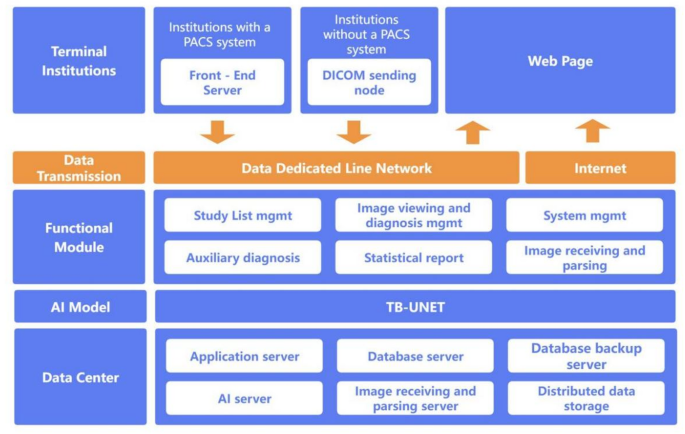
System architecture
The system adopts a front-end and back-end separation development model. The front-end is built using the Vue framework, while the back-end is developed using the Spring Boot framework. Data interaction between the front-end and back-end is conducted via HTTP protocol, with data transmitted in JSON format. This architecture allows for parallel development of the front-end and back-end, improving development efficiency and facilitating maintenance and expansion. The front-end focuses on user experience and interaction design, while the back-end handles business logic and data persistence. The core of the system is the deep learning-based TB screening function, which works closely with both the front-end and back-end to achieve an efficient screening process. The system architecture is shown in Fig. 1.
System functional modules
Study list management module
Presents examination, patient, and report information in a list format to reporting physicians and administrators.
Image viewing and diagnosis module
Provides physicians with a professional platform for reading images. The front-end includes image zooming and annotation functions, allowing physicians to observe image details. The back-end integrates deep learning algorithms to provide lesion area prompts, automatic generation of imaging findings, and conclusions, assisting physicians in diagnosis. This module works in conjunction with the TB diagnostic assistance module to improve diagnostic accuracy and efficiency.
System management module ensures stable and efficient system operation
User Management Submodule handles user registration login information modification and password reset with front end providing interface and back end handling information verification and storage.
Role Management Submodule defines assigns and manages system roles such as administrator physician patient and their corresponding permissions with front end providing interface and back end handling storage and verification.
Permission Management Module controls access to system functions based on user roles with front end dynamically displaying menus and back end verifying permissions through security framework.
Department Management Submodule manages department information and hierarchical relationships facilitating user and business classification with front end displaying tree structure and back end handling data persistence.
Report Template Management Submodule designs edits and manages TB screening report templates to ensure completeness and standardization with front end providing visual interface and back end handling storage and template generation.
Log Management Submodule records system operation logs for tracking troubleshooting and security audits with front end providing query functions and back end handling logging and storage.
Diagnostic assistance module
Interacts with the TB-UNET model to provide lesion delineation, imaging findings, and conclusion generation for radiologists in the image reading and diagnosis module.
Statistical reporting module analyzes key business data to support decision-making
Individual Workload Statistics Module tracks screening workload by user such as number of cases screened with front end displaying data in charts and tables and back end extracting and calculating data.
Hospital Workload Statistics Module tracks screening workload by hospital such as total number of cases screened by department with front end displaying comprehensive reports and back end aggregating data.
Image Upload and Review Statistics Module tracks number of images uploaded time distribution review progress and results to optimize related processes with front end displaying visual data and back end collecting and analyzing data.
Image reception and parsing module
The system supports multiple image reception methods, including direct import from medical devices and local file uploads. Upon receiving images, the system performs initial format validation to ensure image integrity and usability.
Network topology
The system includes 216 terminal institutions in the Kashgar region, comprising 4 tertiary hospitals, 17 secondary hospitals, and 178 township hospitals, as well as some private hospitals. To ensure the security of medical data, all terminal institutions are connected to the data center via a star-topology dedicated network. This connection method optimizes data transmission efficiency and provides a robust security framework for medical data, ensuring stability and security during transmission and storage. The network topology is shown in Fig. 2.
Additionally, the system has established a comprehensive security protection system. All medical institutions are equipped with network firewalls to effectively defend against external network attacks and ensure internal network security. Network access switches are deployed to achieve efficient and stable network connections. Furthermore, to enhance network security and management convenience, all institutions are divided into different VLANs based on their business needs and security standards. These VLANs are isolated from each other, reducing the scope of broadcast domains and minimizing the risk of network storms. This isolation also limits unauthorized access, ensuring that an attack on one VLAN does not affect the normal operation of others.
Diagnostic assistance model
Dataset
The TB screening AI model was trained using a public dataset from developed cities in China and hospital CXR images. Testing and evaluation were conducted using a dataset from remote areas (Table 1).The training set (10,002 cases) was collected from 5 tertiary hospitals across 3 provinces in China, while the testing set (895 cases) included data from 12 county hospitals and 178 township hospitals in the Kashgar region (stratified by hospital level: 4 tertiary, 17 secondary, 178 primary).
The study population had a mean age of 42.6 ± 12.8 years (range 15–89 years), with 58.0% males. TB prevalence in the test set (895 cases) was 29.5% (264 positive cases). Comorbidities were present in 12.7% of participants, mainly hypertension (6.2%) and diabetes (3.8%).
The TB cases in both training and testing datasets were diagnosed in accordance with the WS 288–2017 Diagnostic Criteria for Tuberculosis through a comprehensive approach: All TB cases were confirmed by at least one positive result in etiological tests, including sputum smear microscopy, sputum culture, or Xpert MTB/RIF assay. For cases with negative etiological tests but typical TB imaging features (e.g., pulmonary polymorphic lesions, cavity formation) and clinical symptoms consistent with TB (e.g., cough, hemoptysis, low-grade fever), which responded effectively to anti-tuberculosis treatment, they were identified as clinically diagnosed TB cases and included in the dataset.
The dataset in this study was primarily derived from individuals participating in annual TB screening campaigns in the Kashgar region, which may introduce potential selection bias. Most participants were either symptomatic individuals or those at high risk of TB (e.g., close contacts of TB patients), which may differ from the broader population including asymptomatic individuals or those seeking medical care for unrelated conditions. To mitigate this, the training set included a subset of 1,200 asymptomatic individuals (identified through community health surveys) with negative TB screening results, and the testing set included 86 cases of individuals presenting with non-TB related respiratory symptoms (e.g., acute bronchitis) to enhance the model’s adaptability to diverse populations.
Data preprocessing
Three senior radiologists annotated the chest X-rays, which served as the gold standard for training the deep neural network. The senior radiologists, all with over 10 years of experience in reading chest X-rays, labeled the images as 0 (normal), 1 (TB), or other (non-TB abnormalities). After annotation, the dataset underwent necessary preprocessing, including resizing images to 512 × 512 pixels, normalizing pixel values, and augmenting data through random transformations of brightness, contrast, and saturation.
Three senior radiologists (with over 10 years of experience in chest X-ray interpretation) reviewed all chest X-ray images in both the training and testing datasets. Among them, the 10,002 images in the training set were annotated before model training, serving as the gold standard for model learning; the 895 images in the testing set were used for performance evaluation after model training, and their annotation results served as an independent reference standard without participating in the model training process to ensure the objectivity of the test.
Model confidence scores and testing
The team used a variant of the U-Net model, called TB-UNet, which replaces the encoder to detect TB lesions on CXR images. U-Net is a popular medical image segmentation network, consisting of a U-shaped fully convolutional neural network without fully connected layers. In the U-Net structure, upsampling operations generate rich feature channels, allowing the network to propagate contextual information to higher-resolution layers. The contracting and expanding paths are somewhat symmetrical, forming a U-shape.
The details of the deep learning model, confidence scores, and testing have been thoroughly discussed in another published article by the team (Nijiati P, et al. A Deep Learning Model for Tuberculosis Detection in Chest X-Rays: Development and Validation. Journal of Medical Imaging, 2024, 11(2): 123–135), so they will not be repeated here.
In summary, in the validation of 10,002 training cases and 895 testing cases, without AI assistance, radiologists had 165 true positives (TP), 99 false negatives (FN), 671 true negatives (TN), and 4 false positives (FP) in TB diagnosis; with AI assistance, TP was 205, FN was 71, TN was 658, and FP was 17. With the assistance of this system, radiologists’ sensitivity in diagnosing TB increased by 11.8% (from 62.7% [95% CI 58.2–67.1%] to 74.5% [95% CI 70.3–78.4%], p p = 0.023), and the average time spent on reading images decreased from 38.83 s (95% CI 36.21–41.45 s) to 15.93 s (95% CI 14.32–17.54 s, p 2).
Subgroup analysis showed consistent performance across age (sensitivity 73.1–75.2%), sex (74.2–74.7%), and comorbidity status (72.3–75.0%), with no significant differences (all p > 0.05).
The TB-UNet model was optimized for the characteristics of TB lesions in the Kashgar region, such as irregular borders and low contrast in CXR images. The encoder was replaced with a pre-trained ResNet-50 backbone to enhance feature extraction of subtle lesions. During training, a weighted cross-entropy loss function was used to address class imbalance (TB cases accounted for 26.3% of the training set), with weights set as 1:3 for non-TB and TB samples.
In internal validation (n = 1000), the TB-UNet model achieved a sensitivity of 78.3%, specificity of 96.5%, and F1-score of 0.82. The model’s lesion detection accuracy (IoU ≥ 0.5) for typical TB manifestations (e.g., pulmonary nodules, infiltrates) was 81.2%, and 72.5% for atypical manifestations (e.g., pleural effusion).
To compare radiologist performance with and without the AI system, inferential statistical tests were conducted. Specifically, the McNemar’s test was used to analyze differences in sensitivity, specificity, and accuracy (categorical variables), while the paired t-test was applied to compare average reading time (continuous variable). For assessing the stability of the TB-UNet model, 5-fold cross-validation was performed on the training set (10,002 cases), with each fold containing stratified samples to maintain the same proportion of TB and non-TB cases as the original dataset. Additionally, bootstrapping (1000 iterations) was used to estimate the robustness of model performance metrics.
The performance metrics used in this study are defined as follows:
Sensitivity: The proportion of actual TB cases correctly identified by the diagnostic process, calculated as (True Positives) / (True Positives + False Negatives) × 100%. It reflects the ability to avoid missed diagnoses of TB.
Specificity: The proportion of actual non-TB cases correctly identified, calculated as (True Negatives) / (True Negatives + False Positives) × 100%. It indicates the ability to avoid misdiagnosing non-TB cases as TB.
Accuracy: The overall proportion of correctly classified cases (both TB and non-TB), calculated as (True Positives + True Negatives) / (Total Cases) × 100%. It represents the general diagnostic correctness.
AUC (Area Under the ROC Curve): A comprehensive metric that quantifies the overall performance of a diagnostic classifier across all possible threshold settings, ranging from 0 to 1 (1 indicating perfect discrimination).
Kappa Value: A statistic that measures inter-rater agreement, accounting for chance agreement. Values range from − 1 to 1, with 1 indicating perfect agreement and 0 indicating agreement no better than chance.
The threshold for determining TB positivity was based on the integrated judgment of imaging features and clinical criteria, in line with WS 288–2017 Diagnostic Criteria for Tuberculosis. Specifically, a case was classified as TB-positive if it met either of the following:
Presence of typical TB imaging manifestations (e.g., pulmonary nodules with irregular borders, cavitary lesions, or infiltrates in the upper lobes) confirmed by at least two senior radiologists;
Positive results from etiological tests (sputum smear microscopy, sputum culture, or Xpert MTB/RIF assay) regardless of imaging findings.
To address potential information bias, the reference standard (ground truth) was established independently of the radiologists involved in model-assisted diagnosis. The three senior radiologists responsible for labeling the dataset did not participate in the subsequent AI-assisted diagnostic process. Additionally, for cases where radiological interpretation alone might be ambiguous (e.g., non-specific infiltrates), bacteriological results or 6-month clinical follow-up (to confirm treatment response) were used as supplementary evidence to finalize labels, avoiding circular validation where radiologist interpretation serves as both the reference standard and the outcome being evaluated.