Ethical consideration
The study was approved by the Institutional Review Board (IRB) of Shahid Beheshti University of Medical Sciences (IR.SBMU.RIGLD.REC.004 and IR.SBMU.RIGLD.REC.1399.058). The IRB exempted this study from informed consent. Data were pseudonymized before analysis; patients’ confidentiality and data security were prioritized at all levels. The study was completed under the Helsinki Declaration (2013) guidelines and all experiments were performed in accordance with Iran Ministry of Health regulations. Informed consents were collected from all individuals or their legal guardians. During the generation of LLM predictions, using the OpenAI API and Poe Web interface, we opted out of training on OpenAI and used no training-use models in Poe to maintain the data safety of patient information.
Study aim and experimental summary
The objective of this research is to evaluate the efficacy of CMLs in comparison to LLMs, utilizing a dataset characterized by high-dimensional tabular data. We employed a previously compiled dataset and focused our experimental efforts on the task of classifying COVID-19 mortality. As illustrated in Fig. 1, the primary experiment encompasses the following:
Study Design and Experimental Summary. Image caption: This figure illustrates the design and workflow of the study, which compared large language models (LLMs) and conventional machine learning (CML) approaches for the prediction of COVID-19 patient mortality. The patient data included demographics, symptoms, past medical history, and laboratory results. The data undergo preprocessing before being structured into input‒output instances. The CML pipeline involves training and validation via various algorithms, such as logistic regression, support vector machine, and random forest. Moreover, the LLM pipeline involves a prompt engineering loop. We also fine-tuned one LLM, Mistral-7b, by giving the input and ground truth. The study aims to predict patient outcomes (survival or death) on the basis of the provided information.
-
Assessment of the performance of seven CML models on both internal and external test sets.
-
The assessment of eight LLMs and two pretrained language models on the test set.
-
Assessment of a trained LLM’s performance on both internal and external tests.
Additionally, we investigate the performance of models necessitating training (CML and trained LLM) across varying sample sizes, coupled with an elucidation of model prediction mechanisms through SHAP analysis.
Study context, data collection, and dataset
This study was conducted as part of the Tehran COVID-19 cohort, which included four tertiary centers with dedicated COVID-19 wards and ICUs in Tehran, Iran. The study period was from March 2020 to May 2023 and included two phases of data collection. The protocol and results of the first phase have been published previously. The four COVID-19 peaks during this period covered the alpha, beta, delta, and Omicron variants.
All admitted patients with a positive swab test during the first two days of admission or those with CT scans and clinical symptoms were included in the study. A medical team collected the patients’ symptoms, comorbidities, habitual history, vital signs at admission, and treatment protocol through the hospital information system and reviewed the medical records. Laboratory values during the first and second days of admission were collected and organized from the hospitals’ electronic laboratory records using pandas (v1.5.3), and NumPy (v1.24.1). Patients with a negative PCR result in the first two days of admission or with one missing clinical record in the HIS were excluded.
The dataset included the records of 9,134 patients with COVID-19. The data were filtered to include demographic information, comorbidities, vital signs, and laboratory results collected at the time of admission (first two days).
Computational environment
All classical machine learning (CML) experiments were performed on a workstation equipped with an Intel Core i9-12900 K CPU, 64 GB of RAM, and an NVIDIA RTX 3090 GPU (24 GB VRAM), running Ubuntu 22.04 and Python 3.10. The primary packages utilized include scikit-learn (version 1.2.2), XGBoost (version 1.7.5), pandas (version 1.5.3), and NumPy (version 1.24.1).
The fine-tuning of the Mistral-7b-Instruct model was conducted using an NVIDIA A100 80GB GPU via a cloud-based environment (Google Cloud Platform), utilizing the transformers (v4.37.2), peft (v0.9.0), and bitsandbytes (v0.41.1) libraries. The QLoRA fine-tuning procedure was implemented using 4-bit quantization, gradient accumulation steps, and mixed-precision training to optimize memory usage and reduce computational cost. All LLM zero-shot experiments were conducted via the OpenAI API and Poe interface under controlled sessions to ensure reproducibility.
Data preprocessing
Supplementary Figure S1 illustrates summary of the pipeline from raw data preprocessing through feature engineering and cleaning, to the final training–test data split used for model development and evaluation.
Imputing and normalization
The features in the dataset were divided into categorical and numerical categories. To address the missing values in the numerical features, we used an iterative imputer from the scikit-learn library. This method employs iterative prediction for each feature, considering the multiple imputation by chained equations (MICE) method (16,17). Missing values in the categorical features were imputed via KNN from the scikit-learn library. For optimal model performance, the dataset was normalized via a standard scaler (18). These preprocessing steps were executed independently for the input features of the training, test, and external validation sets, ensuring a consistent approach for handling missing values across the experimental sets without information leakage.
Feature selection
The dataset comprised 81 on-admission features. The dataset was separated into external and internal validations using patient hospitals. Patients from Hospital-4 were used for external validation, whereas patients from the remaining hospitals were used for internal validation. For internal validation, we split the data with a test size of 20% and allocated 80% for training.
The output features in this study include “in-hospital mortality,” “ICU admission,” and “intubation,” with a focus solely on “hospital mortality” as the targeted feature, excluding other output features. Of the 81 features initially available, 76 were employed for training, comprising 53 categorical features and the remaining numerical values. During data wrangement, two duplicate features were dropped.
We strategically employed the lasso method for feature selection because of its effectiveness in handling high-dimensional data. The Lasso method introduces regularization by adding a penalty term to the linear regression objective function, which encourages sparsity in the feature coefficients15,16. This approach proved to be superior to alternative methods, facilitating notable enhancements in our results. Through the application of Lasso, we derived a refined dataset that highlighted the most impactful features on the basis of their importance, aiding dimensionality reduction. We subsequently ranked and selected the top 40 features for further analyses.
Oversampling
To address the issue of class imbalance in our dataset, we employed the synthetic minority oversampling technique (SMOTE), a widely used method in machine learning, particularly for medical diagnosis and prediction tasks17. By applying SMOTE, we mitigated dataset imbalances, resulting in a more robust and reliable analysis for predicting mortality. SMOTE works by creating synthetic samples for the minority class instead of simply duplicating existing samples. It selects samples from the minority class and their nearest neighbors and then generates new synthetic samples by interpolating between these samples and their neighbors. This approach not only increases the number of samples in the minority class but also introduces new data points, improving dataset diversity. In our experiments, the SMOTE technique was applied to the training set (X_train), increasing the number of samples from 6118 to 9760.
Preparing data for the LLM
To prepare the data for input into the LLM, we completed all the previous steps for feature selection and sampling, but normalization was not performed. As shown in Fig. 1, we converted the dataset into text. We categorized the dataset features into symptoms, past medical history, age, sex, and laboratory data. For symptoms and medical history, we considered only positive data. For age, we added ‘the patient’s age is’ before the age number. For sex, we used ‘male’ and ‘female.’ We used the normal range of laboratory data to classify the data into the normal range, higher than the normal range, and lower than the normal range. For example, if blood pressure and oxygen saturation were higher than the normal range, we used the sentence ‘blood pressure and oxygen saturation are higher than the normal range.’ We considered only laboratory data that were higher or lower than the normal range. The exclusion of negative features in symptoms and past medical history, or the normal range in laboratory data, is due to limitations in LLM context windows. We then concatenated the dataset into a single paragraph for each patient, indicating their medical history.
CML predictive performance
We employed five CML algorithms: logistic regression (LR), support vector machine (SVM), decision tree (DT), k-nearest neighbor (KNN), random forest (RF), multilayer perceptron neural network (MLP), and XGBoost. The hyperparameters were optimized via a grid search and cross-validation. The full details of training and hyperparameters are provided in Supplementary Sect. 1.
LLM predictive performance
We utilized open-source and proprietary LLMs to test their predictive power on clinical texts transformed from tabular data. First, we tested different prompts to determine the most efficient prompt to use, as well as the temperature (between 0.1 and 1). Full prompts are listed in Supplementary Table S1. We then sent clinical text and commands, received the unstructured output, and extracted the selected outcome, which could be either “survive” or “die.” We used different sessions for each prediction, limiting the memory of the LLM to remembering previous generations.
We tested open-source, open-weight models of Mistral-7b, Mixtral 8 × 7 B, Llama3-8b, and Llama3-70b via the Poe Chat Interface. OpenAI models, including GPT-3.5T, GPT-4, GPT-4T, and GPT-4o, were utilized via the OpenAI API. We also tested the performance of two pretrained language models, BERT18 and ClincicalBERT19, which are fine-tuned versions of BERT on medical text. A list of all LLMs and times of use, as well as model parameters, is available in Supplementary Table S2.
Zero-shot classification
Zero-shot classification is an approach in prompt engineering in which the prompt is given to the model without any training. This approach is used in transfer learning, where a model used for different purposes is employed instead of fine-tuning a new model, thereby reducing the cost of training the new model. To perform zero-shot classification, we used eight different LLMs and two LMs. We provided each patient’s history as input to predict whether the patient would die or survive and then stored the results.
Fine-tuning LLM
We fine-tuned one of the open-source LLMs, Mistral-7b-Instruct-v0.2, which is a GPT-like large language model with 7 billion parameters. It is trained on a mixture of publicly available and synthetic data and can be used for natural language processing (NLP) tasks. It is also a decoder-only model that is used for text-generation tasks. Fine-tuning an LLM is usually considered time-consuming and expensive; recently, several methods have been introduced to reduce costs. We implemented the QLoRA fine-tuning approach to optimize the LLM while minimizing computational resources20.
The model was configured for 4-bit loading with double quantization, utilizing an “nf4” quantization type and torch.bfloat16 compute data type. A 16-layer model architecture with Lora attention and targeted projection modules was employed. We used the PEFT library to create a LoraConfig object with a dropout rate of 0.1 and task type ‘CAUSAL_LM’. The training pipeline, established via the transformer library, consisted of 4 epochs with a per-device batch size of 1 and gradient accumulation steps of 4. We utilized the “paged_adamw_32bit” optimizer with a learning rate of 2e-4 and a weight decay of 0.001. Mixed-precision training was conducted via fp16, with a maximum gradient norm of 0.3 and a warm-up ratio of 0.03. A cosine learning rate scheduler was employed, and training progress was logged every 25 steps and reported to TensorBoard. This methodology, which combines QLoRA with the Bitsandbytes library, enables efficient enhancement of our language model while significantly reducing resource requirements, demonstrating superior performance across various instruction datasets and model scales. A more detailed description is provided in Supplementary Section S2.
CML and LLM performance on different sample sizes
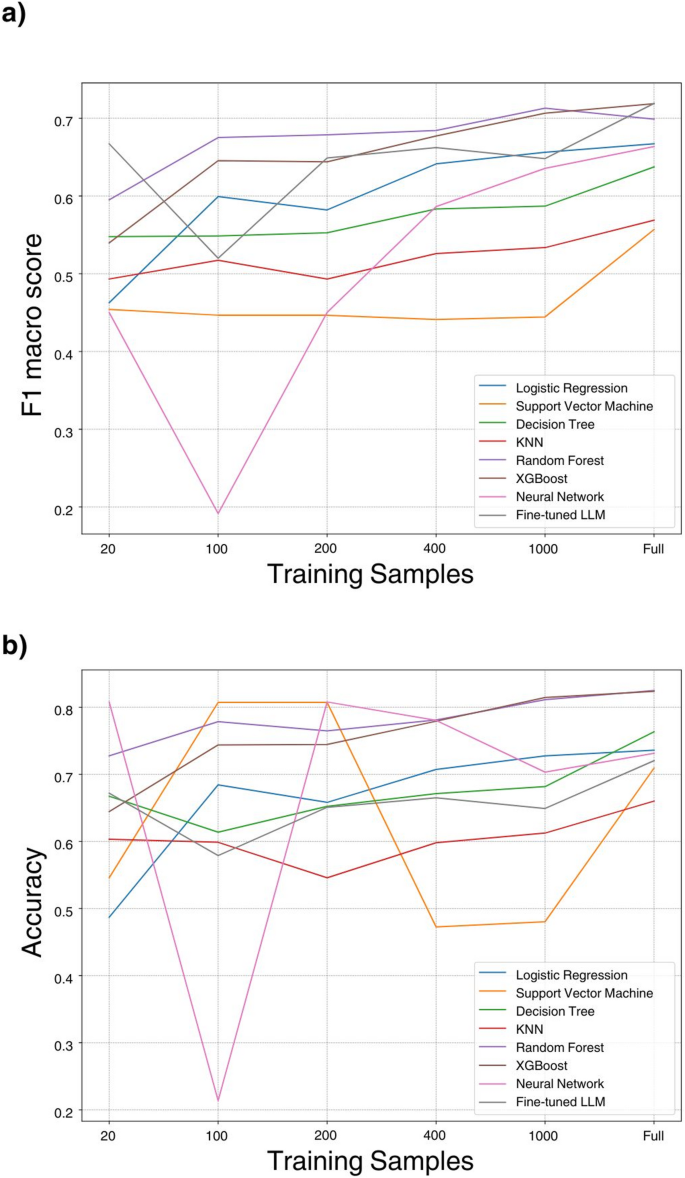
To investigate the influence of training sample sizes on model performance, we conducted a series of experiments using varying sample sizes: 20, 100, 200, 400, 1000, and 6118. Multiple models were trained using these sample sizes, and their performance was evaluated on the basis of the F1 score and accuracy metrics via an internal test set. The objective of this exploration was to gain valuable insights into the correlation between the volume of training data and the accuracy of predictive models.
Evaluation and cross-validation
The accuracy of the outputs was assessed by comparing them against a ground truth that categorized outcomes as either mortality or survival. Outputs from the LLM were similarly classified. If an LLM initially produced an undefined result, the prompt was repeatedly presented up to five times to elicit a defined prediction; these instances are documented in Supplementary Table S2. We evaluated the models’ performance via five critical metrics: specificity, recall, accuracy, precision, and F1 score. To optimize our models, we employed a grid search strategy with accuracy as the primary criterion.
We further implemented 5-fold cross-validation on the training dataset (n = 6,118). The training data were randomly partitioned into five equal-sized subsets. For each fold, four subsets were used for training while the remaining subset served as a validation set. This process was repeated five times, with each subset serving as the validation set once. We calculated performance metrics (accuracy, precision, recall, specificity, F1 score, and AUC) for each fold and reported the mean and standard deviation across all five folds.
Statistical analysis
Baseline characteristics were compared between patients who died and those who survived using appropriate statistical tests based on variable type and distribution. Continuous variables were analyzed using the Mann-Whitney U test (chosen over parametric alternatives due to non-normal distributions typical of clinical data) and presented as mean ± standard deviation. Categorical variables were compared using Pearson’s chi-square test. The area under the receiver operating characteristic curve (AUC) was used to illustrate the predictive capacity of each model. All statistical tests were two-sided with significance set at P < 0.05. Statistical analyses were performed using Python 3.12 with SciPy (v1.16.2).
Explainability
In our study, we employed SHAP (SHapley Additive exPlanations) values to examine both the total (global) and individual (granular) impacts of features on model predictions. We normalized the numerical data via a standard scaler and adopted a model-agnostic methodology. Our model-agnostic approach involved employing XGBoost as the explainer model for LLMs prediction, which was chosen for its robust performance, as demonstrated in prior research and our own findings. SHAP values provide a clear, quantitative assessment of how each feature influences individual predictions, enhancing transparency in the model’s decision-making process.
For our analysis, we used the test set for each model, generated SHAP values for every prediction, and computed the mean and standard deviation of the absolute SHAP scores. We then converted SHAP scores from a range of 0 to 1 into “global impact percentages” by dividing each feature’s score by the total score of all features and multiplying by 100. We calculated the average impact percentages for both CMLs and LLMs by first averaging the SHAP scores and then determining the impact percentages. To compute the standard deviation of the impact percentages, we adjusted the average standard deviation of CML/LLM via a multiplication factor derived from the ratio of the impact score to the SHAP mean. The global impact percentage represents the proportion of each feature’s impact on the predicted class across the entire dataset. A violin plot visually represents the variability of each input feature’s effect on the output.