Analysis with the DDE model
First, we reproduce the results of the DDE model developed in Ref.2. The model consists of six compartments: susceptible (S), exposed (E), infectious (I), confined-to-bed (B), convalescent (C), and removed (R). It assumes that individuals spend fixed durations in the E and I compartments, allows newly infected individuals to bypass the E stage, accounts for pre-existing immunity, and incorporates Erlang-distributed residence times in compartments B and C. The latter is implemented by introducing ten consecutive sub-compartments, \(B_j\) and \(C_j\) (\(j = 1, \ldots , 10\)), connected by equal-rate linear transfer terms. The model is characterized by eight parameters: the initial numbers of immune (\(R_{t=0}\)) and exposed (\(E_{t=0}\)) individuals, the infectivity coefficient (\(\beta _I\)), the probability (\(\rho _{\textrm{noE}}\)) that infected individuals bypass the E stage, the residence times \(T_E\) and \(T_I\) in the exposed and infectious stages, respectively, and the average transition rates \(\delta\) (from B to C) and \(\epsilon\) (from C to R). It is defined by the following set of equations:
$$\begin{gathered} \frac{{dS}}{{dt}} = – \beta _{I} \frac{{SI}}{N}, \hfill \\ \frac{{dE}}{{dt}} = \frac{{(1 – \rho _{{noE}} )\beta _{I} }}{N}(SI – S(t – T_{E} )I(t – T_{E} )), \hfill \\ \frac{{dI}}{{dt}} = \frac{{(1 – \rho _{{noE}} )\beta _{I} }}{N}(S(t – T_{E} )I(t – T_{E} ) – S(t – T_{E} – T_{I} )I(t – T_{E} – T_{I} )) + \frac{{\rho _{{noE}} \beta _{I} }}{N}(SI – S(t – T_{I} )I(t – T_{I} )), \hfill \\ \frac{{dB_{1} }}{{dt}} = \frac{{(1 – \rho _{{noE}} )\beta _{I} }}{N}S(t – T_{E} – T_{I} )I(t – T_{E} – T_{I} ) + \frac{{\rho _{{noE}} \beta _{I} }}{N}S(t – T_{I} )I(t – T_{I} ) – 10\delta B_{1} , \hfill \\ \end{gathered}$$
$$\begin{gathered} \frac{{dB_{j} }}{{dt}} = 10\delta (B_{{j – 1}} – B_{j} ),j = 2,…,10, \hfill \\ \frac{{dC_{1} }}{{dt}} = 10\delta B_{{10}} – 10 \in C_{1} , \hfill \\ \frac{{dC_{j} }}{{dt}} = 10 \in (C_{{j – 1}} – C_{j} ),j = 2,…,10, \hfill \\ \frac{{dR}}{{dt}} = 10 \in C_{{10}} \hfill \\ \end{gathered}$$
with the initial conditions
$$[S,E,I,B_1,…,C_1,…,R](t=0)=[N-E_{t=0}-R_{t=0}, (1-\rho _{noE})E_{t=0},0,…,0,…,R_{t=0}].$$
Following Avilov et al. (2004)2, the DDE model is fitted to the data by minimising the weighted sum of squared residuals for the variables B (the number of confined-to-bed students), C (the number of convalescent students), and AR (the total attack rate), with weights of 1, 0.4, and 10, respectively. Following the protocol in Ref.2, we employ a gradient descent method for numerical optimisation (using the fmincon function in Matlab). The fitting process is repeated 50 times from random starting points drawn from uniform distributions over the permitted parameter range (for more details, see section S1 of the Supplementary Material) to mitigate issues with local minima. Finally, the solution providing the best fit to the data is selected.
Note that the results of the DDE model presented here differ slightly from those reported in Ref.2. This is because we fit the DDE model to the 14 time points provided in the original paper1, whereas Avilov et al. (2004)2 used an extended dataset with 26 time points, assuming that the prevalence of confined-to-bed and convalescent students beyond the original range was zero.
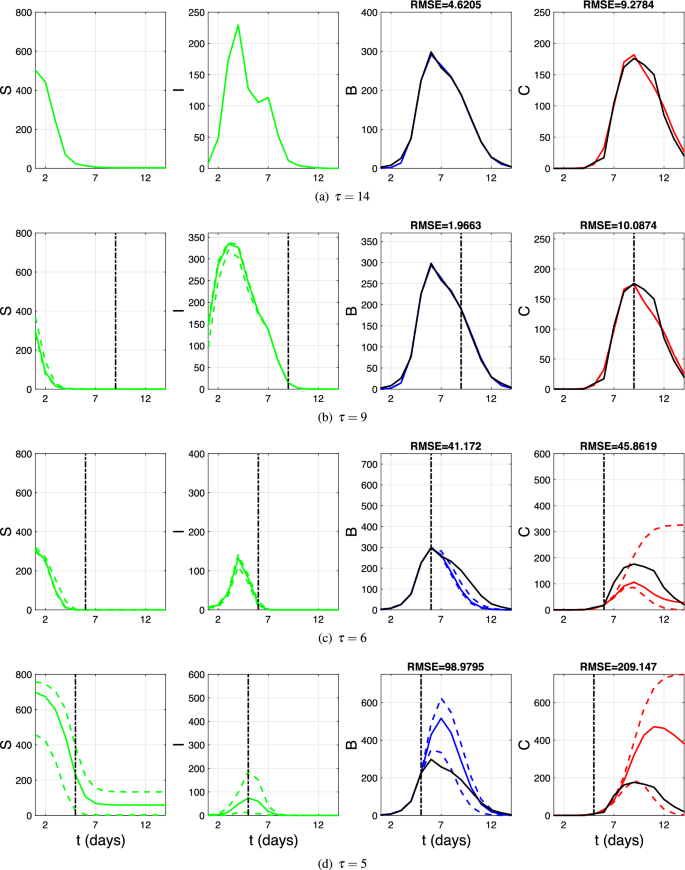
Forward predictions of the DDE model based on the first \(\tau\) time points (training dataset). Shown are the results for the number of susceptible (S), infected (I), confined-to-bed (B), and convalescent (C) individuals, respectively. The model outcomes (non-black lines) are shown alongside the data (solid black lines). Solid non-black lines correspond to the average trajectories, while dashed lines indicate 95% credible intervals. (b–d) The results are based on the top \(M=100\) (out of \(10^5\)) simulations with the best fit in the first \(\tau\) time steps. (a) \(\tau =14\) (entire epidemic); (b) \(\tau =9\) (peaks in curves B and C reached); (c) \(\tau =6\) (only the peak in curve B reached); (d) \(\tau =5\) (neither B nor C peaks reached). For panels (b–d), RMSE values are calculated based on the testing dataset.
Next, we assess the predictive power of the DDE model by training it on only the first \(\tau\) time steps of the epidemic (training dataset), generating forward predictions, and evaluating the goodness-of-fit of these predictions against the remaining time steps (testing dataset). The fitting protocol is adjusted for the reduced dataset size. Specifically, since the total attack rate (AR) is unknown until the epidemic concludes, it is excluded from the DDE fitting procedure. Instead, the model is fitted by minimising the weighted sum of squared residuals for variables B and C using the weights specified in Ref.2.
We consider three values of \(\tau\): \(\tau =9\), \(\tau =6\), and \(\tau =5\), corresponding to the peak in curve C, the peak in curve B, and one step before the peak in curve B, respectively. Notably, in all cases, the DDE model predicts that the peak infection prevalence has already occurred (see Figure 1).
For model fitting, we follow the protocol outlined in Ref.2. However, given the small sample size of the training dataset-where the number of model parameters approaches the number of data points-the objective function exhibits numerous local minima. To address this issue, we conduct \(10^5\) simulations. In each simulation, we apply the Avilov et al.2 protocol with 50 runs. We then select the top M simulations with the best fit, yielding M vectors of parameter estimates that approximate an optimal fit. This sample enables us to compute average parameter estimates and construct credible intervals. The parameter M represents the sample size tolerance level, which is set to \(M=100\) by default.
As shown in Figure 1, the predicted trajectories align well with the data when the peaks of both curves B and C have already been reached. However, further reducing \(\tau\) has a dramatic effect on predictive accuracy. For \(\tau =5\), the root mean square errors for compartments B and C (\(RMSE_B\) and \(RMSE_C\), respectively) increase twentyfold compared with the ex post estimation. Notably, when \(\tau\) is relatively small, the actual trajectories either fall outside the credible intervals produced by the DDE model or fluctuate near their boundaries (see Figures 1c and 1d). This result remains unaffected by the tolerance parameter M (see Figures S4–S6 in the Supplementary Material), indicating that the DDE model is unreliable for predicting epidemic progression in its early stages.
Finally, we assess the robustness of the parameter estimates under these data perturbations-i.e., changes in the number of data points \(\tau\) in the training dataset. The DDE model’s parameter estimates vary dramatically as the training set size decreases (see Table 1 and Figures S1–S3 in the Supplementary Material). In particular, the estimated basic reproduction number \(R_0\) initially triples, rising from 7.51 for \(\tau =14\) (full dataset) to 19.09 for \(\tau =6\), before dropping fivefold to 3.86 for \(\tau =5\). These fluctuations in \(R_0\) are accompanied by significant changes in the estimated number of immune individuals at \(t=0\), \(R_{t=0}\), which first doubles and then decreases sevenfold as \(\tau\) decreases, along with similar variations in the total attack rate (AR).
The key takeaway is that while the DDE model achieves an excellent ex post fit over the full course of the epidemic, it performs poorly in forward prediction, particularly in the early stages. Moreover, its parameter estimates are highly sensitive to the training set size, and combined with biological concerns-such as unrealistically high \(R_0\) values-this raises serious doubts about the reliability of the model’s estimates.
Analysis with the SIR model
As an alternative to the DDE model, we introduce a stochastic SIR-based modelling approach in this section. Given the relatively small population size and the significant role of random events in disease outbreaks within such settings, we adopt a stochastic formulation of the standard SIR model, governed by the following reactions:
$$\begin{aligned} \text {S} + \text {I} \overset{\beta }{\longrightarrow }\ 2{\text {I}},\\ \text {I} \overset{\gamma }{\longrightarrow }\ \text {B},\\ \text {B} \overset{\delta }{\longrightarrow }\ \text {C},\\ \text {C} \overset{\nu }{\longrightarrow }\ \text {R}. \end{aligned}$$
Here, S, I, B, C, and R represent the susceptible, infectious, confined to bed, convalescent, and removed compartments, respectively. We assume that the third reaction is a delayed process with a fixed time delay of 1.5 days. This reflects the fact that students confined to bed were transferred to the convalescent compartment 1.5 days after their temperature returned to normal. Essentially, this means an individual spent a random time duration in compartment B until their fever subsided, followed by an additional 1.5 days before moving to compartment C1.
To estimate the model parameter vector \(\varvec{\theta }=(\beta ,\gamma ,\delta ,\nu )\), we employ the Approximate Bayesian Computation algorithm (vanilla ABC)14. We first sample the proposal parameters \(\varvec{\theta ^{\#}}\) from the following priors:
$$\beta \sim U[0,0.008],$$
$$\gamma \sim U[0.5,3.5],$$
$$\delta \sim U[0,1.6],$$
where U[a, b] denotes the uniform distribution on the interval [a, b]. The sampled proposal parameters are then used to simulate the stochastic SIR model using a modification of the Gillespie algorithm that takes into account delayed reactions15,16,17,18 (see section S3 of the Supplementary Material for more details). For each simulation run, by discretizing time, we obtain realizations of the stochastic process at time steps \(t \in \{1,..,14\}\): \(S_{t}\), \(I_{t}\), \(B_{t}\), \(C_{t}\), and \(R_{t}\). We then determine the summary statistics
$$\varvec{x}=[B_1,…,B_{14},C_1,…,C_{14},AR],$$
where AR is the total attack rate (i.e., the total number of students affected by the epidemic, \(AR = 763 – S(\infty )\)). Comparing \(\varvec{x}\) with the actual number of confined to bed students, convalescent individuals, and the total number of affected individuals-the information stored in the vector \(\varvec{x’}\)-the vector of proposal parameters \(\varvec{\theta ^{\#}}\) is accepted if the distance between the model output and the data is relatively small, i.e., if
$$\psi =\frac{||\varvec{x}-\varvec{x’}||^2}{||\varvec{x’}||^2} \le \epsilon,$$
where \(\epsilon\) is the tolerance. We set \(\epsilon=0.03\) and run our simulations until the sample size reaches \(10^4\).
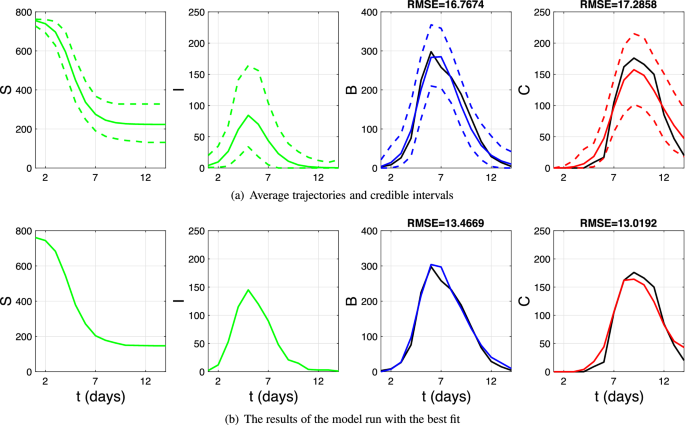
The results of the stochastic SIR model (retrospective fit) for the number of susceptible (S), infected (I), confined-to-bed (B), and convalescent (C) individuals. Shown are the model outputs (non-black lines) and fit to the data (solid black lines). (a) Solid non-black lines correspond to the average trajectories, while dashed lines show the \(95\%\) credible intervals. (b) The results of a simulation run with the smallest sum of the \(RMSE_B\) and \(RMSE_C\) values.
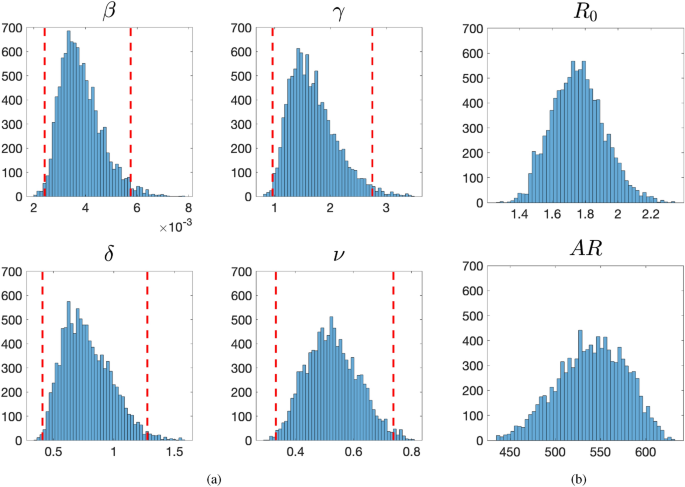
The results of the stochastic SIR model (retrospective fit). (a) Posterior distributions of the model parameters with 95% credible intervals. (b) Posterior distributions of the basic reproduction number \(R_0\) and the total attack rate AR. The posterior distributions are derived from a sample of \(10^4\) simulation runs of the stochastic SIR model, generated using the vanilla Approximate Bayesian Computation (ABC) algorithm with a tolerance parameter of \( \epsilon= 0.03\). Credible intervals were computed via kernel density estimation using Matlab’s mvksdensity function, with bandwidth selection determined according to Silverman’s rule of thumb..
Retrospective fit
As shown in Figure 2a, the stochastic SIR model fits the data quite well. As expected, it has somewhat larger root mean squared errors than the DDE model (16.8 vs 4.6 in the variable B and 17.3 vs 9.3 in the variable C). A better fit can be achieved in some individual SIR simulation runs. For example, the run with the smallest sum of the root mean squared errors in variables B and C has \(RMSE_B=13.5\) and \(RMSE_C=13.0\) (see Figure 2b). Furthermore, as shown in Figure S7 in the Supplementary Material, SIR runs with the best fit in the variables B and C, respectively, are characterised by RMSE values that are comparable with those of the DDE model (\(RMSE_B=9.3\) and \(RMSE_C=8.2\), respectively). This contradicts the claim that the fit of the SIR model, even in the B curve alone, is not satisfactory2. Note that decreasing the tolerance parameter \(\epsilon\) will further improve the goodness-of-fit of the stochastic SIR model (albeit requiring more extensive simulations).
The posterior distributions of the SIR model parameters, along with their 95% credible intervals, are shown in Figure 3a. These intervals are obtained using kernel density estimation19, implemented via the mvksdensity function in Matlab, with bandwidth selection determined by Silverman’s rule of thumb20. The corresponding credible intervals for the stochastic SIR model trajectories are presented in Figure 2a.
The average estimated total attack rate is 540 individuals (see Figure 3b), which closely aligns with the actual value of \(AR=512\) students. Notably, our model estimates the basic reproduction number \(R_0\) at an average value of 1.76 (see Figure 3b), which falls well within the established range of \(R_0 \in [1,4]\) for various influenza outbreaks2,11,12,13. This stands in stark contrast with the DDE model, which produces an unrealistically high estimate of \(R_0 = 7.51\). Moreover, the DDE model’s large \(R_0\) estimate leads to an inferred initial number of immune individuals of \(R_{t=0} = 249\), implying that nearly all uninfected students were classified as immune. In contrast, the stochastic SIR model does not require an initial population of immune individuals, making it less restrictive in terms of underlying biological and social assumptions.
Overall, while the stochastic SIR model does not achieve as good a retrospective fit as the DDE model-an expected outcome given the DDE model’s greater number of parameters and equations-it produces more biologically plausible estimates of \(R_0\) and avoids restrictive assumptions about immunity. This balance between model complexity and interpretability highlights the advantages of the stochastic SIR framework in epidemic modelling (Table 2).
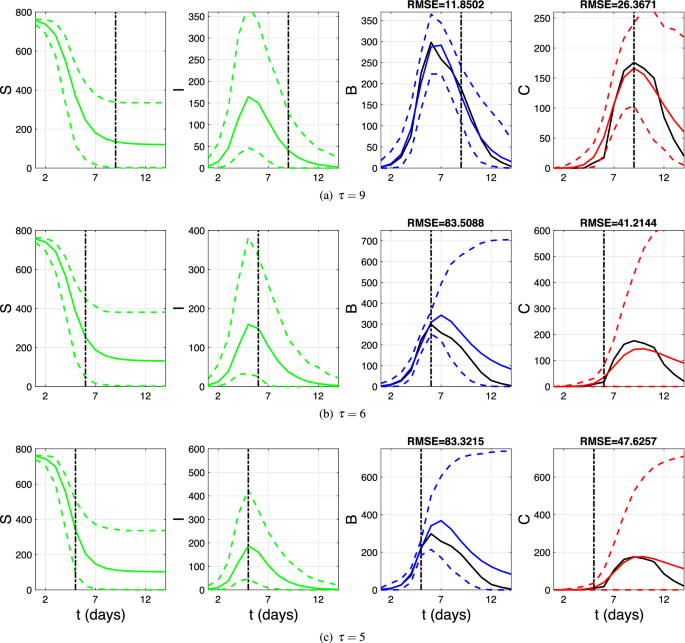
Forward predictions of the stochastic SIR model for the number of susceptible (S), infected (I), confined-to-bed (B), and convalescent (C) individuals based on the first \(\tau\) time points (training dataset). Shown are the model outcomes (non-black lines) and fit to the data (solid black lines). Solid non-black lines correspond to the average trajectories, while dashed lines show 95% credible intervals. (a) \(\tau =14\) (the entire epidemic); (b) \(\tau =9\) (the peaks in curves B and C are reached); (c) \(\tau =6\) (only the peak in curve B is reached); and (d) \(\tau =6\) (the peaks in curves B and C are not reached).
Forward predictions
Here, we evaluate the predictive performance of the stochastic SIR model by training it on only the first \(\tau\) time steps of the epidemic (training dataset), generating forward predictions, and assessing the goodness-of-fit of these predictions against the remaining time steps (testing dataset). To account for the reduced dataset size, we modify our fitting protocol: since the total attack rate (AR) is unknown until the epidemic concludes, it is excluded from the computation of the summary statistic x used in the Bayesian estimation of the stochastic SIR model.
As shown in Figure 4a, decreasing \(\tau\) significantly affects prediction accuracy. For \(\tau =5\), the root mean square errors (RMSE) in compartments B and C increase fivefold and threefold, respectively, compared with the ex post estimation. However, for \(\tau =5\), the stochastic SIR model exhibits slightly lower RMSE in compartment B and four times lower RMSE in compartment C compared with the DDE model, suggesting that the stochastic SIR model produces better forward predictions in the early epidemic stages. Notably, unlike the DDE model, the stochastic SIR model generates credible intervals that consistently contain the observed trajectories.
Furthermore, most parameter estimates of the stochastic SIR model remain stable as the training set size decreases (see Table 1 and Figures S8–S10 in the Supplementary Material). The only exception is the total attack rate, which increases from 540 to 661 as \(\tau\) decreases from 14 to 5. The estimated basic reproduction number \(R_0\) remains relatively robust to data perturbations, rising from 1.76 for \(\tau =14\) (full dataset) to 2.72 for \(\tau =5\), staying within the range of \(R_0\) estimates reported in the literature for various influenza outbreaks.
Overall, our findings lead to two key conclusions. First, while the stochastic SIR model provides a worse ex post fit than the DDE model, it generates more reliable forward predictions, particularly in the early stages of the epidemic. Second, the stochastic SIR model’s parameter estimates are significantly less sensitive to the size of the training set compared with those of the DDE model, further supporting its robustness in predictive applications.