Genomic diversity of heartworms from multiple continents, countries, and hosts
We analysed whole-genome sequencing data from 127 adult heartworms (D. immitis) sampled from nine countries in five regions: Australia, North America (USA), Central America (Costa Rica, Panama), Europe (Greece, Italy, Romania), and Asia (Malaysia, Thailand) (Fig. 1a). Samples were primarily collected from domestic dogs (n = 115), but we also sampled cats (n = 4), foxes (n = 4), a ferret (n = 1), a golden jackal (n = 1), a leopard (n = 1), and a wildcat (n = 1). Across the full cohort, inclusive of outgroup samples, we obtained an average coverage per sample of 72.72× for the nuclear genome, 12,788.18× for the mitochondrial genome, and 1014.25× for the Wolbachia endosymbiont genome (Supplementary Data 1, Supplementary Figs. 1–4). Variant calling, joint genotyping, and filtering of the full cohort revealed 301,310 single nucleotide polymorphisms (SNPs) and 80,514 indels in the nuclear genome, along with 498 SNPs and 24 indels in the mitochondrial genome, and 25,473 SNPs and 1381 indels in the Wolbachia genome; within D. immitis samples, 301,004 nuclear SNPs (3.4 SNPs per kb), 57 mitochondrial SNPs (4.1 SNPs per kb), and 360 Wolbachia SNPs (0.4 SNPs per kb) were found. The SNP frequencies in the nuclear and mitochondrial genomes are comparable, whereas the frequency in the Wolbachia genome is considerably lower. These frequency data are comparable to those of other Wolbachia-infected filarial worms, such as Onchocerca volvulus24 and Wuchereria bancrofti22.
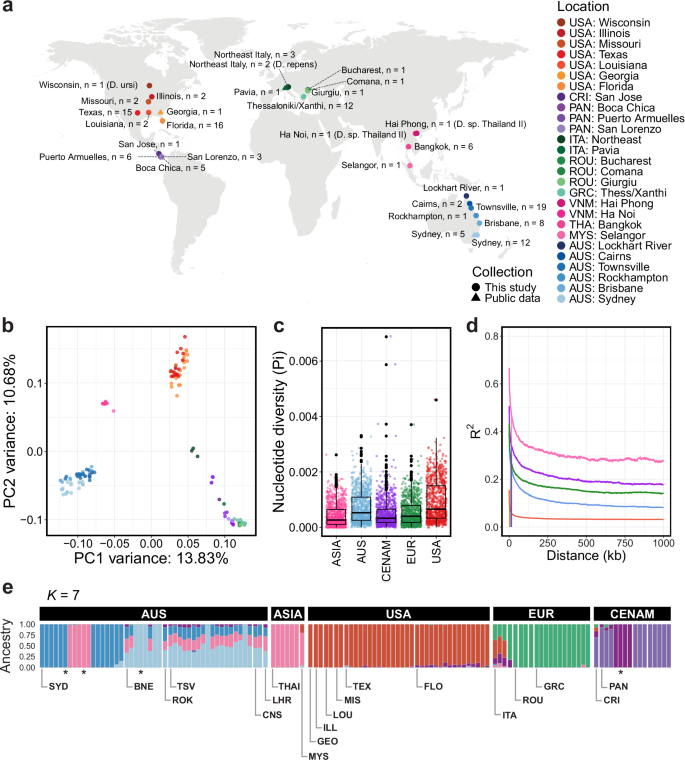
a World map showing the location of adult heartworm specimens used in the study. Adult Dirofilaria ursi, D. repens, and D. sp. ‘Thailand II’ were included as outgroup samples. b Principal component analysis (PCA) of 218,158 autosomal single nucleotide polymorphisms (SNPs) from 124 heartworms. c Box plots showing the nucleotide diversity (Pi) distribution within broad geographical regions, with each point representing a 100 kb sliding window. d Linkage disequilibrium (LD) decay plot for the heartworm cohort, grouped by broad geographic region. Vertical lines show the distance between SNPs where the LD is half the maximum value. e Admixture inferred using NGSadmix with seven clusters (K). Each vertical bar represents an individual sample, with colours representing the clusters. Labels at the top of the bar charts indicate the broad geographic origins of samples, while those at the bottom show their more specific locations. An asterisk (*) denotes a replicate of the preceding sample. USA United States of America (Georgia: GEO, Illinois: ILL, Louisiana: LOU, Missouri: MIS, Texas: TEX, Florida: FLO), CRI Costa Rica, PAN Panama, ITA Italy, ROU Romania, GRC Greece, VNM Vietnam, THAI Thailand, MYS Malaysia, AUS Australia (Sydney: SYD, Brisbane: BNE, Rockhampton: ROK, Townsville: TSV, Cairns: CNS, Lockhart River: LHR).
Nuclear variants reveal distinct continental partitioning of heartworms
Analyses of the heartworm cohort’s genetic diversity revealed distinct groups of samples based on broad geographical regions. Principal component analysis (PCA) using 218,158 high-quality autosomal SNPs revealed four clusters of samples corresponding to the continents from which the samples were collected (Fig. 1b and Supplementary Fig. 5; PC1 variance: 13.83%, PC2 variance: 10.68%). The third and fourth PCs showed a similar pattern of geographical clustering (Supplementary Fig. 6a; PC3 variance: 6.16%, PC4 variance: 4.42%); however, although the signal was comparatively weaker, the European and Central American samples formed separate clusters. Similar findings were observed in a previous study of 31 heartworm samples from Australia, Italy, and the USA, where samples from each country formed distinct clusters25. Independent PCAs of SNPs from each chromosome found the same continental clustering pattern across all chromosomes, including the sex-linked X chromosome (Supplementary Fig. 6b–f), suggesting that this is a genome-wide rather than region-specific observation. The distinct geographical partitioning in our nuclear data was not supported by the mitochondrial (Supplementary Fig. 7a) or Wolbachia data (Supplementary Fig. 7b), likely due to the limited number of variant sites within each dataset. This contrast between the strong nuclear and weak mitochondrial population structure was particularly surprising, however, the underlying biological cause remains unknown.
Analyses of genome-wide nucleotide diversity (Pi) further demonstrated differences between heartworm populations across continents. Pi differed significantly between all populations (Wilcoxon Rank Sum test, P < 0.05) (Fig. 1c). The highest Pi was observed in the USA (median window-averaged Pi = 7.77 × 10−4), while the lowest was in Asia (3.12 × 10−4). The genetic diversity of our USA samples aligned closely with that of a previous D. immitis study (median window-averaged Pi = 7.36 × 10−4)25. However, our Australian samples (median window-averaged Pi = 6.29 × 10−4) exhibited higher genetic diversity than those reported previously (3.9 × 10−5)25, likely due to the larger and geographically broader sampling here. Overall, the nucleotide diversity of our D. immitis samples is comparable to that of the filarial worm W. bancrofti (Pi[mean] = 2.4 × 10−4)22 but lower than that of O. volvulus (Pi[mean] = 4.0 × 10−3)24. Each D. immitis population exhibited different rates of linkage disequilibrium (LD) decay and baseline LD (Fig. 1d), with Asia and the USA showing the highest and lowest LD, respectively. Estimates of admixture provided further support for continental partitioning in heartworms. To infer the ancestry proportions of samples, we used NGSadmix with a range of K clusters (K = 2–10; Supplementary Fig. 8). At K = 7, Australian samples showed the most diverse admixture patterns with strong evidence of shared ancestry with Asia, whereas USA, Europe, and Central America were largely distinct with some evidence of shared ancestry in a minority of samples (Fig. 1e). The strong population structure in the nuclear genome of heartworms suggests that spatial spread has been restricted during the species’ history, resulting in distinct genetic profiles on each continent.
Inference of transmission between diverse heartworm hosts
Heartworms are obligate, vector-borne parasites whose successful establishment in a new environment depends on three key requirements. The first requirement is the presence of a suitable definitive host in which heartworms can develop, establish in the pulmonary arteries and the heart, and sexually reproduce to generate microfilariae, which circulate in the host’s bloodstream. The second requirement is a suitable mosquito vector that can ingest the microfilariae and support their development until they reach their infective larval stages; at least 17 mosquito species, including members of Aedes, Anopheles and Culex spp., are confirmed vectors in Europe26, and perhaps as many as 60 different mosquito species in total27, have been identified as competent vectors of D. immitis. The final requirement relates to the conditions needed for larval development within the mosquito, which is both temperature- and time-dependent. The rate of parasite development within the mosquito, from ingested microfilaria to infective stage larvae, is positively associated with outside ambient temperature, with a minimum temperature of 14 °C28,29. As such, warmer climates, including those induced by climate change, would accelerate this developmental stage inside the mosquito vector. This complex interplay between host, vector, and climate has enabled heartworms to infect both domestic and wild animals on almost every continent of the world.
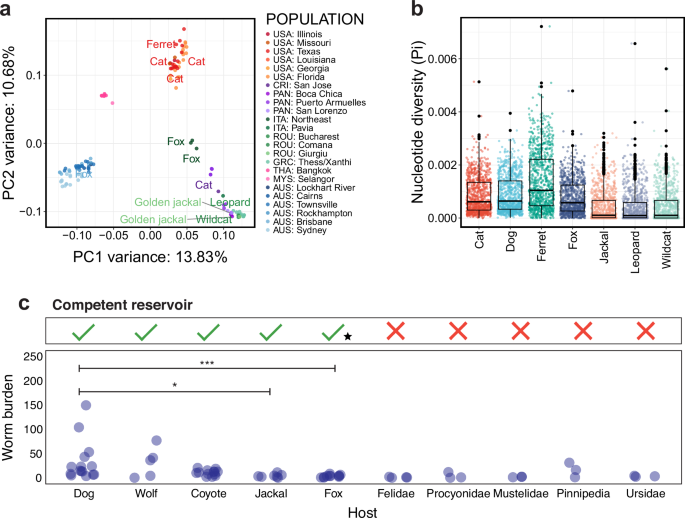
Although most heartworm samples in this study were obtained from dogs, the analysis of heartworm genomes from foxes, cats, a golden jackal, a ferret, and a leopard allowed us to evaluate genetic variation among different host species. We found that geography rather than the host species explained variation in the global heartworm population, evidenced by samples from cat and ferret nested within samples from dogs in the USA, fox and dogs in both Australian and Italian groups, and dogs and jackal, wildcat and leopard from Europe (Fig. 2a). Analyses of nucleotide diversity (Pi), absolute nucleotide divergence (DXY), and genetic differentiation (FST) provided additional insights on the variation within and between different host species. The heartworm sampled from the ferret had the highest diversity (median window-averaged Pi = 1.27 × 10−3) (Fig. 2b) and the greatest divergence from other hosts (Supplementary Fig. 9a). In contrast, the samples from the golden jackal, leopard, and wildcat had the lowest diversity (median window-averaged Pi = 1.46–2.09 × 10−4) and were the least divergent from each other compared with all other host pairs. However, only a single heartworm was sampled from each of these hosts, and the variation observed in the jackal, leopard, and wildcat samples may be confounded as they all originated from Romania. FST values between host pairs were generally low, with foxes and cats showing the highest genetic differentiation from the jackal, leopard, and wildcat (Supplementary Fig. 9b). Collectively, these data suggest an absence of obvious host-specificity and that transmission is likely between domestic animals and wildlife. However, further geographic sampling from non-dog hosts is needed to exclude the possibility of host-specific adaptation.
a Principal component analysis (PCA) of heartworm nuclear diversity from Fig. 1b, with non-dog hosts labelled. The geographic origin of samples is indicated on the right (i.e. COUNTRY: city or region). Country abbreviations: USA = United States of America; CRI = Costa Rica; PAN = Panama; ITA = Italy; ROU = Romania; GRC = Greece; THA = Thailand; MYS = Malaysia; AUS = Australia. b Box plots show the distribution of nucleotide diversity (Pi) within host populations. Each data point represents a sliding window size of 100 kb. c Support for the heartworm life cycle and adult worm burdens in carnivoran hosts. The top panel highlights hosts that are frequently microfilaremic and are hence considered competent reservoirs for heartworms. A black star indicates cases where microfilaremia has been reported in the host, but they are generally considered to pose a low risk of being competent reservoirs. The bottom panel is a plot showing adult heartworm burdens across various hosts. Each point represents data from a study, sourced from the literature (Supplementary Data 2). Significant differences in worm burdens among competent reservoirs were assessed using a Kruskal–Wallis test with Dunn’s multiple comparisons. P-values ≤ 0.05 were considered statistically significant (*≤0.05, **≤0.01, ***≤0.001).
We next questioned which host may have harboured and dispersed heartworms before the emergence of modern dogs. We hypothesised that longer co-evolution between parasites and hosts would result in the latter being able to carry higher parasite loads. Felids (e.g. domestic cats, wild cats, snow leopards), procyonids (e.g. raccoon dogs), mustelids (e.g. European badger, ferrets), pinnipeds (e.g. harbour seal, brown fur seal), and ursids (e.g. brown bear, black bear) have been described as being infected with heartworms (Supplementary Data 2); however, they typically carry fewer adult worms and fail to support the parasite’s entire life cycle, and so they are not considered primary reservoirs30,31,32,33 (Fig. 2c). In contrast, wild canids are commonly microfilaremic and have high worm burdens. Wolves and coyotes have worm burdens similar to dogs, while jackals and foxes have significantly lower burdens than dogs (Kruskal–Wallis test with Dunn’s multiple comparisons; Dog vs Jackal: P ≤ 0.05, Dog vs Fox: P ≤ 0.001) (Fig. 2c). These findings imply that canids, particularly ancestral wolves or coyotes, were most likely the primary hosts for heartworm during ancient times34.
Coevolution of canids and heartworms: distinguishing modern from ancient origins and dispersal
Previous work had proposed that canids originated in North America ~40 Mya when the continent was completely isolated14, whereas mosquitoes originated earlier in the mid-Cretaceous (~106 Mya)19. Phylogenetic analyses of mitochondrial DNA suggest that heartworms diverged from their closest relatives in the genus Onchocerca ~25 Mya during the Oligocene35, i.e. when canids and mosquito vectors were already well-established. Together, this timeline supports a long co-evolutionary history between heartworms and ancient canids.
The formation of the Bering Land Bridge and the Isthmus of Panama during the Miocene and Pliocene facilitated the first migrations of canids into Eurasia (~7 Mya)14 and South America (3.5–3.9 Mya)36, respectively. Throughout history, fluctuations in sea level have repeatedly exposed and submerged the Bering Land Bridge. Fossil records and genomic data provide robust evidence that these periods of exposure enabled ancient canids to migrate between Asia and North America, facilitating population connectivity and supporting their widespread presence across the Northern Hemisphere. Examples include Pleistocene wolves, which were highly connected across North America and Eurasia16,37,38, Xenocyon lycaonoides, a large canid distributed across North America and Eurasia in the mid-Pleistocene39 and Eucyon, a jackal-sized canid distributed across North America, Eurasia, and Africa in the late Miocene40. The widespread presence of ancient canids across continents, together with evidence of their population connectivity, suggests that heartworms may have similarly formed highly connected populations alongside their hosts at that time.
The current hypothesis for the distribution of heartworms around the world is that it has been largely influenced by human movement in contemporary times. Alternative hypotheses suggest that heartworms spread with humans during or shortly after dog domestication (~14–40 kya), or that they spread with wild canids before domestication. To further explore the demographic history of heartworms, we inferred their effective population (Ne) size histories using SMC++ over the last 1 million years. Given the domestication hypothesis, we broadly characterise the population demographics into three periods: post-domestication (<14 kya), domestication (14–40 kya), and pre-domestication (>40 kya). Contemporary estimates of Ne after domestication (Fig. 3a) were consistent with current levels of genetic diversity (Fig. 1c) in each population, with USA and Australian populations showing the highest Ne and Pi, and Asia the lowest. Differences between populations in demographic analyses broadly support the DXY and FST results (Supplementary Fig. 10), which indicate the greatest differentiation and, therefore, infer population splits from Asia, which diverged from the populations of Europe, the USA, and Central America. Considering the variation in population demographic curves >40 kya (Fig. 3a), we propose that there were substantial differences in some populations that were evolving independently before dog domestication16,17,18. Given that modern global human movement would drive populations to be more similar, the exacerbation of differences in Ne post-domestication would further suggest that these differences in populations existed before domestication.
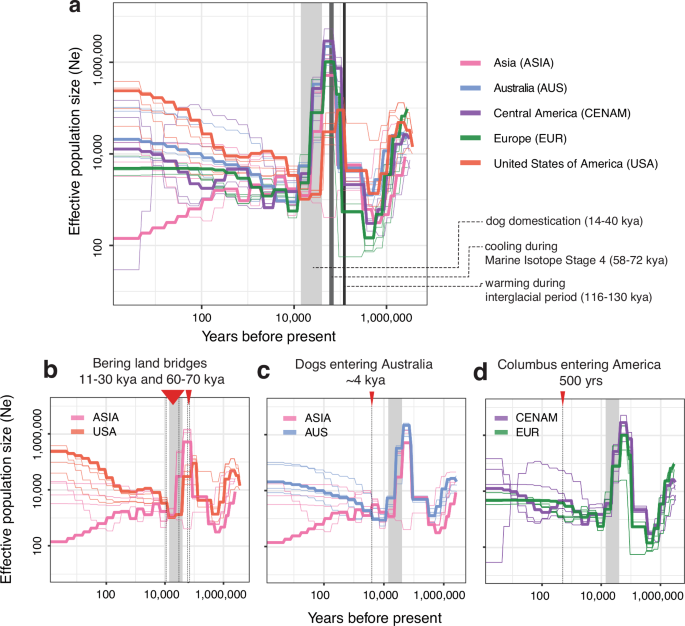
a Effective population size history of heartworm populations inferred using SMC++ based on a 2.5-year generation time. Each thick line represents the effective population size of heartworms on each continent over time. Thin lines represent jackknife sampling of chromosomes, iteratively removing one of four autosomal chromosomes, to visualise variance in Ne. The light grey box spans ~14–40 kya, highlighting the previously suggested period of dog domestication, as well as dark grey boxes highlighting global cooling (58–72 kya) and warming (116–130 kya) periods. b–d Reconfigured datasets from a, highlighting key pairwise comparisons to explore hypotheses presented in the text. b Comparison of the effective population size histories of heartworm populations from Asia and the USA. The red arrows and dotted lines represent periods of the Bering Land Bridge exposure. c Comparison of effective population size histories of Asian and Australian heartworm populations. The red arrow and dotted lines indicate the suggested period of dingo introduction in Australia. d Comparison of effective population size histories of Central American and European heartworm populations. The red arrow with a dotted line indicates the period of European colonisation of the Americas.
If heartworms were once one population, when did they diverge, and why? Our estimates of Ne do not extend earlier than 1 million years ago. However, between 1 million and ~50,000 years ago, significant climatic events correlate with changes in the heartworm population demographics. For example, we observe a substantial increase in Ne of all populations roughly aligning with the last interglacial period (Fig. 3a), when Earth experienced a climate as warm or even warmer than today (116–130 kya)41. Such an environment could have been suitable for the development and transmission of heartworms via mosquito vectors. Following the warm climates of the last interglacial period, the Earth began to cool. By Marine Isotope Stage 4 (58–72 kya), large continental ice sheets had formed across North America, Beringia, and northern Eurasia42. These ice sheets separated ancient canid populations37,43, potentially fragmenting the associated ancient heartworm population. This time period was associated with a fall in Ne, particularly in the USA samples, but for all populations, a considerable decrease was observed immediately before or during dog domestication, which occurred shortly afterwards (Fig. 3a–d). This fragmentation would have reduced gene flow for extended periods, explaining the continental grouping of samples in our study. Further insights might be obtained by exploring the presence of heartworm DNA in the remains of ancient canids; if successfully extracted, these ancient worm DNA sequences may fill in the temporal and geographic gaps needed to understand heartworm divergence in more detail. Similarly, further sampling in unstudied regions, including South America and Africa, and finer sampling within Asia, would enhance our understanding of heartworms’ evolutionary history.
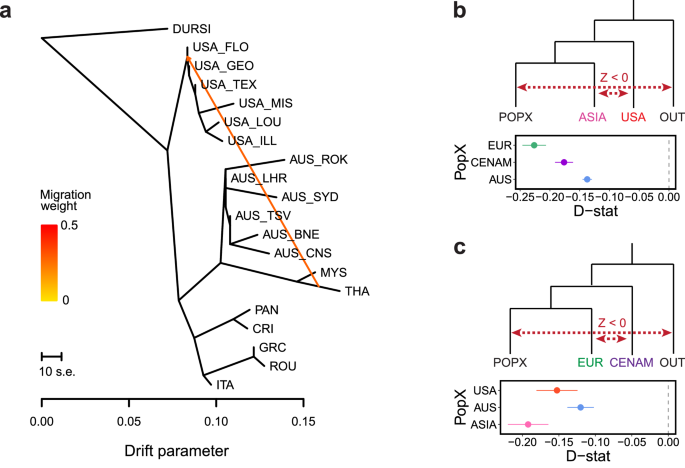
Considering global warming and cooling cycles, the fragmentation of canid and heartworm populations would have occurred multiple times. We found evidence of secondary admixture between heartworms from Asia and the USA after the initial population divergence, evident from both the admixture analyses and gene flow between populations (Fig. 4a; delta m = 4.1; all tree migration edges are presented in Supplementary Fig. 11). D-statistics (Fig. 4b, c) provided further support for excessive allele sharing between the USA and Asia relative to that between the USA and other regions (Fig. 4b). This admixture may reflect multiple waves of dispersal with canids across the Bering Land Bridge in the Late Pleistocene during periods of ice sheet contraction16 between 11–30 kya and 60–70 kya. Our population demographic data provide some support for this hypothesis, at least during 11–30 kya, when USA and Asian populations experienced a close overlap in Ne before undergoing rapid and distinct divergence thereafter (Fig. 3b).
a Maximum-likelihood tree of heartworm generated using TreeMix, with Dirofilaria ursi (DURSI) as an outgroup and one migration edge. b, c D-statistics for hypothetical allele-sharing scenarios between heartworm populations using Admixtools qpDstat with 20,392 single nucleotide polymorphisms (SNPs) and D. ursi as the outgroup (OUT). D-statistics test for admixture between four populations (W, X, Y, Z). A Z-score (D-statistic / standard error) is calculated, which provides information about the direction of gene flow (red dotted lines) (Z-score > 0 = gene flow between either [W and Y], or [X and Z]; Z-score < 0 = gene flow between either [W and Z] or [X and Y]). For all data presented, the Z score was significant (|Z| > 3). Error bars show the standard error.
The origin and spread of heartworms in Australia
Heartworms are endemic in Australia, but the mechanisms and timing of their arrival in the country have remained an enigma. Consistent with the modern human dispersal hypothesis, one means of introduction could have been after European colonisation, which began ~235 years ago. However, considering our genetic evidence of older, canid-mediated dispersal, another likely scenario is that heartworms were brought to Australia via dingos. The dingo, an ancient lineage from Asia, was the only canid present on the Australian continent until European settlement. Recent morphometric analysis indicates an East Asian origin of dingoes via Melanesia44, whereas genetic studies suggest that dingoes likely came from Island Southeast Asia and were brought to Australia by seafarers ~4 kya45.
Multiple lines of evidence support the introduction of heartworms into Australia from Asia. Both admixture and TreeMix analyses highlighted the close relationship, evidenced by partial or complete admixture profiles (Fig. 1e) and mutual monophyly (Fig. 4a) between Asian samples and samples collected along the East Coast of Australia (Fig. 5a). Calculations of DXY showed that AUS-ASIA (median window-averaged DXY = 5.81 × 10−4) was the least divergent pair, suggesting that Australian worms are more closely related to those in Asia than elsewhere (Supplementary Fig. 10a). D-statistics solidified this relationship by showing excessive allele sharing between the two continents relative to that between Australia and other regions (Fig. 5b). Finally, samples from Australia and Asia exhibit very similar population demographic trajectories of Ne until ~10 kya, which begin to diverge in the post-domestication period, consistent with the timing of the arrival of dingoes. Together, our findings align with an Asian origin of Australian heartworms, possibly transported with dingo hosts (Fig. 3c). We note that our sampling of Asian heartworms was from Thailand and Peninsular Malaysia, which are geographically distinct from Indonesia, from which dingoes would have entered Australia from ~4 kya, potentially explaining the small discrepancy in the estimated time of divergence. Sampling across a broader geographic region, particularly throughout the Indonesian archipelago and Melanesia, could bridge the gap between our Australian and Asian samples and provide further granularity of the Asia–Australia divergence, potentially coinciding more closely with the timeline of dingo arrival. An alternative, plausible scenario is the post-colonial introduction of heartworms into Australia through the importation of modern dog breeds via Asia. Importation of infected dogs from regions such as China or Japan could also explain the genetic divergence we currently observe between our Australian and Asian (represented only by Thailand and Malaysia) samples. If the genetic link between Asia and Australia is due to modern translocation, we might expect genetic links to other regions, not just Asia, which we do not find. Nonetheless, broader sampling across the Asian continent is needed to determine the relative genetic contribution of heartworms from ancient dingoes to more recent importation of modern dog breeds from Asia.
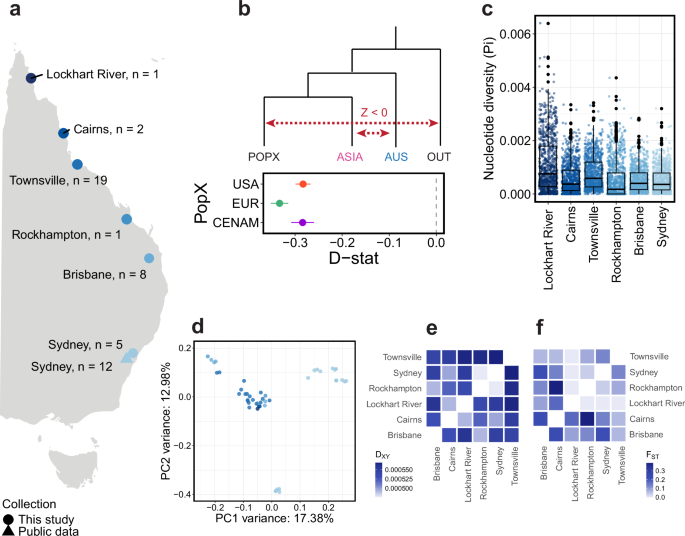
a Distribution of adult heartworm samples (n = 48) collected along the east coast of Australia. b D-statistics showing excessive allele sharing between Australia (AUS) and ASIA relative to that between AUS and other regions (POPX). Admixtools qpDstat was used to analyse 20,392 single nucleotide polymorphisms (SNPs), with Dirofilaria ursi as the outgroup (OUT). D-statistics test for admixture between four populations (W, X, Y, Z). A Z-score (D-statistic / standard error) is calculated, which provides information about the direction of gene flow (red dotted lines) (Z-score > 0 = gene flow between [W and Y], or [X and Z]; Z-score < 0 = gene flow between [W and Z] or [X and Y]). For all data presented, the Z score was significant (|Z| > 3). Error bars show the standard error. c Box plots showing the nucleotide diversity (Pi) distribution from dog hosts per city. Each data point represents a 100 kb sliding window. d Principal component analysis (PCA) of 131,508 nuclear SNPs in the Australian cohort. e Absolute nucleotide divergence (DXY) and f genetic differentiation (FST) between Australian cities using a median of 100 kb sliding windows. Only heartworms from dog hosts were included in the analyses for c, e, f.
Considering the extensive sampling along the north–south transect of eastern Australia, we sought to identify factors that might explain the current genetic diversity. Australian heartworms were the second most genetically diverse population globally (Fig. 1c). Within Australia, diversity was highest in Lockhart River (median window-averaged Pi = 8.87 × 10−4) and lowest in Rockhampton (2.60 × 10−4) (Fig. 5c). Heartworms from the northern state of Queensland were highly admixed, with two unique sources of ancestry that were largely absent from other continents (Fig. 1e). Interestingly, ten of 16 heartworms from Sydney, the southernmost site, had a single fixed ancestry that was detected in Queensland samples. Another four Sydney heartworms had a different fixed ancestry identical to that found in Asia, potentially reflecting modern movement, though the exact cause remains unclear. Previously, it was believed that heartworm cases in temperate Sydney and broader New South Wales were primarily caused by infected pets or mosquitoes travelling down from tropical and subtropical Queensland46, where climates are more favourable to heartworm transmission. However, our nuclear PCA showed a large cluster of samples primarily from Queensland, alongside two distinct clusters from Sydney (Fig. 5d). This suggests that Sydney heartworms may not necessarily originate from Queensland, as previously believed. Further investigations are required to determine whether there are local endemic populations of heartworms in Sydney. Differences in DXY and FST between Australian heartworm subpopulations were subtle, showing no clear relationship between genetic and geographical distance as might be expected for a vector-borne parasite (Fig. 5e, f).
Modern heartworm dispersal between Europe and Central America
Our genomic analysis revealed a close genetic relationship between worms in Europe and Central America. This finding was most evident in our PCA of nuclear SNPs, where samples from Greece, Italy, and Romania clustered with those from Costa Rica and Panama (Fig. 1b). In addition, estimates of DXY revealed that EUR-CENAM (median window-averaged DXY = 5.93 × 10−4) was the second least divergent compared with other populations. Admixture analysis using NGSadmix further supported this trend as the European and Central American samples shared high levels of ancestry at lower K values (K = 2–4; Supplementary Fig. 8). Although increasing K helped to differentiate worms from these two continents, low levels of admixture were still present in some samples (Fig. 1e). These findings were corroborated using D-statistics which provided strong evidence of excessive allele sharing between heartworms from Central America and Europe relative to that between Central America and other regions (Fig. 4c). Finally, Europe and Central America shared considerable overlap in Ne over deep evolutionary timescales (Fig. 3d). These findings support a modern migration event between European and Central American heartworms which took place after dog domestication.
One plausible modern migration event between Europe and Central America was the transatlantic transportation of domestic animals, including dogs, during the European colonisation of the Americas, which began in the late 15th century. Animals introduced to the Americas during the Age of Discovery were primarily livestock species, including cattle, sheep, pigs, goats, horses, and mules. However, dogs such as greyhounds and mastiffs were also introduced for hunting, guarding, protection, shepherding, and battle purposes47,48,49. The introduction of European dogs into the Americas is perhaps best documented from Columbus’ voyages, with historical records indicating the presence of at least 20 dogs on board during the second voyage to Hispaniola47. Therefore, infected dogs from Europe may have transported local strains of this parasite to the Americas. Alternatively, the shared ancestry between European and Central American heartworms could result from the more recent migration of humans and their pets.