Ethics statement
This study was approved by the Institutional Review Board (Project title: A Randomized, Open-Label, Controlled Clinical Trial of Remdecivir Plus Ipratropium Bromide Versus Remdecivir Alone in Patients With Severe COVID-19, approval number: 2021-01-011-001). According to the principles of the Declaration of Helsinki, signed written consent was obtained from each participant or their legal guardian for the use of data and samples for scientific purposes.
All animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of Jeonbuk National University (approval title: “Evaluation of the Efficacy of Ipratropium Bromide Against SARS-CoV-2”; approval No. JBNU 2021 − 0184; Ethics Committee approval No. NON 2022 102) and were conducted in accordance with institutional regulations and the ARRIVE guidelines and checklist.
Preparation of total RNA of COVID-19 patients’ blood samples
Eight COVID-19 patients (≥ 19 years old) in South Korea were enrolled from February 2020 to January 2021. The diagnosis of COVID-19 in the enrolled patients was confirmed using molecular methods. Specifically, we utilized our in-house-designed reverse transcription polymerase chain reaction (RT-PCR) targeting the N gene, along with a commercial kit (Kogene Biotech, Seoul, South Korea) targeting the E and RdRp genes, following the manufacturer’s protocol. A positive COVID-19 diagnosis was determined if more than two genes were detected at a Ct value 19. Regarding lung imaging, all severe cases exhibited pneumonic changes in chest imaging, whereas mild cases did not show significant abnormalities. Among them, five had mild symptoms, and three had severe symptoms on admission. Disease severity was classified based on the World Health Organization (WHO) COVID-19 severity criteria (https://bestpractice.bmj.com/topics/en-gb/3000201/criteria):
-
Mild illness: Symptomatic patients meeting the case definition for COVID-19 without evidence of hypoxia or pneumonia.
-
Severe disease (adolescent or adult): Clinical signs of pneumonia (i.e., fever, cough, dyspnea, fast breathing) plus one of the following:
All severe cases in our study met at least one of these criteria, while mild cases did not exhibit hypoxia or pneumonia. We have clarified this classification in the revised manuscript.Blood samples were collected from each patient at diagnosis and convalescence. Human peripheral blood mononuclear cells (PBMCs) were isolated using Ficoll Paque Plus (Sigma-Aldrich, Burlington, MA, USA) following density gradient centrifugation20. Total RNAs were isolated from PBMCs using the TRI Reagent (Molecular Research Center, Inc., Cincinnati, OH, USA).
|
Age/sex |
Disease severity |
Comorbidities |
Pneumonia on imaging |
SpO2 in admission |
Oxygen therapy |
Steroid use |
|---|---|---|---|---|---|---|
|
35/F |
Mild |
None |
No |
98% |
No |
No |
|
22/M |
Mild |
None |
No |
98% |
No |
No |
|
46/M |
Mild |
HTN, Hyperlipidemia |
No |
97% |
No |
No |
|
30/M |
Mild |
HTN |
No |
97% |
No |
No |
|
30/M |
Mild |
None |
No |
98% |
No |
No |
|
79/F |
Severe |
HTN, DM, Guillain-Barre syndrome |
Yes |
85% |
Yes |
Yes |
|
61/F |
Severe |
None |
Yes |
89% |
Yes |
No |
|
75/F |
Severe |
HTN, old CVA, Lt.hemiparesis |
Yes |
88% |
Yes |
No |
Data sources for drug screening analysis
The mRNA levels of the collected sixteen blood samples were measured by QuantSeq 3′ mRNA sequencing conducted by e-Biogen (Seoul, Korea)21. The raw FASTQ RNA sequencing data were processed using conventional NGS pipeline tools, including Trimmomatic (version 0.39-1) and Salmon (version 0.7.2), for trimming and alignment processes, respectively, producing quantified transcriptome data in transcripts per million (TPA). Transcriptome datasets for 23,277 chemicals were collected from large-scale databases, including the Connectivity Map and Library of Integrated Network-based Cellular Signatures L1000.
RAPTOR AI
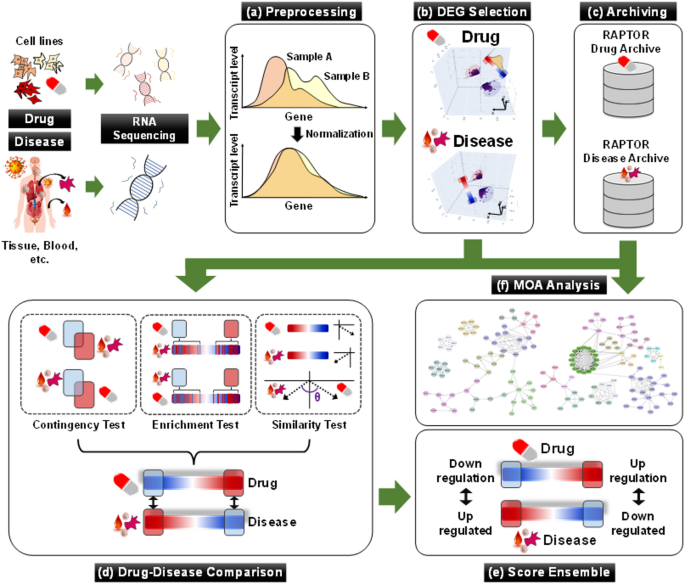
RAPTOR AI (ONCOCROSS Co., Ltd., Seoul, Korea) is a commercial AI drug-screening platform with four analytical processes: collection and preprocessing of transcriptome datasets, differently expressed gene (DEG) selection, drug-disease comparison, and score ensemble (Fig. 1)22. The analysis utilized both drug- and disease-derived RNA expression datasets to identify ideal drug candidates (see details in Supplementary Methods).
Overall analytic process of RAPTOR AI. (a) RNA datasets are standardized and normalized through preprocessing before use in the drug-screening process. (b) Statistically significant DEGs are identified between normal and disease samples, as well as between drug-treated samples and their negative controls. (c and f) All processed data is stored in proprietary databases for future studies, including MOA analysis. (d) The Drug-disease comparison analyzes the complementary correlation between drug-derived and disease-derived DEGs using three statistical tests: the contingency test (CT), the enrichment test (ET), and the similarity test (ST). (e) The final chemical score is determined in the score ensemble process.
Signaling pathways and mode of action analysis
A web-based pathway analysis tool, the Consensus Pathway Analysis (CPA), was utilized for the pathway analysis23. Detailed analysis of mode of actions (MOAs) among DEGs (Table S1) identified by RAPTOR AI within pathways was conducted using the Kyoto Encyclopedia of Genes and Genomes pathway database, following the standard analytical protocol24. P-values
Compounds
IB and remdesivir were purchased from Selleck Chemicals, LLC (Houston, TX, USA). IB was administered to the whole body using a nebulizer, and remdesivir was dissolved in dimethyl sulfoxide (DMSO; Sigma-Aldrich, St. Louis, MO, USA) and subsequently diluted with saline to the final concentration of the target dose.
Cell and virus
Vero cells were purchased from the Korean Cell Line BANK. The SARS-CoV-2 strain was obtained from the National Culture Collection for Pathogens of the Korea Disease Control and Prevention Agency. The virus was proliferated in Vero cells at 37 °C, 5% CO2, and subsequently harvested for our experiments. Genome details of the virus strain are available at GISAID (identifier: BetaCoV/Wuhan/IVDC-HB-01/2020| EPI_ISL_402119) and the China Microbiology Data Center (accession number NMDC10013001 and genome accession number MDC60013002-01).
Cell culture
Vero cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, Waltham, MA, USA), supplemented with 10% FBS, 1% penicillin-streptomycin, at 37 °C, 5% CO2. A549-ACE2-TMPRSS2 and THP-1 cells were co-cultured in Roswell Park Memorial Institute 1640 medium, supplemented with 10% FBS, 1% penicillin-streptomycin, at 37 °C, 5% CO2.
Cell viability assay
Compound cytotoxicity was assessed using WST-1 cell proliferation reagent (Roche, Basel, Switzerland). Vero cells (2 × 104 cells/well) were seeded in a 96-well clear flat-bottom TC-treated microplate with DMEM and incubated at 37 °C, 5% CO2. The next day, cells were washed once with phosphate-buffered saline (PBS) and serial two-fold dilutions of the cell suspension were inoculated into fresh medium. At 48 h post-infection, 10 µL of WST-1 was added to each well, followed by 2 h of incubation at 37 °C and 5% CO2. Cell viability was determined by measuring absorbance using a microplate reader.
Virus infection and cell harvest
Vero cells were seeded in 24-well plates until attachment. After a cell wash with fresh DMEM, the cells were inoculated with 0.01 multiplicity of infection (MOI) SARS-CoV-2, or with DMEM for negative control, then incubated at 33 °C for 1 h for virus adsorption. After drug treatment tests, the cell pellets were collected at specified time points and stored at -80 °C for RNA extraction.
Animal study
Inbred SH101 hamsters of the same genetic background were utilized in assessment of IB efficacy in treating COVID-19. Four-month-old hamsters were purchased from Alpha Biochemicals (Torrance, CA, USA) and were randomly divided into groups of no more than five hamsters per cage in the Animal Biosafety Level 3 Animal Breeding Room. All animal maintenance and experimental procedures were performed according to the regulations and procedures of the facility.
Evaluation of therapeutic efficacy
The COVID-19 animal model in SH101 hamsters was established by intranasal injection of 50 µL SARS-CoV-2 aqueous solution (105 TCID50). Treatment details for each group are in Table S2. The healthy control received 50 µL DMEM, while infection control group was given the viral solution. Remdesivir or IB administered in a weight-dependent manner. According to the scheduled experimental timeline, all hamsters euthanized under anesthesia with isoflurane followed by cervical dislocation.
A remdesivir solution was prepared by dissolving the drug in DMSO, filtering through a 0.45-µm syringe filter, and diluting with saline to a 15 mg/kg concentration. The prepared remdesivir solution was intravenously administered to appropriate groups from 1 to 8 day post-infection (dpi).
For IB treatment, a 500 µg/2 mL stock solution was diluted to 62.5 µg/kg with saline. Hamsters in the IB group received IB in nasal cavity via nebulization using a peristaltic pump (Fig. S1).
|
Group |
Sacrifice |
Total |
|||
|---|---|---|---|---|---|
|
2 dpi |
5 dpi |
8 dpi |
|||
|
Normal animal |
Control |
5 |
5 |
5 |
15 |
|
Disease animal |
Vehicle |
5 |
5 |
5 |
15 |
|
Remdesivir |
5 |
5 |
5 |
15 |
|
|
Ipratropium bromide |
5 |
5 |
5 |
15 |
|
Hamster clinical observation
A thermal imaging camera (FLIR, Wilsonville, OR, USA) was utilized to capture the thermal images of the whole bodies of hamsters from 0 to 8 dpi. The DirA (FLIR Tools) was then employed to identify the highest temperature point.
Body weight was measured at 0–8 dpi using a scale. The rate of weight change (%) was calculated as [current weight/0 dpi weight × 100%].
Student’s t-test was conducted using the mean and standard deviation values to identify significance on differences between groups.
Blood analysis
At 2, 5, and 8 dpi, whole blood was collected from each hamster group by cardiac puncture. Whole blood was centrifuged at 2,500 g for 10 min to separate the serum. The serum was immediately stored at -80 °C.
The D-dimer and fibrin degradation products (FDP) levels were quantified by enzyme-linked immunosorbent assay (ELISA) using ELISA kits (D-dimer: Cat. No. MBS012417, FDP: Cat. No. MBS005821, MyBioSource, San Diego, CA, USA). The samples were diluted 10-fold following the manufacturer’s instructions. Subsequently, a 50 µL of the diluted sample was dispensed into a well on a D-dimer or FDP antibody-coated micro-ELISA plate and incubated at 37 °C for 60 min. Then, the chromogen solution was added and the mixture reacted for 15 min. Immediately after terminating the reaction by adding the stop solution, the OD value was measured at 450 nm using an ELISA reader (iMark microplate reader, Cat No. #1681130; Biorad, Hercules, CA, USA). The OD450 values were normalized based on a blank OD450. The protein concentrations were calculated from the OD450 values using standard curves generated using six standard samples for D-dimer (31.2 ng/mL to 1,000 ng/mL) or FDP (0.625 pg/mL to 10 pg/mL) respectively.
The interleukin (IL)-6and tumor necrosis factor (TNF)-⍺ levels were quantified using ELISA kits (IL-6: Cat. No. MBS7606648, TNF-⍺: Cat. No. MBS7606475). The sample loading procedures were identical to those described above, except that a 100 µL sample was used instead of 50 µL, and the incubation time on the antibody-coated plate was extended to 90 min instead of 60 min. Then, a biotin-labeled antibody solution was added and allowed to react, followed by washing with a wash buffer. Horseradish peroxidase-streptavidin conjugate was added, reacted, and then washed. After adding the chromogenic substrate, the mixture reacted according to the manufacturer’s instructions, until the stop solution was added to stop the reaction. Calculation IL-6 and TNF-⍺ levels were calculated using the same procedure as described above, utilizing the OD450 values and standard curves of IL-6 (7.8 pg/mL to 500 pg/mL) and TNF-⍺ (3.125 pg/mL to 200 pg/mL).
Autopsy and histopathology
SH101 hamsters at 2, 5, and 8 dpi were euthanized and lungs were isolated by laparotomy. Whole-lung images were captured using a digital camera. Immediately, the right lung lobe was fixed using 10% neutral formalin. After washing with water followed by dehydration with ethanol, the tissue was made transparent using xylene and embedded in paraffin via a penetration process. Sections of 5 μm were cut, attached to glass slides. After deparaffinization and dehydration, lung tissue samples were subjected to H&E and IHC staining following general procedures (Supplementary Methods) and observed under optical microscopy.
Quantification of the SARS-CoV-2 virus in lung tissues of SH101 hamsters
At 2, 5, and 8 dpi, SH101 hamsters were euthanized and dissected to isolate the lungs. Left lung lobe was homogenized with a tissue homogenizer and centrifuged at 3,000 rpm for 10 min at 4 °C. The supernatant was carefully collected, and total RNA was extracted using the RNeasy Mini Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s manuals. The isolated total RNA was used as template in the subsequent RT-qPCR (Supplementary Methods).
Real-time PCR standard curves generated using serial dilution of recombinant plasmids. The log10 transformations of the equivalent nucleotide copy number per mL of the samples were used to express the viral titer. Statistical significance between groups was evaluated by a one-way ANOVA multiple comparison test at P 0.05 using the mean ± standard deviation.
Bronchoalveolar lavage fluid (BALF) test
At 2, 5, or 8 dpi, neck of pre-euthanized hamster was dissected and a catheter was inserted into the bronchus and secured with a suture. Subsequently, 1 mL PBS was circulated three times to obtain BALF. Cytospin centrifugation was used to separate the cells from the BALF solution before drying the slides. Giemsa staining solution (Cat No. GS-500, Sigma-Aldrich) was used to stain the dried slides. The stained slides were covered with mounting solution and cover glass, and infiltrated neutrophils were examined under an optical microscope.