodology see Materials and Methods and Supplementary Fig. S1). C Spatiotemporal patterns of mosquito-viral suitability across Thailand showed by the average monthly values of Index P from 2014 to 2023 elucidating the geographical variation in vector suitability, highlighting regions and times of heightened risk. For detailed monthly/year maps see animations provided in Supplementary Video 1. D Spatiotemporal distribution of DENV cases. Delineation of temporal dynamics of DENV transmission, identifying the geographic distribution of the cases per season considering the onset of the transmission in January, an initial peak in March, the highest peak in September, and the season’s end in December (determined in panel A). The monthly mapped cases are available in Supplementary Video 2. E Synthesis of case trajectories place alongside with population density, offering insights into the spatial clustering of outbreaks, red arrows illustrate the directional progression of the cases over the specified period. The directional progression of cases were derived from statistically determined central positions (centroids) of case distributions using kernel density estimation (KDE) across successive time periods.
IndexP, a novel suitability index based on a climate-driven mathematical expression for the basic reproductive number of mosquitoes-borne viruses20. Index P has been previously used to estimate mosquito population dynamics and virus–mosquito transmission efficiency for dengue20, yellow fever virus21 and zika virus22 in Brazil, as well as west nile virus23 in Israel. In the current study Index P was calculated from several factors, including 1) climate: temperature, precipitation, humidity, 2) mosquito prevalence: rate of bites, mating, number of females, and 3) human susceptibility: number of bites * life expectancy. Thailand exhibits a distinct IndexP pattern, characterized by a gradual migratory shift from southern regions toward the north and back to the south, corresponding to peaks in March and September, respectively. During the peak period spanning July-October, summer conditions are most conducive to mosquito breeding and survival and create heightened potential for dengue transmission. As the year progresses, index values recede in the north to signal a decrease in abundance of mosquito populations with the higher values in the southern regions for the months of November through May. This cyclical movement of mosquito populations24 suggests a strong seasonal component driven by climatic factors that undoubtedly drives dengue transmission (Fig. 1C and Supplementary Video 1).
While this spatiotemporal analysis seeks to approximate the relationship between ecological transitions and mosquito prevalence and has the potential to identify hotspots and periods of intensified dengue outbreak risk, it does not singularly account for the observed reduction in dengue cases in 2021 across Thailand. Additional factors may have contributed to this outcome: herd immunity, vector control initiatives, public health interventions, or changes in human behavior (e.g. reduced contact between human and mosquito due to covid lockdowns). To determine whether any of these anthropogenic factors influence infection trends in Thailand, we conducted a detailed, yearly analysis of the geospatial distribution of dengue cases from 2014 to 2023 and related this to population density (Fig. 1D–E and Supplementary. Video 2). During the 2014-2020 period, patterns mirrored that of the typical IndexP cycle, migrating from the south to more densely populated central and northern provinces where the most significant aggregation of dengue cases were found. Notably, in 2021, the distribution of dengue cases was largely confined to the western regions with fewer inhabitants, and notably absent from the most populated urban areas. This unexpected containment pattern suggests that partial immunity may have offered protection against widespread transmission common to the central and northern provinces. By 2023 the pattern of distribution of cases have been restored (Fig. 1E).
Dynamics and temporal diversification of dengue serotypes in Thailand
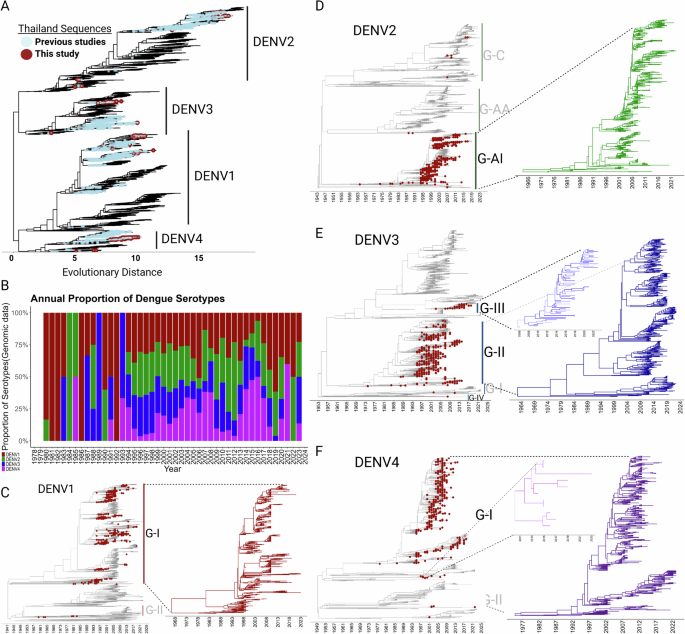
Understanding levels of genetic diversity and estimating trajectories will indicate which serotypes or lineages therein are expanding whereas others could be experiencing an evolutionary bottleneck that could explain the fluctuations in the number of DENV cases in the country. To investigate the genetic diversity of DENV in Thailand, we conducted a Maximum Likelihood (ML) phylogenetic analysis utilizing all available, nearly complete genomic sequences ( > 70% coverage) from across the globe present in public databases. We enriched this dataset by adding approximately 200 full genome sequences from Thailand obtained in this study by metagenomic NGS and viral target enrichment (CVRP), which includes representatives from all four serotypes. This comprehensive analysis aimed to delineate the evolutionary relationships and track the potential emergence of new viral variants over time in the country. Our phylogenetic analysis confirmed the endemic circulation of all four DENV serotypes within Thailand (Fig. 2A). Contrary to the findings suggested by Poltep et al.25, which used partial sequences from envelope compared to our full genome analysis, we did not detect a high level of intra-serotype diversity among Thai strains. Rather, we observed the formation of homogenous clusters within the serotypes, which were composed of sequences from both previous studies (blue) and those characterized in our current research (maroon) submitted for this study. This phylogenetic pattern strongly suggests that the DENV landscape in Thailand is defined by stable, endemic circulation with sporadic introductions rather than by widespread genetic variation. The clusters formed by previous Thai sequences and reinforced by the addition of our current data containing strains from the same lineages, suggest a relatively constrained evolutionary dynamic within the country (Fig. 2A).
A Phylogenetic reconstruction of dengue Virus. Maximum likelihood (ML) phylogenetic tree, constructed from all sequences in Dataset A (see Materials and Methods). The tree illustrates the presence of all four DENV serotypes (DENV1-4) in Thailand, revealing patterns of diversification and cluster formation that suggest endemic circulation. Each serotype is denoted in the tree, and sequences from both previous studies and the current study are highlighted. B Serotype distribution and prevalence over time. This panel displays the distribution and frequency of the four dengue serotypes in Thailand from 1980 to 2023. It highlights the temporal shifts in serotype dominance and prevalence, reflecting evolving transmission dynamics, vector-host interactions, and population immunity levels. C–F Time-stamped phylogenies represented by MCC-tree for all four DENV serotypes. Sequences from Thailand are highlighting with red tips. Genotypes within each serotype that include sequences from Thailand are denoted (the distribution of all genotypes within each serotype please see Supplementary Material Fig. S1). Insets show detailed views of the endemic lineages in Thailand across the years to highlight the different evolutionary trajectories. Lineages that didn’t share the same Thai ancestor were treated as distinct, and sequences without further diversification were considered external introductions (confirmed in Fig. 4).
Our longitudinal analysis of the annual proportion of dengue serotypes (Fig. 2B), revealed a highly dynamic pattern in serotype prevalence over a 45-year span. The co-circulation of serotypes appears to be an established pattern in the country and raises the specter of DENV co-infections, which could influence disease manifestation and epidemic behavior. The early 1980s were marked by the predominance of DENV1, nearly vanishing in subsequent years, and then resurging and persisting at ~25% from the mid-1990s onwards. From 2016 onward, it has regained its status as the most prevalent serotype. DENV2 also exhibited epochs of dominance (mid-1980s to early 1990s) and absence (1991 to 1993), but it too maintained its presence at steady levels for 25 years (1995–2020). However, no genomic data for DENV2 was identified in 2021which should indicate a reduced circulation for this serotype since it reappears in 2022. DENV3 circulation was sporadic throughout the 80’s, with several years of absence and other years where it appears to have been the only serotype present. Over the period of 1994-2018, yearly prevalence steadied at 10–25%, after which DENV3 seems to have largely disappeared (2021–2022) but reemerging in 2023. DENV4 showed a similar pattern as DENV3, characterized by occasional peaks during the 80’s, consistent levels (5–50%) during the 1992-2018 period, and sporadic levels in recent years (Fig. 2B).
Time-calibrated phylogenetic trees further illustrate the concurrent circulation and evolutionary history of all four DENV serotypes in Thailand (Fig. 2C–F). Thai sequences, denoted by red dots on the trees, formed distinct clusters that signify periods of localized evolution, suggesting a scenario where endemic diversification has occurred. However, the distribution of these sequences could either indicate there were multiple introductions of DENV into Thailand or represent their persistent endemic presence punctuated by phases of expansion. The formation of discrete sub-clades within principal clades indicates there are lineage-specific attributes that potentially impact transmission and pathogenicity (Fig. 2C–F). In the case of DENV1, there are 5 genotypes globally, but only two are present in Thailand. Most sequences belong to Genotype I (GI), with a smaller representation from Genotype II (GII) (Fig. 2C). In GI some clades persisted for decades whereas others went extinct after a short period of time. Genotype II (GII) sequences appear to have been introduced between 1961 to 1984 but failed to diversify within Thailand, as evidenced by the absence of emerging lineages from these sequences (Fig. 2C). A notable feature within the DENV1 GI strains is the ancestral composition of Thai sequences within each cluster, supporting the hypothesis that diversification of these clades occurred primarily within Thailand and lead to their outward spread to other regions. To test this hypothesis, we constructed a temporal phylogeny to scrutinize the emergence and demographic expansion of this genotype in the country dating its emergence back to 1968 (Fig. 2C, inset). For DENV2, sequences were almost exclusively grouped into the Asian I (G-AI) genotype, with a smaller subset corresponding to introductions from the Asian-American (G-AA) and Cosmopolitan (G-C) genotypes, which did not show further diversification (Fig. 2D). Like DENV1-GI strains, the DENV2 G-AI strains constitute a monophyletic clade that emerged ~1966 (Fig. 2D, Inset). By contrast, DENV3’s genetic landscape in Thailand was characterized by the diversification of two primary lineages. Genotype II sequences have been established since 1977, whereas a secondary, smaller lineage of Genotype III sequences was introduced more recently in 2006, alongside Genotype IV strains that displayed limited diversification (Fig. 2E). DENV4 temporal phylogeny mirrors that of DENV1 and DENV2, with a predominant monophyletic cluster of Genotype I sequences circulating since 1977. A secondary, genetically distinct cluster within the same genotype emerged or was introduced in 2011. Additionally, sporadic sequences from Genotype II (GII) appear to have been introduced between 2001 and 2021, yet these did not undergo further diversification within Thailand, as suggested by the static lineage structure (Fig. 2F). It is unclear if these genotypes were introduced to or emerged from Thailand, but as a result, all four serotypes have been present in the country since the late 60’s.
The Bayesian birth-death skyline plot explicitly estimates the rate of transmission, recovery, and sampling allowing the inference of the effective reproductive number (Re) directly from genetic data26. This approach has been extensively used to described fitness competition between lineages in different viral species including SARS-CoV2 in Senegal27, HIV in North America28 and dengue in Brazil29. An estimation of the Re for all six endemic Thai lineages exhibited a Re > 1, accounting for the mean value, (Supplementary Material Fig. S2), indicating that on average, each infected individual is capable of transmitting the disease to more than one other individual. These levels highlight the potential for not only sustained growth, but epidemic growth as well. Notably, the Re values across different lineages remained relatively constant over time, suggesting no discernible differential impact on the transmission dynamics among the lineages within Thailand. This consistency implies that the observed genetic diversity is likely the result of competitive interactions between lineages, rather than attributed to increased fitness or a transmission advantage. In agreement with the notion of an equilibrium having been reached between vectors and hosts is that we observe stable, elevated evolutionary rates for all six lineages (Supplementary Table S1). If one or more lineages had any evolutionary advantage or disadvantage over each other, these values for Re and mutational rates would cease to maintain uniformity. These observations are important for understanding the evolutionary pressures at work and suggests that inter-lineage competition may be a significant factor shaping the patterns of genetic diversity in the DENV lineages circulating in Thailand.
Impact of pervasive and episodic selection on the genetic makeup of dengue serotypes in Thailand
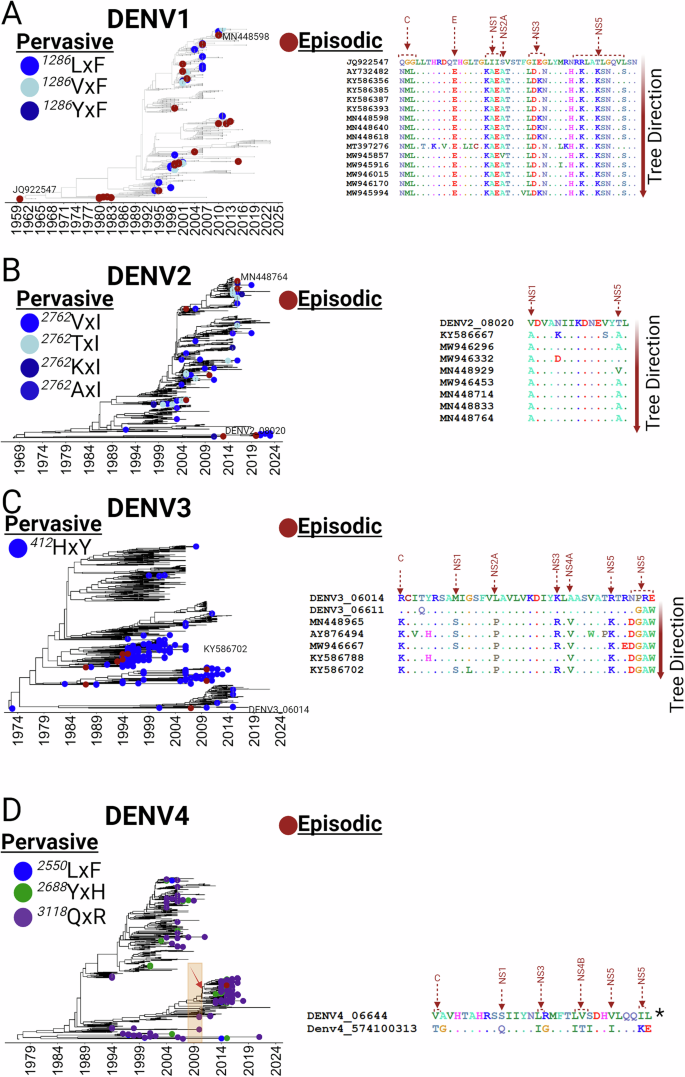
To assess which mutations may have enabled each lineage to compete more effectively, a positive selection analysis was performed. Pervasive evolutionary changes, or those driven by forces acting continuously over time and across populations, were statistically significant at only one position (1286 in NS2A) in DENV1 (Fig. 3A and Supplementary. Data1). This amino acid, as indicated by the FuBAR analysis, has undergone multiple replacements (L1286F, V1286F, or Y1286F) in DENV1 strains that have circulated in Thailand since 1992 (Fig. 3A, central panel). By contrast, an aBSREL analysis for measuring episodic selection, driven by sporadic or distinct events, identified 23 branches that were significantly affected (p ≤ 0.05) (Fig. 3A, central panel, Supplementary. Data2). Subsequent MEME analysis identified 45 sites under episodic selection, with a noticeable preference for nonstructural proteins. Indeed, 13 substitutions in non-structural proteins were maintained over time compared to 4 in structural proteins (Fig. 3A, right panel, Supplementary. Data3 and Supplementary. Data13). For DENV2, the Fubar analysis indicated pervasive selection once again at only one position (2762 in the RdRp; NS5) (Fig. 3B and Supplementary. Data4 and Supplementary. Data13), with replacements V2762I, T2762I, K2762I, or A2762I observed in post-1991 Thai strains (Fig. 3B, central panel). The aBSREL analysis revealed 10 branches under episodic selection (p ≤ 0.05) (Fig. 3B, central panel, Supplementary. Data5) and MEME pinpointed 15 sites under episodic selection, again predominantly in nonstructural proteins, specifically NS1 and NS5 (Fig. 3B, right panel, Supplementary. Data6 and Supplementary. Data13). For DENV3, pervasive selection was evident at position 412 in the envelope protein E (Fig. 3C and Supplementary. Data7 and Supplementary. Data13), leading to the H412Y replacement. This mutation was unique among serotypes for its impact on a structural protein and also unlike other serotypes, both pervasive and episodic selections were restricted to specific sub-lineages (Fig. 3C, central panel, Supplementary. Data8 and Supplementary. Data13). MEME analysis confirmed several sites were under episodic selection, predominantly in nonstructural proteins, except for the positively selected site K104R in the capsid (Fig. 3C, right panel, and Supplementary. Data9 Supplementary. Data13). DENV4 strains exhibited pervasive selection at three positions: 2255 in NS4A, and 2688 and 3118 in NS5 (Fig. 3D and Supplementary. Data10 and Supplementary. Data13). These sites, particularly in RdRp, were widespread across the phylogenetic tree, suggesting an effective replicative advantage. Interestingly, the NS4A site was limited to strains circulating between 2004 and 2014, reflecting a transient advantage (Fig. 3D, right panel). Only two branches, coinciding with a significant CHIKV outbreak in 2008-200930, were identified under episodic selection (denoted by orange dashed lines) (Fig. 3D, Supplementary. Data10). Similar to the other serotypes, MEME analysis confirmed several sites in nonstructural proteins were under episodic selection (Fig. 3D, right panel, and Supplementary. Data12 and Supplementary. Data13). Despite decades of circulation, only a few mutations were positively selected for each serotype. These resided primarily in non-structural proteins, suggesting that inter-serotype competition in Thailand is driven by replicative advantages rather than immune evasion.
Selection dynamics within the coding regions of four dengue virus (DENV) serotypes (A) DENV1, B DENV2, C DENV3, and D DENV4 as obtained from the time-stamped phylogenies. Each panel displays a Maximum Clade Credibility (MCC) tree at the center, trees were obtained from the coding regions of each DENV serotype (see Materials and Methods for details). The left panel highlights pervasive positively selected sites for each serotype, identified using the FUBAR method (see Supplementary Data 1–13). Each positively selected site is represented by a unique color, with varying shades indicating different amino acid replacements. These sites are integrated into the time-stamped phylogenies to trace the timing of each selection event accurately. The right panel shows episodically selected sites determined by the MEME method, with sites mapped onto specific branches that were identified using the aBSREL method for detecting branch-specific selection (see Supplementary Data 1–13). Sites that persisted throughout the evolutionary history are marked with arrows. Additionally, the locations of the viral proteins are also indicated on the trees (see Supplementary Data 14, mapping). Strains within branches identified under episodic positive selection are represented with the red dots across all the temporal trees (see Supplementary Data 1–13). (*Branches belonged to the same node therefore there is not a clear evolutionary direction).
Phylogeographic origins and dispersal patterns of dengue serotypes in Asia demonstrates that endemic circulation has driven diversification in Thailand
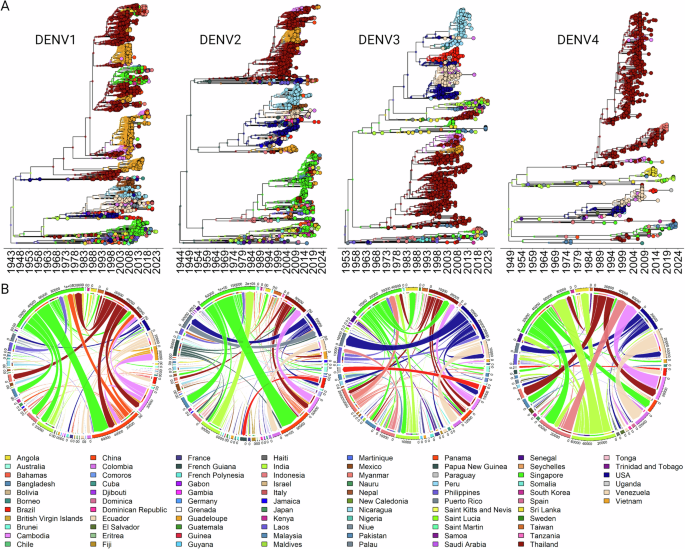
The monophyletic clustering of lineages and the presence of Thai strains at ancestral nodes suggest that these lineages may have originated in Thailand. (Fig. 2). To further substantiate the premise that competition between resident lineages has sustained DENV circulation in Thailand and that external introductions have played a minimal role, a discrete phylogeographic analysis was undertaken. Maximum Clade Credibility (MCC) trees (Fig. 4A) were constructed to visualize the transmission patterns and geographical dispersal of the four DENV serotypes beginning with their emergence to the present day. According to the phylogenetic rooting, the initial emergence of all four serotypes is traced back to Asia, with DENV1 and DENV3 originating from the Philippines, DENV2 from Papua New Guinea, and DENV4 from India, spanning the period from 1944 to 1955. The MCC trees demonstrate temporal clusters of geographically diverse sequences, indicating periods of increased transmission and spread. These clusters reflect not only persistent endemic circulation within certain regions but also cross-border dissemination of the virus. Distribution and phylogenetic branching patterns are known to align with human movement, vector ecology, and environmental conditions conducive to DENV propagation9,10,31. The analysis substantiates our initial hypothesis that the majority of the DENV diversification in Thailand can be attributed to sustained local transmission. This is determined by the identification of internal nodes which designate Thailand as the location of diversification for several main clades. Conversely, Thai sequences that cluster within genotypes less represented in the region, and which exhibit limited diversification, are indicative of external introductions into the country (Fig. 4A).
A Phylogeographic Relationships displayed using Maximum Clade Credibility (MCC) trees, this analysis elucidates the endemic diversification of DENV in Thailand and reveal the trajectories of all four DENV serotypes across the globe. The time-scaled phylogenies display ancestral nodes and current geographic locations (tips) as discrete states, illustrating the spatial and temporal spread of the virus. B Dynamic Pathways of Geographical Movement displayed as circular plots generated by the ‘circlize’ package in R to depict the Markov-jump trajectories of DENV movement. The visualization highlights the frequency and routes of viral importation, exportation, and intra-country dispersal, providing the role of Thailand the global dispersal of DENV. (Color legend for all the countries is provided at the bottom of the Figure).
The Markov jump count analysis, as shown in the circular plots of Fig. 4B, quantifies and visualizes the transmission events (not total number of sequences) of each dengue virus (DENV) serotype between countries. Here, Singapore emerges as a significant focal point for the global dissemination of DENV, implicated in numerous exportation events involving all four serotypes. The circular plots suggest that Thailand is a substantial contributor to the international spread of DENV1, DENV2, and DENV3. However, the transmission of DENV4 from Thailand appears to be predominantly regional, with most dissemination events involving only neighboring countries such as Myanmar, Cambodia, and Singapore. Corroborating the insights obtained from the MCC trees, the Markov jump analyses further support the observation that Thailand has a relatively low incidence of DENV importation from external sources. Not surprisingly, the primary contributors are its neighboring countries, Myanmar, Cambodia, and Singapore. This pattern underscores the regional dynamics of DENV transmission and highlights the interconnected nature of DENV spread within Southeast Asia.
Impact of CHIKV on genetic diversity and Force of infection on dengue Serotypes in Thailand
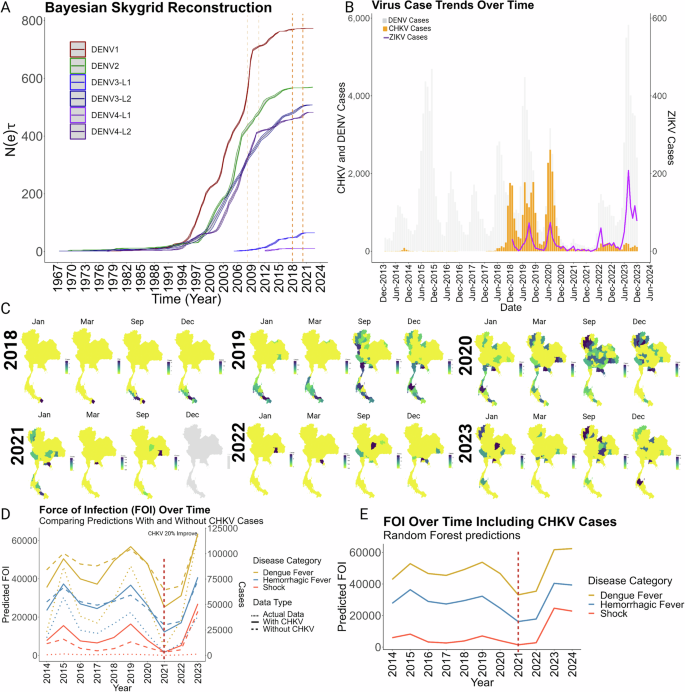
Finally, we asked what factors or events could account for the dearth of DENV infections in 2021 (Fig. 1A). Considering the potential impact of the CHIKV outbreak of 2009-2010 on the selection pressure of DENV4 (Fig. 3D) we first evaluated the trends in genetic diversity over time for all four serotypes in Thailand. Bayesian Skygrid reconstructions (Fig. 5A) represent the effective population sizes (Neτ) across DENV serotypes and lineages over an extended time frame (1967-2023) which provide insights into the genetic diversity and transmission dynamics of DENV within Thailand. DENV1 began its upward surge in 1994, with DENV2-4 serotypes following suit a few years later, each indicative of increased viral transmission and expansion in the country. What is notable in these Neτ plots are the ‘pauses’ or ‘plateaus’ that have interrupted the upward trajectories or sharp increases that have accelerated them, influenced by cyclical variations in transmission, environmental changes, intervention strategies, or inter-serotype competitions, which affect the equilibrium between rates of new infections and clearance. Upward jumps in diversity are required for a lineage to survive after events that quelled infections. DENV1 and DENV4 (lineage 1) exhibit inflection points around 2009 (dashed line; Fig. 5A), wherein these drastic changes in genetic diversity coincided with the major outbreaks of CHIKV reported in Thailand30,32. A similar flattening of the genetic diversity curve between 2018-2020 was observed for all four serotypes (Fig. 5A).
A Demographic reconstruction using skygrid analysis. The demographic history of the six identified DENV lineages in Thailand are shown. The major epidemic waves of Chikungunya virus (CHIKV) are marked with dashed orange lines, providing context for the temporal overlay of viral expansions. B Comparative analysis of chikungunya and zika virus transmission. Temporal distributions of Chikungunya (CHIKV) and Zika virus (ZIKV) cases are also shown illustrating concurrent outbreaks with DENV (in background gray). C Spatiotemporal distribution of CHIKV Cases. Similar framework as for DENV were represented the transmission dynamics of CHIKV, with seasonal patterns beginning in January, peaking in March and September, and concluding in December. The comprehensive spatial analysis for CHIKV is available in Supplementary Video 3) D Force of infection (FOI) of DENV obtained from the random forest model compared with and without the incorporation of the CHKV cases contrasted against all cases for the different clinical categories of DENV. The drop in the cases by 2021 is denoted with a red dashed line. E Application of the selected model (incorporating the CHKV cases to forecast the dynamics of dengue virus infection determined by the estimated FOI for DENV over time, for the year 2024. The year 2021 is marked with a red dashed line.
Recent studies by Brito et al.33 and Pinotti et al.34 have suggested that short-term immunity induced by Zika virus (ZIKV) infections may contribute to the temporary suppression of dengue virus (DENV) cases. Our analysis aligns with these findings, showing an increase in ZIKV cases in 2020 (Fig. 5B) preceding the significant decline in DENV cases in 2021 (Fig. 5B). However, given the relatively low overall incidence of ZIKV across Thailand, it seems unlikely that ZIKV alone could confer widespread partial protection against dengue. Interestingly, Brito et al.33 also reported a surge in chikungunya virus (CHIKV) cases prior to the decrease in DENV cases. Despite CHIKV (Togavirus) being taxonomically more distant from DENV than Zika virus (both flaviviruses), the temporal co-circulation is noteworthy. There were three consecutive outbreaks of CHIKV infections from 2018 to 2020, peaking in 2020, just before the significant reduction in DENV cases observed in 2021 (Fig. 5B). Other illnesses monitored at this time did not experience a similar drop in cases including scrub typhus35 and leptospirosis36, suggesting reporting of DENV would not have been deficient during the SARS-CoV-2 pandemic. Whether other factors related to mobility, such as travel restrictions, sheltering in place orders, and limited gatherings played a role cannot be ruled out37. Nevertheless, the sequential pattern of viral epidemics hints at complex interactions between circulating arboviruses that could drive down cases through cross-immunity or competition for vector resources.
To further explore the dynamic interactions between chikungunya virus (CHIKV) and dengue virus (DENV), including potential competition or partial protective effects, we conducted a detailed analysis of the geospatial distribution of CHIKV cases across Thailand. From 2018 to 2020, the geographic spread of CHIKV closely mirrored the distribution patterns of the IndexP, suggesting that regions with prior outbreaks often became the epicenters for subsequent year’s epidemics (Fig. 1B and Supplementary Video 1–3). These patterns were consistent with those observed for DENV, tracking with shifts in the IndexP. However, unlike the distribution trends observed with DENV, the dispersal pattern for CHIKV was notably disrupted and did not revert to its previous trajectory in the following years of 2022 and 2023 (Figs. 1C and 5B). This deviation from the expected geographic pattern could indicate factors beyond simple vector availability, such as viral interference or cross-immunity, might play critical roles in shaping the epidemiology of these viruses.
We developed a random forest model to estimate the force of infection (FOI) for dengue virus (DENV). Our variable impact analysis (Fig. 5C), supported that the infection category (dengue fever (DF), dengue hemorrhagic fever (DHF), and severe dengue fever (SDF)), yearly case data, and total case counts significantly increased node purity and therefore improved the statistical power of the model (Supplementary Fig. S3) Across all categories, the actual observed case numbers in Thailand from 2014 to 2021 matched trends estimated by FOI calculations, which use mathematical and statistical models based on epidemiologic factors and assumptions (Fig. 5D). Confident in the model’s ability to predict DENV dynamics, we next assessed the potential impact of including other variables, such as concurrent chikungunya virus (CHIKV) cases. Inclusion of CHIKV case data into the model yielded a 20% improvement in its predictive ability, suggesting this factor likely influenced the observed decline in cases, notably accentuating the sharp decline in cases in 2021.The application of the random forest model to forecast DENV infection trends for 2024 predicts a moderate increase in dengue fever (DF) cases, continuing the trend observed in previous years. This rise is not only limited to DF but also extends to more severe clinical manifestations such as shock and hemorrhagic fever (Fig. 5E). Significantly, the projected values for 2024 are anticipated to surpass the incidence rates recorded in 2019. This analysis, underscores the persistent upward trajectory in both the frequency and severity of DENV cases, highlighting a critical area of concern in public health monitoring and response strategies. These results also suggest that the concurrent circulation of multiple arboviruses may drive a selective dynamic that influences the genetic composition of dengue virus strains.