Identification of DEGs between the ACHBLF and normal groups
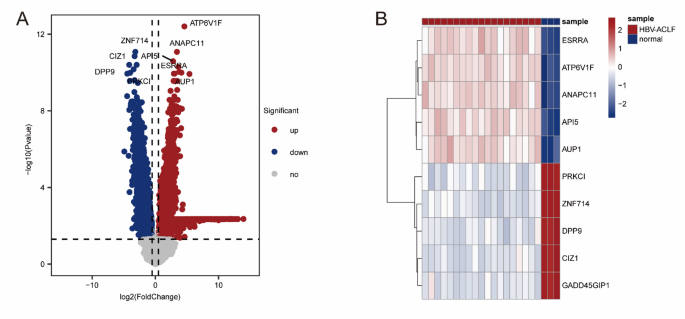
A total of 7461 DEGs were found between the ACHBLF and normal groups when |log2Fold Change|> 0.5 and P Value 1 A, Table S2). The top 5 most significantly upregulated DEGs were ESRRA, ATP6V1F, ANAPC11, API5 and AUP1, and the top 5 most significantly downregulated DEGs were PRKCI, ZNF714, DPP9, CIZ1 and GADD45GIP1(Fig. 2B).
Screening and functional enrichment of NRGs-related DEGs
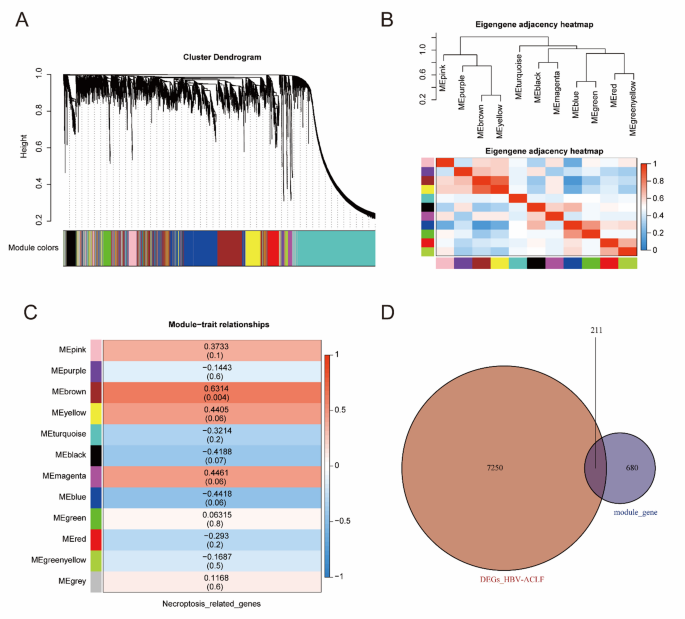
To screen NRG-related DEGs, WGCNA was performed, and 11 co-expression modules were acquired. Genes that were not assigned to any specific module were categorized into grey modules and excluded from further analysis (Fig. 3A). Module eigengenes (MEs) were used to explore the relationships and correlations of each module, and a dendrogram and heatmap were used to visualize the eigengene network (Fig. 3B). To elucidate the physiological significance of genes within modules, we correlated 12 MEs with NRGs and identified the most significant associations. On the basis of the module-trait relationships heatmap (Fig. 3C, Table S3), genes enriched in the brown module (r = 0.6314, P 3D, Table S5).
Screening of the NRGs-related DEGs in ACHBLF. (A) WGCNA modules with different colors under the clustering tree. (B) Correlations and relationships among modules. (C) Heat map of module-trait relationships. (D) The Venn plot identified 211 NRGs-related DEGs among 680 module genes and 7250 DEGs of ACHBLF.
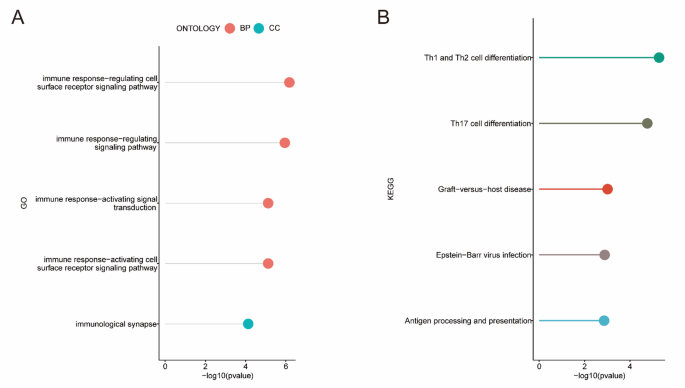
GO functional enrichment analyses revealed possible GO terms, including the CC terms “immunological synapse”, and BP terms “immune response-activating cell surface receptor signaling pathway”, “immune response-activating signal transduction”, “immune response-regulating signaling pathway”, and “immune response-regulating cell surface receptor signaling pathway” (Fig. 4B, Table S6). KEGG12,13,14 analysis revealed that 211 NRG-related DEGs were strongly associated with Th1 and Th2 cell differentiation, Th17 cell differentiation, graft-versus-host disease, Epstein-Barr virus infection and antigen processing and presentation (Fig. 4B, Table S7).
Screening by machine learning and functional enrichment of the hub genes
Three machine learning algorithms were used to screen hub genes. LASSO regression model identified 19 ACHBLF-related signature NRG-related DEGs (Fig. 5A and B), the RF algorithm was used to choose the top 30 genes as signature genes according to the feature weights (MDA and MDG) (Fig. 5C, D and E), and SVM-RFE identified 199 ACHBLF-related signature genes (Fig. 5F and G). A Venn diagram was used to identify the genes common to the three machine learning methods, and 5 NRG-signature genes (FCRL3, CDC14A, KLHL22, RALY, and MAP4K1) were identified as hub genes for subsequent analyses (Fig. 5H, Table S8).
Machine learning identified the signature NRGs-related DEGs of ACHBLF. (A, B) LASSO regression model screening of the signature NRGs-related DEGs of ACHBLF. (C) Random forest error rate versus the number of classification trees. (D) RF methods screen the signature genes according to MDA. (E) RF methods screen the signature genes according to MDG. (F, G) SVM-RFE selected the signature NRGs-related DEGs of ACHBLF. (H) Venn plot among three machine learning methods resulted in five hub genes of ACHBLF.
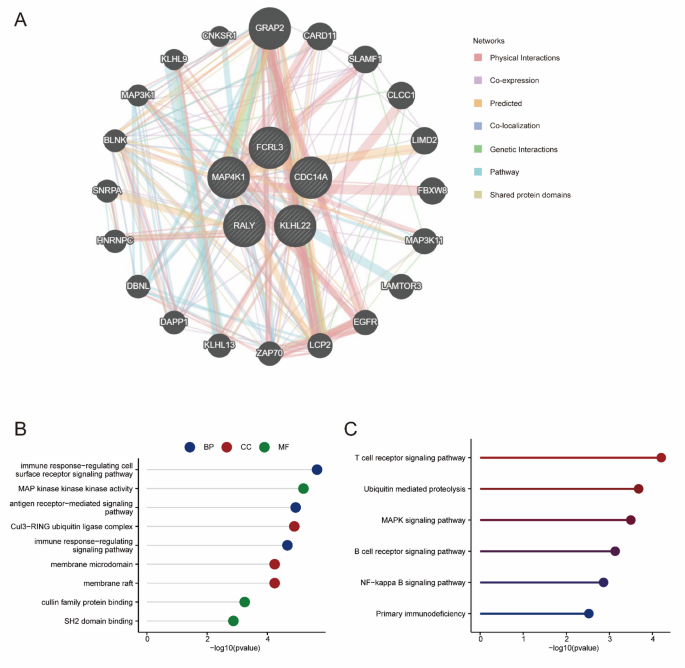
To investigate the biological roles of the hub genes in ACHBLF, protein-protein interaction (PPI) networks were constructed using GeneMANIA and analyzed by Cytoscape (Fig. 6A). GO and KEGG analyses were conducted to elucidate the potential signal pathways associated with the 5 hub genes and 20 hub gene-related genes. As indicated by the GO analysis, the genes were enriched in BPs, including the immune response-regulating cell surface receptor signaling pathway, the antigen receptor-mediated signaling pathway and the immune response-regulating signaling pathway. In terms of cellular components, these genes were predominantly enriched in the Cul3-RING ubiquitin ligase complex, membrane microdomains and membrane rafts. The genes were also enriched in molecular functions (MF), including MAP kinase kinase kinase activity, cullin family protein binding, and SH2 domain binding (Fig. 6B, Table S9). KEGG analysis revealed that these genes were strongly activated in the T cell receptor signaling pathway, ubiquitin-mediated proteolysis, MAPK signaling pathway, the B cell receptor signaling pathway, the NF-kappa B signaling pathway, and primary immunodeficiency (Fig. 6C, Table S10).
Validation the expression of the hub genes with the GEO databases
Because no data have been reported on FCRL3, CDC14A, KLHL22, RALY, or MAP4K1 in ACHBLF, we compared the expression of FCRL3, CDC14A, KLHL22, RALY, and MAP4K1 between PBMCs from ACHBLF patients and normal controls. As shown in Fig. 7A, the expression of MAP4K1 was significantly greater in ACHBLF patients than in normal controls; in contrast, the expression levels of FCRL3, CDC14A, KLHL22, and RALY were significantly lower in ACHBLF patients than in normal controls. Moreover, Spearman correlation analysis revealed that CDC14A was significantly positively associated with FCRL3 and KLHL22, and RALY was significantly negatively correlated with MAP4K1 (Fig. 7B).
In the analysis of the molecular pathways of the hub genes in ACHBLF, ssGSEA revealed that CDC14A, the expression of which was upregulated in ACHBLF, was enriched in primary immunodeficiency, autoimmune thyroid disease, graft versus host disease, antigen processing presentation and ribosome (Fig. 8A); FCRL3, the expression of which was upregulated in ACHBLF, was enriched in the intestinal immune network for IgA production, antigen processing and presentation, graft versus host disease, autoimmune thyroid disease and allograft rejection (Fig. 8B); KLHL22, the expression of which was upregulated in ACHBLF, was enriched in graft versus host disease, ribosome, allograft rejection, autoimmune thyroid disease and asthma (Fig. 8C); MAP4K1, the expression of which was upregulated in ACHBLF, was enriched in graft versus host disease, primary immunodeficiency, antigen processing presentation, allograft rejection and viral myocarditis (Fig. 8D); and RALY, the expression of which was upregulated in ACHBLF, was enriched in lysosome, epithelial cell signaling in helicobacter pylori infection, hematopoietic cell lineage, viral myocarditis and cell adhesion molecules (Fig. 8E).
Constructing a regulatory network of ceRNA, RBP and TFs
To explore the regulatory mechanisms of the hub genes in ACHBLF, an mRNA-miRNA-lncRNA interaction network was constructed, and the results revealed that the signature gene targeted by miR-424-5p is CDC14A and that the targeted lncRNA is XIST (Fig. 9A). Next, the starBase online database was used to search the mRNA/RBP pairs of the signature NRG-related DEGs, and the results revealed that 4 hub genes (CDC14A, MAP4K1, KLHL22 and RALY) had relevant pairing information. An RBP-mRNA regulatory network was generated using the starBase online database, which identified 24 nodes, 20RBPs, 4 mRNAs and 41 edges (Fig. 9B, Table S11). TFs are important regulators of genes expression. According to the hTFtarget database, interaction between 3 hub genes (CDC14A, MAP4K1, and RALY) and 2 TFs (GATA1 and CEBPB) were predicted (Fig. 9C, Table S12).
Constructing regulatory network of ceRNA, RBP and TFs. (A) An interaction network of mRNA-miRNA-lncRNA. Red color represented signature gene, yellow represented microRNA, and blue represented lncRNA. (B) RBP-mRNA regulatory network. Red color represented signature gene and blue represented RBP. (C) Predicted potential TFs of signature NRGs-related DEGs. Red color represented TFs and blue represented signature gene.
Evaluation of immune infiltration and correlation analysis of the hub genes
To compare immune infiltration between ACHBLF patients and normal controls, ssGSEA was conducted with 28 immune infiltration cell markers from the TISIDB database (Table S13). As shown in in Fig. 10A, the proportion of 28 immune cell subsets differed from one individual to another individual with ACHBLF (Fig. 10A), implying that immune cell may have crucial impacts on the pathophysiology of ACHBLF. Thus, immune cell infiltration was compared between ACHBLF patients and normal controls (Table S14). Compared with normal controls, ACHBLF patients had higher enrichment scores for immune infiltration of type 17 T helper cell, activated dendritic cell, plasmacytoid dendritic cell and monocytes (Fig. 10B). Additionally, correlations between the hub genes and immune cell infiltration were evaluated. As a consequence, the expression level of FCRL3 was negatively associated with plasmacytoid dendritic cells (R=−0.723, P R=−0.774, P R=−0.814, P R = 0.621, P = 0.002), activated CD8 T cells (R = 0.603, P = 0.003) and type 2 T helper cells (R = 0.648, P = 0.001). CDC14A expression was negatively correlated with activated dendritic cells (R=−0.674, P R=−0.621, P = 0.002). KLHL22 expression was negatively correlated with only activated dendritic cells (R=−0.673, P 11).
The correlations between hub genes and the infiltration level of the immune cells. (A) Correlation between FCRL3 and plasmacytoid dendritic cell. (B) Correlation between CDC14A and activated dendritic cell. (C) Correlation between FCRL3 and Monocyte. (D) Correlation between FCRL3 and activated dendritic cell. (E) Correlation between KLHL22 and activated dendritic cell. (F) Correlation between CDC14A and plasmacytoid dendritic cell. (G) Correlation between FCRL3 and natural killer cell. (H) Correlation between FCRL3 and activated CD8 T cell. (I) Correlation between FCRL3 and type 2 T helper cell.
Association of the hub genes with cancer hallmark pathways and correlation analyses
Comparisons of the 50 cancer hallmark pathways were performed between ACHBLF patients and normal controls. DNA repair, MTORC1 signaling, the P53 pathway, PI3K-AKT-MTOR signaling, TGF-β signaling and Wnt/β-catenin signaling were significantly activated in ACHBLF patients compared with normal controls. However, Hedgehog signaling was suppressed in ACHBLF patients compared with normal controls (Fig. 12, Table S15). We also analyzed the correlations between the 5 signature NRG-related DEGs and 50 cancer hallmark pathways. The expression of CDC14A was significantly negatively correlated with coagulation, adipogenesis, the xenobiotic metabolism pathway, etc. The expression levels of FCRL3 were negatively associated with the reactive oxygen species pathway, glycolysis, peroxisome, etc. KLHL22 expression was negatively associated with many pathways, such as spermatogenesis, myogenesis, and the late estrogen response. MAP4K1 expression was significantly associated with adipogenesis, angiogenesis and pancreas β cells. In contrast, RALY expression was positively correlated with the reactive oxygen species pathway, glycolysis, fatty acid metabolism, etc. (Fig. 13)
Identification of small-molecule drugs
MAP4K1 and highly associated genes were investigated to predict candidate drugs for ACHBLF therapy from the CMAP database (Table S16). The top 10 target drugs were tiaprofenic-acid, guanabenz, imatinib, etacrynic-acid, brivanib, vincamine, amiloride, lypressin, pizotifen and leflunomide (Table 1). However, no drugs have been approved by the National Medical Products Administration.